A Conjoint Analysis of Epilepsy and Migraine Through Network- And-Pathway-Based Method
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Anti-GRIK1 / Glur5 Antibody (ARG59676)
Product datasheet [email protected] ARG59676 Package: 50 μg anti-GRIK1 / GluR5 antibody Store at: -20°C Summary Product Description Rabbit Polyclonal antibody recognizes GRIK1 / GluR5 Tested Reactivity Hu, Ms, Rat Tested Application IHC-P, WB Host Rabbit Clonality Polyclonal Isotype IgG Target Name GRIK1 / GluR5 Species Human Immunogen Recombinant protein corresponding to R271-I450 of Human GRIK1. Conjugation Un-conjugated Alternate Names GluR5; GluK1; GLUR5; EEA3; GluR-5; Excitatory amino acid receptor 3; Glutamate receptor ionotropic, kainate 1; EAA3; Glutamate receptor 5; GLR5 Application Instructions Application table Application Dilution IHC-P 1:200 - 1:1000 WB 0.1 - 0.5 µg/ml Application Note IHC-P: Antigen Retrieval: Heat mediation was performed in Citrate buffer (pH 6.0) for 20 min. * The dilutions indicate recommended starting dilutions and the optimal dilutions or concentrations should be determined by the scientist. Properties Form Liquid Purification Affinity purification with immunogen. Buffer 0.9% NaCl, 0.2% Na2HPO4, 0.05% Sodium azide and 5% BSA. Preservative 0.05% Sodium azide Stabilizer 5% BSA Concentration 0.5 mg/ml Storage instruction For continuous use, store undiluted antibody at 2-8°C for up to a week. For long-term storage, aliquot and store at -20°C or below. Storage in frost free freezers is not recommended. Avoid repeated freeze/thaw cycles. Suggest spin the vial prior to opening. The antibody solution should be gently mixed before use. Note For laboratory research only, not for drug, diagnostic or other use. www.arigobio.com 1/4 Bioinformation Gene Symbol GRIK1 Gene Full Name glutamate receptor, ionotropic, kainate 1 Background Glutamate receptors are the predominant excitatory neurotransmitter receptors in the mammalian brain and are activated in a variety of normal neurophysiologic processes. -

A Guide to Glutamate Receptors
A guide to glutamate receptors 1 Contents Glutamate receptors . 4 Ionotropic glutamate receptors . 4 - Structure ........................................................................................................... 4 - Function ............................................................................................................ 5 - AMPA receptors ................................................................................................. 6 - NMDA receptors ................................................................................................. 6 - Kainate receptors ............................................................................................... 6 Metabotropic glutamate receptors . 8 - Structure ........................................................................................................... 8 - Function ............................................................................................................ 9 - Group I: mGlu1 and mGlu5. .9 - Group II: mGlu2 and mGlu3 ................................................................................. 10 - Group III: mGlu4, mGlu6, mGlu7 and mGlu8 ............................................................ 10 Protocols and webinars . 11 - Protocols ......................................................................................................... 11 - Webinars ......................................................................................................... 12 References and further reading . 13 Excitatory synapse pathway -

Chr21 Protein-Protein Interactions: Enrichment in Products Involved in Intellectual Disabilities, Autism and Late Onset Alzheimer Disease
bioRxiv preprint doi: https://doi.org/10.1101/2019.12.11.872606; this version posted December 12, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Chr21 protein-protein interactions: enrichment in products involved in intellectual disabilities, autism and Late Onset Alzheimer Disease Julia Viard1,2*, Yann Loe-Mie1*, Rachel Daudin1, Malik Khelfaoui1, Christine Plancon2, Anne Boland2, Francisco Tejedor3, Richard L. Huganir4, Eunjoon Kim5, Makoto Kinoshita6, Guofa Liu7, Volker Haucke8, Thomas Moncion9, Eugene Yu10, Valérie Hindie9, Henri Bléhaut11, Clotilde Mircher12, Yann Herault13,14,15,16,17, Jean-François Deleuze2, Jean- Christophe Rain9, Michel Simonneau1, 18, 19, 20** and Aude-Marie Lepagnol- Bestel1** 1 Centre Psychiatrie & Neurosciences, INSERM U894, 75014 Paris, France 2 Laboratoire de génomique fonctionnelle, CNG, CEA, Evry 3 Instituto de Neurociencias CSIC-UMH, Universidad Miguel Hernandez-Campus de San Juan 03550 San Juan (Alicante), Spain 4 Department of Neuroscience, The Johns Hopkins University School of Medicine, Baltimore, MD 21205 USA 5 Center for Synaptic Brain Dysfunctions, Institute for Basic Science, Daejeon 34141, Republic of Korea 6 Department of Molecular Biology, Division of Biological Science, Nagoya University Graduate School of Science, Furo, Chikusa, Nagoya, Japan 7 Department of Biological Sciences, University of Toledo, Toledo, OH, 43606, USA 8 Leibniz Forschungsinstitut für Molekulare Pharmakologie -

