Cargo Sorting Zones in the Trans-Golgi Network Visualized by Super-Resolution Confocal Live Imaging Microscopy in Plants
Total Page:16
File Type:pdf, Size:1020Kb

Load more
Recommended publications
-

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

Protein Identities in Evs Isolated from U87-MG GBM Cells As Determined by NG LC-MS/MS
Protein identities in EVs isolated from U87-MG GBM cells as determined by NG LC-MS/MS. No. Accession Description Σ Coverage Σ# Proteins Σ# Unique Peptides Σ# Peptides Σ# PSMs # AAs MW [kDa] calc. pI 1 A8MS94 Putative golgin subfamily A member 2-like protein 5 OS=Homo sapiens PE=5 SV=2 - [GG2L5_HUMAN] 100 1 1 7 88 110 12,03704523 5,681152344 2 P60660 Myosin light polypeptide 6 OS=Homo sapiens GN=MYL6 PE=1 SV=2 - [MYL6_HUMAN] 100 3 5 17 173 151 16,91913397 4,652832031 3 Q6ZYL4 General transcription factor IIH subunit 5 OS=Homo sapiens GN=GTF2H5 PE=1 SV=1 - [TF2H5_HUMAN] 98,59 1 1 4 13 71 8,048185945 4,652832031 4 P60709 Actin, cytoplasmic 1 OS=Homo sapiens GN=ACTB PE=1 SV=1 - [ACTB_HUMAN] 97,6 5 5 35 917 375 41,70973209 5,478027344 5 P13489 Ribonuclease inhibitor OS=Homo sapiens GN=RNH1 PE=1 SV=2 - [RINI_HUMAN] 96,75 1 12 37 173 461 49,94108966 4,817871094 6 P09382 Galectin-1 OS=Homo sapiens GN=LGALS1 PE=1 SV=2 - [LEG1_HUMAN] 96,3 1 7 14 283 135 14,70620005 5,503417969 7 P60174 Triosephosphate isomerase OS=Homo sapiens GN=TPI1 PE=1 SV=3 - [TPIS_HUMAN] 95,1 3 16 25 375 286 30,77169764 5,922363281 8 P04406 Glyceraldehyde-3-phosphate dehydrogenase OS=Homo sapiens GN=GAPDH PE=1 SV=3 - [G3P_HUMAN] 94,63 2 13 31 509 335 36,03039959 8,455566406 9 Q15185 Prostaglandin E synthase 3 OS=Homo sapiens GN=PTGES3 PE=1 SV=1 - [TEBP_HUMAN] 93,13 1 5 12 74 160 18,68541938 4,538574219 10 P09417 Dihydropteridine reductase OS=Homo sapiens GN=QDPR PE=1 SV=2 - [DHPR_HUMAN] 93,03 1 1 17 69 244 25,77302971 7,371582031 11 P01911 HLA class II histocompatibility antigen, -

Supplementary Table S1. Correlation Between the Mutant P53-Interacting Partners and PTTG3P, PTTG1 and PTTG2, Based on Data from Starbase V3.0 Database
Supplementary Table S1. Correlation between the mutant p53-interacting partners and PTTG3P, PTTG1 and PTTG2, based on data from StarBase v3.0 database. PTTG3P PTTG1 PTTG2 Gene ID Coefficient-R p-value Coefficient-R p-value Coefficient-R p-value NF-YA ENSG00000001167 −0.077 8.59e-2 −0.210 2.09e-6 −0.122 6.23e-3 NF-YB ENSG00000120837 0.176 7.12e-5 0.227 2.82e-7 0.094 3.59e-2 NF-YC ENSG00000066136 0.124 5.45e-3 0.124 5.40e-3 0.051 2.51e-1 Sp1 ENSG00000185591 −0.014 7.50e-1 −0.201 5.82e-6 −0.072 1.07e-1 Ets-1 ENSG00000134954 −0.096 3.14e-2 −0.257 4.83e-9 0.034 4.46e-1 VDR ENSG00000111424 −0.091 4.10e-2 −0.216 1.03e-6 0.014 7.48e-1 SREBP-2 ENSG00000198911 −0.064 1.53e-1 −0.147 9.27e-4 −0.073 1.01e-1 TopBP1 ENSG00000163781 0.067 1.36e-1 0.051 2.57e-1 −0.020 6.57e-1 Pin1 ENSG00000127445 0.250 1.40e-8 0.571 9.56e-45 0.187 2.52e-5 MRE11 ENSG00000020922 0.063 1.56e-1 −0.007 8.81e-1 −0.024 5.93e-1 PML ENSG00000140464 0.072 1.05e-1 0.217 9.36e-7 0.166 1.85e-4 p63 ENSG00000073282 −0.120 7.04e-3 −0.283 1.08e-10 −0.198 7.71e-6 p73 ENSG00000078900 0.104 2.03e-2 0.258 4.67e-9 0.097 3.02e-2 Supplementary Table S2. -

