A Systematic Review of Molecular Imaging Agents Targeting Bradykinin B1 and B2 Receptors
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Molecular Dissection of G-Protein Coupled Receptor Signaling and Oligomerization
MOLECULAR DISSECTION OF G-PROTEIN COUPLED RECEPTOR SIGNALING AND OLIGOMERIZATION BY MICHAEL RIZZO A Dissertation Submitted to the Graduate Faculty of WAKE FOREST UNIVERSITY GRADUATE SCHOOL OF ARTS AND SCIENCES in Partial Fulfillment of the Requirements for the Degree of DOCTOR OF PHILOSOPHY Biology December, 2019 Winston-Salem, North Carolina Approved By: Erik C. Johnson, Ph.D. Advisor Wayne E. Pratt, Ph.D. Chair Pat C. Lord, Ph.D. Gloria K. Muday, Ph.D. Ke Zhang, Ph.D. ACKNOWLEDGEMENTS I would first like to thank my advisor, Dr. Erik Johnson, for his support, expertise, and leadership during my time in his lab. Without him, the work herein would not be possible. I would also like to thank the members of my committee, Dr. Gloria Muday, Dr. Ke Zhang, Dr. Wayne Pratt, and Dr. Pat Lord, for their guidance and advice that helped improve the quality of the research presented here. I would also like to thank members of the Johnson lab, both past and present, for being valuable colleagues and friends. I would especially like to thank Dr. Jason Braco, Dr. Jon Fisher, Dr. Jake Saunders, and Becky Perry, all of whom spent a great deal of time offering me advice, proofreading grants and manuscripts, and overall supporting me through the ups and downs of the research process. Finally, I would like to thank my family, both for instilling in me a passion for knowledge and education, and for their continued support. In particular, I would like to thank my wife Emerald – I am forever indebted to you for your support throughout this process, and I will never forget the sacrifices you made to help me get to where I am today. -

Supplementary Table 3 Complete List of RNA-Sequencing Analysis of Gene Expression Changed by ≥ Tenfold Between Xenograft and Cells Cultured in 10%O2
Supplementary Table 3 Complete list of RNA-Sequencing analysis of gene expression changed by ≥ tenfold between xenograft and cells cultured in 10%O2 Expr Log2 Ratio Symbol Entrez Gene Name (culture/xenograft) -7.182 PGM5 phosphoglucomutase 5 -6.883 GPBAR1 G protein-coupled bile acid receptor 1 -6.683 CPVL carboxypeptidase, vitellogenic like -6.398 MTMR9LP myotubularin related protein 9-like, pseudogene -6.131 SCN7A sodium voltage-gated channel alpha subunit 7 -6.115 POPDC2 popeye domain containing 2 -6.014 LGI1 leucine rich glioma inactivated 1 -5.86 SCN1A sodium voltage-gated channel alpha subunit 1 -5.713 C6 complement C6 -5.365 ANGPTL1 angiopoietin like 1 -5.327 TNN tenascin N -5.228 DHRS2 dehydrogenase/reductase 2 leucine rich repeat and fibronectin type III domain -5.115 LRFN2 containing 2 -5.076 FOXO6 forkhead box O6 -5.035 ETNPPL ethanolamine-phosphate phospho-lyase -4.993 MYO15A myosin XVA -4.972 IGF1 insulin like growth factor 1 -4.956 DLG2 discs large MAGUK scaffold protein 2 -4.86 SCML4 sex comb on midleg like 4 (Drosophila) Src homology 2 domain containing transforming -4.816 SHD protein D -4.764 PLP1 proteolipid protein 1 -4.764 TSPAN32 tetraspanin 32 -4.713 N4BP3 NEDD4 binding protein 3 -4.705 MYOC myocilin -4.646 CLEC3B C-type lectin domain family 3 member B -4.646 C7 complement C7 -4.62 TGM2 transglutaminase 2 -4.562 COL9A1 collagen type IX alpha 1 chain -4.55 SOSTDC1 sclerostin domain containing 1 -4.55 OGN osteoglycin -4.505 DAPL1 death associated protein like 1 -4.491 C10orf105 chromosome 10 open reading frame 105 -4.491 -