Case-Control Study of GRIA1 and GRIA3 Gene Variants in Migraine Jie Fang1, Xingkai An1, Shuai Chen2, Zhenzhen Yu1, Qilin Ma1,2* and Hongli Qu1*
Fang et al. The Journal of Headache and Pain (2016) 17:2 DOI 10.1186/s10194-016-0592-2 RESEARCH ARTICLE Open Access Case-control study of GRIA1 and GRIA3 gene variants in migraine Jie Fang1, Xingkai An1, Shuai Chen2, Zhenzhen Yu1, Qilin Ma1,2* and Hongli Qu1* Abstract Background: As the most abundant excitatory neurotransmitter in the central nervous system, glutamate has been accepted to play a major role in the pathophysiology of migraine. The previous studies have reported the glutamate receptor ionotropic GRIA1 and GRIA3 genes variants associated with migraine. The project aims to investigate the polymorphisms in both genes for their association with migraine in the Chinese Han population. Methods: A Han-Chinese case-control population, including 331 unrelated female migraine patients and 330 matched controls, was studied. Variants in genes (GRIA1 and GRIA3) were genotyped by Multiplex SNaPshot assay. Results: In the group of patients, the frequency of allele C was 84.1 % (557 C alleles) and allele T was 15.9 % (105 T alleles) for the GRIA1 (rs2195450) in migraineurs, this was significantly as compared with the controls (P = .001, OR = 1.786, 95 % CI: 1.28–2.49). And an association was also seen in the migraine with aura (MA) subtype (P = .012, OR = 2.092, 95 % CI: 1.17–3.76) and migraine without aura (MO) subtype (P =.002,OR=1.737, 95 % CI: 1.23–2.45). However, no evidence was found that GRIA1 (rs548294) or GRIA3 (rs3761555) is associated with migraine. Conclusion: Our data of this study confirmed the association of GRIA1 (rs2195450) to female migraine (MA, MO) susceptibility in the Chinese Han population. -

Kainate Receptors in Health and Disease
View metadata, citation and similar papers at core.ac.uk brought to you by CORE provided by Elsevier - Publisher Connector Neuron Review Kainate Receptors in Health and Disease Juan Lerma1,* and Joana M. Marques1 1Instituto de Neurociencias, CSIC-UMH, San Juan de Alicante, 03550 Spain *Correspondence: [email protected] http://dx.doi.org/10.1016/j.neuron.2013.09.045 Our understanding of the molecular properties of kainate receptors and their involvement in synaptic phys- iology has progressed significantly over the last 30 years. A plethora of studies indicate that kainate receptors are important mediators of the pre- and postsynaptic actions of glutamate, although the mechanisms under- lying such effects are still often a topic for discussion. Three clear fields related to their behavior have emerged: there are a number of interacting proteins that pace the properties of kainate receptors; their activity is unconventional since they can also signal through G proteins, behaving like metabotropic recep- tors; they seem to be linked to some devastating brain diseases. Despite the significant progress in their importance in brain function, kainate receptors remain somewhat puzzling. Here we examine discoveries linking these receptors to physiology and their probable implications in disease, in particular mood disorders, and propose some ideas to obtain a deeper understanding of these intriguing proteins. A Historical Overview The absence of specific antibodies against different KAR Most excitatory synapses in the brain use the amino acid gluta- subunits has been a significant limitation in terms of exploring re- mate as a neurotransmitter. Since the excitatory properties of ceptor distribution. Thus, most of the information regarding their glutamate were postulated nearly 40 years ago, an extraordinary tissue expression comes from in situ hybridization studies that, wealth of data has accumulated on the types of synaptic re- although informative, cannot reveal the subcellular distribution sponses triggered by this neurotransmitter. -