Supplementary Table S4. FGA Co-Expressed Gene List in LUAD
Supplementary Table S4. FGA co-expressed gene list in LUAD tumors Symbol R Locus Description FGG 0.919 4q28 fibrinogen gamma chain FGL1 0.635 8p22 fibrinogen-like 1 SLC7A2 0.536 8p22 solute carrier family 7 (cationic amino acid transporter, y+ system), member 2 DUSP4 0.521 8p12-p11 dual specificity phosphatase 4 HAL 0.51 12q22-q24.1histidine ammonia-lyase PDE4D 0.499 5q12 phosphodiesterase 4D, cAMP-specific FURIN 0.497 15q26.1 furin (paired basic amino acid cleaving enzyme) CPS1 0.49 2q35 carbamoyl-phosphate synthase 1, mitochondrial TESC 0.478 12q24.22 tescalcin INHA 0.465 2q35 inhibin, alpha S100P 0.461 4p16 S100 calcium binding protein P VPS37A 0.447 8p22 vacuolar protein sorting 37 homolog A (S. cerevisiae) SLC16A14 0.447 2q36.3 solute carrier family 16, member 14 PPARGC1A 0.443 4p15.1 peroxisome proliferator-activated receptor gamma, coactivator 1 alpha SIK1 0.435 21q22.3 salt-inducible kinase 1 IRS2 0.434 13q34 insulin receptor substrate 2 RND1 0.433 12q12 Rho family GTPase 1 HGD 0.433 3q13.33 homogentisate 1,2-dioxygenase PTP4A1 0.432 6q12 protein tyrosine phosphatase type IVA, member 1 C8orf4 0.428 8p11.2 chromosome 8 open reading frame 4 DDC 0.427 7p12.2 dopa decarboxylase (aromatic L-amino acid decarboxylase) TACC2 0.427 10q26 transforming, acidic coiled-coil containing protein 2 MUC13 0.422 3q21.2 mucin 13, cell surface associated C5 0.412 9q33-q34 complement component 5 NR4A2 0.412 2q22-q23 nuclear receptor subfamily 4, group A, member 2 EYS 0.411 6q12 eyes shut homolog (Drosophila) GPX2 0.406 14q24.1 glutathione peroxidase -