Binding Mode Exploration of B1 Receptor Antagonists' by the Use of Molecular Dynamics and Docking Simulation—How Different T
International Journal of Molecular Sciences Article Binding Mode Exploration of B1 Receptor Antagonists’ by the Use of Molecular Dynamics and Docking Simulation—How Different Target Engagement Can Determine Different Biological Effects Marica Gemei 1,*, Carmine Talarico 1 , Laura Brandolini 1, Candida Manelfi 1, Lorena Za 2, Silvia Bovolenta 2, Chiara Liberati 2, Luigi Del Vecchio 3, Roberto Russo 4 , Carmen Cerchia 4, Marcello Allegretti 1 and Andrea Rosario Beccari 1 1 Dompé Farmaceutici SpA, via Campo di Pile, 67100 L’Aquila, Italy; [email protected] (C.T.); [email protected] (L.B.); candida.manelfi@dompe.com (C.M.); [email protected] (M.A.); [email protected] (A.R.B.) 2 Axxam, Via Meucci 3, Bresso, 20091 Milano, Italy; [email protected] (L.Z.); [email protected] (S.B.); [email protected] (C.L.) 3 Ceinge Biotecnologie Avanzate, via G. Salvatore 486, 80145 Napoli, Italy; [email protected] 4 Department of Pharmacy, University of Naples “Federico II”, via D. Montesano, 49, 80131 Napoli, Italy; [email protected] (R.R.); [email protected] (C.C.) * Correspondence: [email protected]; Tel.: +34-06-465916 Received: 26 August 2020; Accepted: 12 October 2020; Published: 16 October 2020 Abstract: The kinin B1 receptor plays a critical role in the chronic phase of pain and inflammation. The development of B1 antagonists peaked in recent years but almost all promising molecules failed in clinical trials. Little is known about these molecules’ mechanisms of action and additional information will be necessary to exploit the potential of the B1 receptor. -

Properties of Chemically Oxidized Kininogens*
Vol. 50 No. 3/2003 753–763 QUARTERLY Properties of chemically oxidized kininogens*. Magdalena Nizio³ek, Marcin Kot, Krzysztof Pyka, Pawe³ Mak and Andrzej Kozik½ Faculty of Biotechnology, Jagiellonian University, Kraków, Poland Received: 30 May, 2003; revised: 01 August, 2003; accepted: 11 August, 2003 Key words: bradykinin, N-chlorosuccinimide, chloramine-T, kallidin, kallikrein, reactive oxygen species Kininogens are multifunctional proteins involved in a variety of regulatory pro- cesses including the kinin-formation cascade, blood coagulation, fibrynolysis, inhibi- tion of cysteine proteinases etc. A working hypothesis of this work was that the prop- erties of kininogens may be altered by oxidation of their methionine residues by reac- tive oxygen species that are released at the inflammatory foci during phagocytosis of pathogen particles by recruited neutrophil cells. Two methionine-specific oxidizing reagents, N-chlorosuccinimide (NCS) and chloramine-T (CT), were used to oxidize the high molecular mass (HK) and low molecular mass (LK) forms of human kininogen. A nearly complete conversion of methionine residues to methionine sulfoxide residues in the modified proteins was determined by amino acid analysis. Production of kinins from oxidized kininogens by plasma and tissue kallikreins was significantly lower (by at least 70%) than that from native kininogens. This quenching effect on kinin release could primarily be assigned to the modification of the critical Met-361 residue adja- cent to the internal kinin sequence in kininogen. However, virtually no kinin could be formed by human plasma kallikrein from NCS-modified HK. This observation sug- gests involvement of other structural effects detrimental for kinin production. In- deed, NCS-oxidized HK was unable to bind (pre)kallikrein, probably due to the modifi- cation of methionine and/or tryptophan residues at the region on the kininogen mole- cule responsible for the (pro)enzyme binding. -
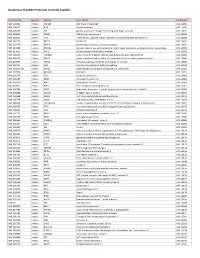
Quantigene Flowrna Probe Sets Currently Available
QuantiGene FlowRNA Probe Sets Currently Available Accession No. Species Symbol Gene Name Catalog No. NM_003452 Human ZNF189 zinc finger protein 189 VA1-10009 NM_000057 Human BLM Bloom syndrome VA1-10010 NM_005269 Human GLI glioma-associated oncogene homolog (zinc finger protein) VA1-10011 NM_002614 Human PDZK1 PDZ domain containing 1 VA1-10015 NM_003225 Human TFF1 Trefoil factor 1 (breast cancer, estrogen-inducible sequence expressed in) VA1-10016 NM_002276 Human KRT19 keratin 19 VA1-10022 NM_002659 Human PLAUR plasminogen activator, urokinase receptor VA1-10025 NM_017669 Human ERCC6L excision repair cross-complementing rodent repair deficiency, complementation group 6-like VA1-10029 NM_017699 Human SIDT1 SID1 transmembrane family, member 1 VA1-10032 NM_000077 Human CDKN2A cyclin-dependent kinase inhibitor 2A (melanoma, p16, inhibits CDK4) VA1-10040 NM_003150 Human STAT3 signal transducer and activator of transcripton 3 (acute-phase response factor) VA1-10046 NM_004707 Human ATG12 ATG12 autophagy related 12 homolog (S. cerevisiae) VA1-10047 NM_000737 Human CGB chorionic gonadotropin, beta polypeptide VA1-10048 NM_001017420 Human ESCO2 establishment of cohesion 1 homolog 2 (S. cerevisiae) VA1-10050 NM_197978 Human HEMGN hemogen VA1-10051 NM_001738 Human CA1 Carbonic anhydrase I VA1-10052 NM_000184 Human HBG2 Hemoglobin, gamma G VA1-10053 NM_005330 Human HBE1 Hemoglobin, epsilon 1 VA1-10054 NR_003367 Human PVT1 Pvt1 oncogene homolog (mouse) VA1-10061 NM_000454 Human SOD1 Superoxide dismutase 1, soluble (amyotrophic lateral sclerosis 1 (adult)) -