Coupling of Autism Genes to Tissue-Wide Expression and Dysfunction of Synapse, Calcium Signalling and Transcriptional Regulation
PLOS ONE RESEARCH ARTICLE Coupling of autism genes to tissue-wide expression and dysfunction of synapse, calcium signalling and transcriptional regulation 1 2,3 4 1,5 Jamie ReillyID *, Louise Gallagher , Geraldine Leader , Sanbing Shen * 1 Regenerative Medicine Institute, School of Medicine, Biomedical Science Building, National University of a1111111111 Ireland (NUI) Galway, Galway, Ireland, 2 Discipline of Psychiatry, School of Medicine, Trinity College Dublin, Dublin, Ireland, 3 Trinity Translational Medicine Institute, Trinity Centre for Health SciencesÐTrinity College a1111111111 Dublin, St. James's Hospital, Dublin, Ireland, 4 Irish Centre for Autism and Neurodevelopmental Research a1111111111 (ICAN), Department of Psychology, National University of Ireland (NUI) Galway, Galway, Ireland, a1111111111 5 FutureNeuro Research Centre, Royal College of Surgeons in Ireland (RCSI), Dublin, Ireland a1111111111 * [email protected] (JR); [email protected] (SS) Abstract OPEN ACCESS Citation: Reilly J, Gallagher L, Leader G, Shen S Autism Spectrum Disorder (ASD) is a heterogeneous disorder that is often accompanied (2020) Coupling of autism genes to tissue-wide with many co-morbidities. Recent genetic studies have identified various pathways from expression and dysfunction of synapse, calcium hundreds of candidate risk genes with varying levels of association to ASD. However, it is signalling and transcriptional regulation. PLoS ONE unknown which pathways are specific to the core symptoms or which are shared by the co- 15(12): e0242773. https://doi.org/10.1371/journal. pone.0242773 morbidities. We hypothesised that critical ASD candidates should appear widely across dif- ferent scoring systems, and that comorbidity pathways should be constituted by genes Editor: Nirakar Sahoo, The University of Texas Rio Grande Valley, UNITED STATES expressed in the relevant tissues. -

Mechanisms Underlying the EEG Biomarker in Dup15q Syndrome Joel Frohlich1,2,3* , Lawrence T
Frohlich et al. Molecular Autism (2019) 10:29 https://doi.org/10.1186/s13229-019-0280-6 RESEARCH Open Access Mechanisms underlying the EEG biomarker in Dup15q syndrome Joel Frohlich1,2,3* , Lawrence T. Reiter4, Vidya Saravanapandian2, Charlotte DiStefano2, Scott Huberty2,5, Carly Hyde2, Stormy Chamberlain6, Carrie E. Bearden7, Peyman Golshani8, Andrei Irimia9, Richard W. Olsen10, Joerg F. Hipp1† and Shafali S. Jeste2† Abstract Background: Duplications of 15q11.2-q13.1 (Dup15q syndrome), including the paternally imprinted gene UBE3A and three nonimprinted gamma-aminobutyric acid type-A (GABAA) receptor genes, are highly penetrant for neurodevelopmental disorders such as autism spectrum disorder (ASD). To guide targeted treatments of Dup15q syndrome and other forms of ASD, biomarkers are needed that reflect molecular mechanisms of pathology. We recently described a beta EEG phenotype of Dup15q syndrome, but it remains unknown which specific genes drive this phenotype. Methods: To test the hypothesis that UBE3A overexpression is not necessary for the beta EEG phenotype, we compared EEG from a reference cohort of children with Dup15q syndrome (n = 27) to (1) the pharmacological effects of the GABAA modulator midazolam (n = 12) on EEG from healthy adults, (2) EEG from typically developing (TD) children (n = 14), and (3) EEG from two children with duplications of paternal 15q (i.e., the UBE3A-silenced allele). Results: Peak beta power was significantly increased in the reference cohort relative to TD controls. Midazolam administration recapitulated the beta EEG phenotype in healthy adults with a similar peak frequency in central channels (f = 23.0 Hz) as Dup15q syndrome (f = 23.1 Hz). -

Characterization of Zebrafish GABAA Receptor Subunits
www.nature.com/scientificreports OPEN Characterization of zebrafsh GABAA receptor subunits Kenichiro Sadamitsu, Leona Shigemitsu, Marina Suzuki, Daishi Ito, Makoto Kashima & Hiromi Hirata* γ-Aminobutyric acid (GABA), the major inhibitory neurotransmitter in the central nervous system, exerts its efect through the activation of GABA receptors. GABAA receptors are ligand-gated chloride channels composed of fve subunit proteins. Mammals have 19 diferent GABAA receptor subunits (α1–6, β1–3, γ1–3, δ, ε, π, θ, and ρ1–3), the physiological properties of which have been assayed by electrophysiology. However, the evolutionary conservation of the physiological characteristics of diverged GABAA receptor subunits remains unclear. Zebrafsh have 23 subunits (α1, α2a, α2b, α3–5, α6a, α6b, β1–4, γ1–3, δ, π, ζ, ρ1, ρ2a, ρ2b, ρ3a, and ρ3b), but the electrophysiological properties of these subunits have not been explored. In this study, we cloned the coding sequences for zebrafsh GABAA receptor subunits and investigated their expression patterns in larval zebrafsh by whole- mount in situ hybridization. We also performed electrophysiological recordings of GABA-evoked currents from Xenopus oocytes injected with one or multiple zebrafsh GABAA receptor subunit cRNAs and calculated the half-maximal efective concentrations (EC50s) for each. Our results revealed the spatial expressions and electrophysiological GABA sensitivities of zebrafsh GABAA receptors, suggesting that the properties of GABAA receptor subunits are conserved among vertebrates. γ-Aminobutyric acid (GABA), the major inhibitory neurotransmitter in the central nervous system of vertebrates, 1 controls the excitability of neural networks mainly through GABA A receptors . Te GABAA receptor mediates two types of inhibition, known as phasic and tonic inhibition2. -