Gamma Adaptin (AP1G1) Rabbit Polyclonal Antibody Product Data
OriGene Technologies, Inc. 9620 Medical Center Drive, Ste 200 Rockville, MD 20850, US Phone: +1-888-267-4436 [email protected] EU: [email protected] CN: [email protected] Product datasheet for TA323388 gamma Adaptin (AP1G1) Rabbit Polyclonal Antibody Product data: Product Type: Primary Antibodies Applications: IHC Recommended Dilution: ELISA: 1:1000-5000, IHC: 1:50-200 Reactivity: Human, Mouse, Rat Host: Rabbit Isotype: IgG Clonality: Polyclonal Immunogen: Synthetic peptide corresponding to a region derived from 802-814 amino acids of Human Adapter-related protein complex 1 subunit gamma-1 Formulation: PBS pH7.3, 0.05% NaN3, 50% glycerol Concentration: lot specific Purification: Antigen affinity purification Conjugation: Unconjugated Storage: Store at -20°C as received. Stability: Stable for 12 months from date of receipt. Gene Name: adaptor related protein complex 1 gamma 1 subunit Database Link: NP_001119 Entrez Gene 11765 MouseEntrez Gene 171494 RatEntrez Gene 164 Human O43747 Background: Adaptins are important components of clathrin-coated vesicles transporting ligand-receptor complexes from the plasma membrane or from the trans-Golgi network to lysosomes. The adaptin family of proteins is composed of four classes of molecules named alpha; beta-; beta prime- and gamma- adaptins. Adaptins; together with medium and small subunits; form a heterotetrameric complex called an adaptor; whose role is to promote the formation of clathrin-coated pits and vesicles. The protein encoded by this gene is a gamma-adaptin protein and it belongs to the adaptor complexes large subunits family. Two transcript variants encoding different isoforms have been found for this gene. Synonyms: ADTG; CLAPG1 This product is to be used for laboratory only. -

Entrez ID 1 Symbol 1 Entrez ID 2 Symbol 2 Data Source (R
Supporting Information Table 4. List of human protein-protein interactons. Entrez ID 1 Symbol 1 Entrez ID 2 Symbol 2 Data Source (R: Rual et al; S: Stelzl et al; L: Literature curation) 1 A1BG 10321 CRISP3 L 2 A2M 259 AMBP L 2 A2M 348 APOE L 2 A2M 351 APP L 2 A2M 354 KLK3 L 2 A2M 567 B2M L 2 A2M 1508 CTSB L 2 A2M 1990 ELA1 L 2 A2M 3309 HSPA5 L 2 A2M 3553 IL1B L 2 A2M 3586 IL10 L 2 A2M 3931 LCAT L 2 A2M 3952 LEP L 2 A2M 4035 LRP1 L 2 A2M 4803 NGFB L 2 A2M 5047 PAEP L 2 A2M 7045 TGFBI L 2 A2M 8728 ADAM19 L 2 A2M 9510 ADAMTS1 L 2 A2M 10944 SMAP S 2 A2M 55729 ATF7IP L 9 NAT1 8260 ARD1A L 12 SERPINA3 351 APP L 12 SERPINA3 354 KLK3 L 12 SERPINA3 1215 CMA1 L 12 SERPINA3 1504 CTRB1 L 12 SERPINA3 1506 CTRL L 12 SERPINA3 1511 CTSG L 12 SERPINA3 1990 ELA1 L 12 SERPINA3 1991 ELA2 L 12 SERPINA3 2064 ERBB2 L 12 SERPINA3 2153 F5 L 12 SERPINA3 3817 KLK2 L 12 SERPINA3 4035 LRP1 L 12 SERPINA3 4485 MST1 L 12 SERPINA3 5422 POLA L 12 SERPINA3 64215 DNAJC1 L 14 AAMP 51497 TH1L S 15 AANAT 7534 YWHAZ L 18 ABAT 7915 ALDH5A1 L 19 ABCA1 335 APOA1 L 19 ABCA1 6645 SNTB2 L 19 ABCA1 8772 FADD L 20 ABCA2 55755 CDK5RAP2 L 22 ABCB7 2235 FECH L 23 ABCF1 3692 ITGB4BP S 24 ABCA4 1258 CNGB1 L 25 ABL1 27 ABL2 L 25 ABL1 472 ATM L 25 ABL1 613 BCR L 25 ABL1 718 C3 L 25 ABL1 867 CBL L 25 ABL1 1501 CTNND2 L 25 ABL1 2048 EPHB2 L 25 ABL1 2547 XRCC6 L 25 ABL1 2876 GPX1 L 25 ABL1 2885 GRB2 L 25 ABL1 3055 HCK L 25 ABL1 3636 INPPL1 L 25 ABL1 3716 JAK1 L 25 ABL1 4193 MDM2 L 25 ABL1 4690 NCK1 L 25 ABL1 4914 NTRK1 L 25 ABL1 5062 PAK2 L 25 ABL1 5295 PIK3R1 L 25 ABL1 5335 PLCG1 L 25 ABL1 5591 -