Sex Differences in Glutamate Receptor Gene Expression in Major Depression and Suicide
Molecular Psychiatry (2015) 20, 1057–1068 © 2015 Macmillan Publishers Limited All rights reserved 1359-4184/15 www.nature.com/mp IMMEDIATE COMMUNICATION Sex differences in glutamate receptor gene expression in major depression and suicide AL Gray1, TM Hyde2,3, A Deep-Soboslay2, JE Kleinman2 and MS Sodhi1,4 Accumulating data indicate that the glutamate system is disrupted in major depressive disorder (MDD), and recent clinical research suggests that ketamine, an antagonist of the N-methyl-D-aspartate (NMDA) glutamate receptor (GluR), has rapid antidepressant efficacy. Here we report findings from gene expression studies of a large cohort of postmortem subjects, including subjects with MDD and controls. Our data reveal higher expression levels of the majority of glutamatergic genes tested in the dorsolateral prefrontal cortex (DLPFC) in MDD (F21,59 = 2.32, P = 0.006). Posthoc data indicate that these gene expression differences occurred mostly in the female subjects. Higher expression levels of GRIN1, GRIN2A-D, GRIA2-4, GRIK1-2, GRM1, GRM4, GRM5 and GRM7 were detected in the female patients with MDD. In contrast, GRM5 expression was lower in male MDD patients relative to male controls. When MDD suicides were compared with MDD non-suicides, GRIN2B, GRIK3 and GRM2 were expressed at higher levels in the suicides. Higher expression levels were detected for several additional genes, but these were not statistically significant after correction for multiple comparisons. In summary, our analyses indicate a generalized disruption of the regulation of the GluRs in the DLPFC of females with MDD, with more specific GluR alterations in the suicides and in the male groups. -