Supplemental Digital Content 2 List of Down-Regulated Transcripts In
Supplemental Digital Content 2 List of down-regulated transcripts in colonic mucosal UC-associated dysplasia and pancolitis compared to left-sided colitis Probe Set ID Gene Title Gene Symbol 225655_at ubiquitin-like, containing PHD and RING finger domains, 1 UHRF1 216640_s_at protein disulfide isomerase family A, member 6 PDIA6 1555358_a_at ectonucleoside triphosphate diphosphohydrolase 4 ENTPD4 201340_s_at ectodermal-neural cortex (with BTB-like domain) ENC1 203843_at ribosomal protein S6 kinase, 90kDa, polypeptide 3 RPS6KA3 206668_s_at secretory carrier membrane protein 1 SCAMP1 207791_s_at RAB1A, member RAS oncogene family RAB1A 228996_at roquin KIAA2025 Transcribed locus, weakly similar to XP_517454.1 PREDICTED: 228623_at similar to hypothetical protein MGC45438 [Pan troglodytes] --- 202817_s_at synovial sarcoma translocation, chromosome 18 SS18 235234_at hypothetical protein FLJ36874 FLJ36874 201514_s_at Ras-GTPase-activating protein SH3-domain-binding protein G3BP 218238_at GTP binding protein 4 GTPBP4 241343_at Ribonuclease H1 RNASEH1 1553033_at synaptotagmin-like 5 SYTL5 203210_s_at replication factor C (activator 1) 5, 36.5kDa RFC5 213875_x_at chromosome 6 open reading frame 62 C6orf62 1553108_at hypothetical protein FLJ37562 FLJ37562 215820_x_at sorting nexin 13 SNX13 230130_at Slit homolog 2 (Drosophila) SLIT2 200607_s_at RAD21 homolog (S. pombe) RAD21 235296_at eukaryotic translation initiation factor 5A2 EIF5A2 1552656_s_at U2AF homology motif (UHM) kinase 1 UHMK1 214155_s_at La ribonucleoprotein domain family, member 4 LARP4 -

Identification of Podocyte Cargo Proteins by Proteomic Analysis of Clathrin-Coated
Kidney360 Publish Ahead of Print, published on April 16, 2020 as doi:10.34067/KID.0000212020 Identification of Podocyte Cargo Proteins by Proteomic Analysis of Clathrin-coated Vesicles Marwin Groener1,2,3, Ying Wang1, Elizabeth Cross1, Xuefei Tian1, Karen Ebenezer1, Eunice Baik1, Christopher Pedigo1, Mario Schiffer2,3,4, Kazunori Inoue1, and Shuta Ishibe1 1. Department of Internal Medicine, Yale University School of Medicine, New Haven, Connecticut, USA 2. Department of Nephrology and Hypertension, University Hospital Erlangen, Erlangen, Germany 3. Friedrich-Alexander-Universität Erlangen-Nürnberg (FAU), Erlangen, Germany 4. Mount Desert Island Biological Laboratories, Salisbury Cove, Maine, USA Corresponding Author: Dr. Shuta Ishibe, Section of Nephrology, Yale University School of Medicine, PO Box 208029, 333 Cedar Street, New Haven, CT 06520-8029. Email: [email protected]. Copyright 2020 by American Society of Nephrology. Abstract Background: Clathrin-mediated endocytosis (CME) plays a fundamental role in podocyte health. Genetic ablation of genes implicated in CME have been shown to cause severe proteinuria and foot process effacement in mice. However, little is known about the cargo of clathrin-coated vesicles (CCVs) in podocytes. The goal of this study was to isolate CCVs from podocytes and identify their cargo by proteomic analysis. Methods: Glomeruli isolated from Podocin-Cre Rosa-DTRflox mouse kidneys were seeded and treated with diphtheria toxin to obtain pure primary podocyte cultures. CCVs were isolated by differential gradient ultracentrifugation and enrichment of CCVs was assessed by immunoblotting and electron microscopy (EM). Liquid chromatography-mass spectrometry (LC-MS) was performed for proteomic analysis. Proteins with higher abundance than transferrin receptor protein 1 were evaluated for CCV cargo potential against previously published literature. -
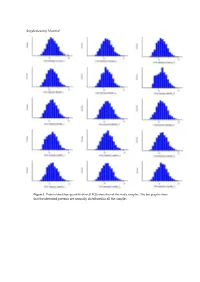
Supplementary Material
Supplementary Material Figure 1. Protein label-free-quantification (LFQ) intensities of the study samples. The bar graphs show that the identified proteins are normally distributed in all the samples. Figure 2. IPA network analysis of the modified proteins in response to BK 24h stimulation relative to control in RASMC. Involvement of TNF, Jnk, and SOD2 downstream of BK stimulation. Figure 3. IPA network analysis of the modified proteins in response to BK 48h stimulation relative to control in RASMC. Activation of SERPINE1 downstream of BK stimulation. Figure 4. Immunocytochemistry staining assessing the expression of LeptR (FITC, Green) in response to BK time point stimulations in RASMC. Figure 5. IPA network analysis of the modified proteins in response to Leptin 24h stimulation relative to control in RASMC. Involvement of IL1β and TNF downstream of Leptin stimulation. Figure 6. IPA network analysis of the modified proteins in response to Leptin 48h stimulation relative to control in RASMC. Involvement of NFκB, TNF, Jn, and SOD2 downstream of Leptin stimulation. Table 1. Comparative list of proteins in the BK-stimulated for 24h compared to controls. Fold Number Accession Name q-value change 1 B2RYL3 Tmem119 - Transmembrane protein 119 precursor 0.002 6.18 2 A0A140TAI9 Leprotl1 - Leptin receptor overlapping transcript-like 1 0.006 1.83 3 Q9JLS8 Leprot - Leptin receptor gene-related protein 0.040 1.82 4 D4A5L9 LOC690675 - Electron carrier protein 0.033 1.66 5 Q5BJU0 Rras2 - Ras-related protein R-Ras2 0.044 1.62 6 B2RYP6 Luc7l2 - LUC7-like -

A Network-Based, Integrative Study to Identify Core Biological Pathways That Drive Breast Cancer Clinical Subtypes
British Journal of Cancer (2012) 106, 1107 – 1116 & 2012 Cancer Research UK All rights reserved 0007 – 0920/12 www.bjcancer.com A network-based, integrative study to identify core biological pathways that drive breast cancer clinical subtypes 1 *,2 3 4 4 2 2 5 2 2 B Dutta , L Pusztai ,YQi, F Andre´ , V Lazar , G Bianchini , N Ueno , R Agarwal , B Wang , CY Shiang , 2 1 6 *,1 GN Hortobagyi , GB Mills , WF Symmans and G Bala´zsi 1Department of Systems Biology – Unit 950, The University of Texas MD Anderson Cancer Center, 7435 Fannin Street, Houston, TX 77054, USA; 2 Department of Breast Medical Oncology – Unit 1354, The University of Texas MD Anderson Cancer Center, PO Box 301439, Houston, 3 4 TX 77230-1439, USA; Department of Bioinformatics, The University of Texas MD Anderson Cancer Center, Houston, TX 77054, USA; Institute Gustave 5 6 Roussy, Villejuif, France; Department of Medical Oncology, Imperial College, London, UK; Department of Pathology, The University of Texas MD Anderson Cancer Center, Houston, TX 77054, USA BACKGROUND: The rapid collection of diverse genome-scale data raises the urgent need to integrate and utilise these resources for biological discovery or biomedical applications. For example, diverse transcriptomic and gene copy number variation data are currently collected for various cancers, but relatively few current methods are capable to utilise the emerging information. METHODS: We developed and tested a data-integration method to identify gene networks that drive the biology of breast cancer clinical subtypes. The method simultaneously overlays gene expression and gene copy number data on protein–protein interaction, transcriptional-regulatory and signalling networks by identifying coincident genomic and transcriptional disturbances in local network neighborhoods. -