Genetics of Migraine: Insights Into the Molecular Basis of Migraine Disorders
This may be the author’s version of a work that was submitted/accepted for publication in the following source: Sutherland, Heidi& Griffiths, Lyn (2017) Genetics of migraine: Insights into the molecular basis of migraine disor- ders. Headache, 57(4), pp. 537-569. This file was downloaded from: https://eprints.qut.edu.au/105633/ c Consult author(s) regarding copyright matters This work is covered by copyright. Unless the document is being made available under a Creative Commons Licence, you must assume that re-use is limited to personal use and that permission from the copyright owner must be obtained for all other uses. If the docu- ment is available under a Creative Commons License (or other specified license) then refer to the Licence for details of permitted re-use. It is a condition of access that users recog- nise and abide by the legal requirements associated with these rights. If you believe that this work infringes copyright please provide details by email to [email protected] Notice: Please note that this document may not be the Version of Record (i.e. published version) of the work. Author manuscript versions (as Sub- mitted for peer review or as Accepted for publication after peer review) can be identified by an absence of publisher branding and/or typeset appear- ance. If there is any doubt, please refer to the published source. https://doi.org/10.1111/head.13053 Genetics of Migraine: insights into the molecular basis of migraine disorders Heidi G. Sutherland, PhD and Lyn R. Griffiths, PhD Genomics Research Centre, Institute of Health and Biomedical Innovation, QUT, Musk Ave, Kelvin Grove, QLD 4059, Australia The authors declare no conflicts of interest. -
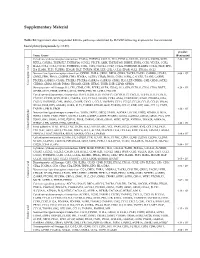
Supplementary Material
Supplementary Material Table S1: Significant downregulated KEGGs pathways identified by DAVID following exposure to five cinnamon- based phenylpropanoids (p < 0.05). p-value Term: Genes (Benjamini) Cytokine-cytokine receptor interaction: FASLG, TNFSF14, CXCL11, IL11, FLT3LG, CCL3L1, CCL3L3, CXCR6, XCR1, 2.43 × 105 RTEL1, CSF2RA, TNFRSF17, TNFRSF14, CCNL2, VEGFB, AMH, TNFRSF10B, INHBE, IFNB1, CCR3, VEGFA, CCR2, IL12A, CCL1, CCL3, CXCL5, TNFRSF25, CCR1, CSF1, CX3CL1, CCL7, CCL24, TNFRSF1B, IL12RB1, CCL21, FIGF, EPO, IL4, IL18R1, FLT1, TGFBR1, EDA2R, HGF, TNFSF8, KDR, LEP, GH2, CCL13, EPOR, XCL1, IFNA16, XCL2 Neuroactive ligand-receptor interaction: OPRM1, THRA, GRIK1, DRD2, GRIK2, TACR2, TACR1, GABRB1, LPAR4, 9.68 × 105 GRIK5, FPR1, PRSS1, GNRHR, FPR2, EDNRA, AGTR2, LTB4R, PRSS2, CNR1, S1PR4, CALCRL, TAAR5, GABRE, PTGER1, GABRG3, C5AR1, PTGER3, PTGER4, GABRA6, GABRA5, GRM1, PLG, LEP, CRHR1, GH2, GRM3, SSTR2, Chlorogenic acid Chlorogenic CHRM3, GRIA1, MC2R, P2RX2, TBXA2R, GHSR, HTR2C, TSHR, LHB, GLP1R, OPRD1 Hematopoietic cell lineage: IL4, CR1, CD8B, CSF1, FCER2, GYPA, ITGA2, IL11, GP9, FLT3LG, CD38, CD19, DNTT, 9.29 × 104 GP1BB, CD22, EPOR, CSF2RA, CD14, THPO, EPO, HLA-DRA, ITGA2B Cytokine-cytokine receptor interaction: IL6ST, IL21R, IL19, TNFSF15, CXCR3, IL15, CXCL11, TGFB1, IL11, FLT3LG, CXCL10, CCR10, XCR1, RTEL1, CSF2RA, IL21, CCNL2, VEGFB, CCR8, AMH, TNFRSF10C, IFNB1, PDGFRA, EDA, CXCL5, TNFRSF25, CSF1, IFNW1, CNTFR, CX3CL1, CCL5, TNFRSF4, CCL4, CCL27, CCL24, CCL25, CCL23, IFNA6, IFNA5, FIGF, EPO, AMHR2, IL2RA, FLT4, TGFBR2, EDA2R, -

Research Article Microarray-Based Comparisons of Ion Channel Expression Patterns: Human Keratinocytes to Reprogrammed Hipscs To
Hindawi Publishing Corporation Stem Cells International Volume 2013, Article ID 784629, 25 pages http://dx.doi.org/10.1155/2013/784629 Research Article Microarray-Based Comparisons of Ion Channel Expression Patterns: Human Keratinocytes to Reprogrammed hiPSCs to Differentiated Neuronal and Cardiac Progeny Leonhard Linta,1 Marianne Stockmann,1 Qiong Lin,2 André Lechel,3 Christian Proepper,1 Tobias M. Boeckers,1 Alexander Kleger,3 and Stefan Liebau1 1 InstituteforAnatomyCellBiology,UlmUniversity,Albert-EinsteinAllee11,89081Ulm,Germany 2 Institute for Biomedical Engineering, Department of Cell Biology, RWTH Aachen, Pauwelstrasse 30, 52074 Aachen, Germany 3 Department of Internal Medicine I, Ulm University, Albert-Einstein Allee 11, 89081 Ulm, Germany Correspondence should be addressed to Alexander Kleger; [email protected] and Stefan Liebau; [email protected] Received 31 January 2013; Accepted 6 March 2013 Academic Editor: Michael Levin Copyright © 2013 Leonhard Linta et al. This is an open access article distributed under the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited. Ion channels are involved in a large variety of cellular processes including stem cell differentiation. Numerous families of ion channels are present in the organism which can be distinguished by means of, for example, ion selectivity, gating mechanism, composition, or cell biological function. To characterize the distinct expression of this group of ion channels we have compared the mRNA expression levels of ion channel genes between human keratinocyte-derived induced pluripotent stem cells (hiPSCs) and their somatic cell source, keratinocytes from plucked human hair. This comparison revealed that 26% of the analyzed probes showed an upregulation of ion channels in hiPSCs while just 6% were downregulated.