The Endosome Is a Master Regulator of Plasma Membrane Collagen Fibril Assembly
bioRxiv preprint doi: https://doi.org/10.1101/2021.03.25.436925; this version posted March 25, 2021. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. The endosome is a master regulator of plasma membrane collagen fibril assembly 1Joan Chang*, 1Adam Pickard, 1Richa Garva, 1Yinhui Lu, 2Donald Gullberg and 1Karl E. Kadler* 1Wellcome Centre for Cell-Matrix Research, Faculty of Biology, Medical and Health, University of Manchester, Michael Smith Building, Oxford Road, Manchester M13 9PT UK, 2Department of Biomedicine and Center for Cancer Biomarkers, Norwegian Center of Excellence, University of Bergen, Norway. * Co-corresponding authors: JC email: [email protected] (orcid.org/0000-0002-7283- 9759); KEK email: [email protected] (orcid.org/0000-0003-4977-4683) Keywords: collagen-I, endocytosis, extracellular matrix, fibril, fibrillogenesis, integrin-a11, trafficking, VPS33b, [abstract] [149 word max] Collagen fibrils are the principal supporting elements in vertebrate tissues. They account for 25% of total protein mass, exhibit a broad range of size and organisation depending on tissue and stage of development, and can be under circadian clock control. Here we show that the remarkable dynamic pleomorphism of collagen fibrils is underpinned by a mechanism that distinguishes between collagen secretion and initiation of fibril assembly, at the plasma membrane. Collagen fibrillogenesis occurring at the plasma membrane requires vacuolar protein sorting (VPS) 33b (which is under circadian clock control), collagen-binding integrin-a11 subunit, and is reduced when endocytosis is inhibited. Fibroblasts lacking VPS33b secrete soluble collagen without assembling fibrils, whereas constitutive over-expression of VPS33b increases fibril number with loss of fibril rhythmicity. -

Autocrine IFN Signaling Inducing Profibrotic Fibroblast Responses by a Synthetic TLR3 Ligand Mitigates
Downloaded from http://www.jimmunol.org/ by guest on September 28, 2021 Inducing is online at: average * The Journal of Immunology published online 16 August 2013 from submission to initial decision 4 weeks from acceptance to publication http://www.jimmunol.org/content/early/2013/08/16/jimmun ol.1300376 A Synthetic TLR3 Ligand Mitigates Profibrotic Fibroblast Responses by Autocrine IFN Signaling Feng Fang, Kohtaro Ooka, Xiaoyong Sun, Ruchi Shah, Swati Bhattacharyya, Jun Wei and John Varga J Immunol Submit online. Every submission reviewed by practicing scientists ? is published twice each month by http://jimmunol.org/subscription Submit copyright permission requests at: http://www.aai.org/About/Publications/JI/copyright.html Receive free email-alerts when new articles cite this article. Sign up at: http://jimmunol.org/alerts http://www.jimmunol.org/content/suppl/2013/08/20/jimmunol.130037 6.DC1 Information about subscribing to The JI No Triage! Fast Publication! Rapid Reviews! 30 days* Why • • • Material Permissions Email Alerts Subscription Supplementary The Journal of Immunology The American Association of Immunologists, Inc., 1451 Rockville Pike, Suite 650, Rockville, MD 20852 Copyright © 2013 by The American Association of Immunologists, Inc. All rights reserved. Print ISSN: 0022-1767 Online ISSN: 1550-6606. This information is current as of September 28, 2021. Published August 16, 2013, doi:10.4049/jimmunol.1300376 The Journal of Immunology A Synthetic TLR3 Ligand Mitigates Profibrotic Fibroblast Responses by Inducing Autocrine IFN Signaling Feng Fang,* Kohtaro Ooka,* Xiaoyong Sun,† Ruchi Shah,* Swati Bhattacharyya,* Jun Wei,* and John Varga* Activation of TLR3 by exogenous microbial ligands or endogenous injury-associated ligands leads to production of type I IFN.