Differential Gene Expression in Oligodendrocyte Progenitor Cells, Oligodendrocytes and Type II Astrocytes
Tohoku J. Exp. Med., 2011,Differential 223, 161-176 Gene Expression in OPCs, Oligodendrocytes and Type II Astrocytes 161 Differential Gene Expression in Oligodendrocyte Progenitor Cells, Oligodendrocytes and Type II Astrocytes Jian-Guo Hu,1,2,* Yan-Xia Wang,3,* Jian-Sheng Zhou,2 Chang-Jie Chen,4 Feng-Chao Wang,1 Xing-Wu Li1 and He-Zuo Lü1,2 1Department of Clinical Laboratory Science, The First Affiliated Hospital of Bengbu Medical College, Bengbu, P.R. China 2Anhui Key Laboratory of Tissue Transplantation, Bengbu Medical College, Bengbu, P.R. China 3Department of Neurobiology, Shanghai Jiaotong University School of Medicine, Shanghai, P.R. China 4Department of Laboratory Medicine, Bengbu Medical College, Bengbu, P.R. China Oligodendrocyte precursor cells (OPCs) are bipotential progenitor cells that can differentiate into myelin-forming oligodendrocytes or functionally undetermined type II astrocytes. Transplantation of OPCs is an attractive therapy for demyelinating diseases. However, due to their bipotential differentiation potential, the majority of OPCs differentiate into astrocytes at transplanted sites. It is therefore important to understand the molecular mechanisms that regulate the transition from OPCs to oligodendrocytes or astrocytes. In this study, we isolated OPCs from the spinal cords of rat embryos (16 days old) and induced them to differentiate into oligodendrocytes or type II astrocytes in the absence or presence of 10% fetal bovine serum, respectively. RNAs were extracted from each cell population and hybridized to GeneChip with 28,700 rat genes. Using the criterion of fold change > 4 in the expression level, we identified 83 genes that were up-regulated and 89 genes that were down-regulated in oligodendrocytes, and 92 genes that were up-regulated and 86 that were down-regulated in type II astrocytes compared with OPCs. -

140503 IPF Signatures Supplement Withfigs Thorax
Supplementary material for Heterogeneous gene expression signatures correspond to distinct lung pathologies and biomarkers of disease severity in idiopathic pulmonary fibrosis Daryle J. DePianto1*, Sanjay Chandriani1⌘*, Alexander R. Abbas1, Guiquan Jia1, Elsa N. N’Diaye1, Patrick Caplazi1, Steven E. Kauder1, Sabyasachi Biswas1, Satyajit K. Karnik1#, Connie Ha1, Zora Modrusan1, Michael A. Matthay2, Jasleen Kukreja3, Harold R. Collard2, Jackson G. Egen1, Paul J. Wolters2§, and Joseph R. Arron1§ 1Genentech Research and Early Development, South San Francisco, CA 2Department of Medicine, University of California, San Francisco, CA 3Department of Surgery, University of California, San Francisco, CA ⌘Current address: Novartis Institutes for Biomedical Research, Emeryville, CA. #Current address: Gilead Sciences, Foster City, CA. *DJD and SC contributed equally to this manuscript §PJW and JRA co-directed this project Address correspondence to Paul J. Wolters, MD University of California, San Francisco Department of Medicine Box 0111 San Francisco, CA 94143-0111 [email protected] or Joseph R. Arron, MD, PhD Genentech, Inc. MS 231C 1 DNA Way South San Francisco, CA 94080 [email protected] 1 METHODS Human lung tissue samples Tissues were obtained at UCSF from clinical samples from IPF patients at the time of biopsy or lung transplantation. All patients were seen at UCSF and the diagnosis of IPF was established through multidisciplinary review of clinical, radiological, and pathological data according to criteria established by the consensus classification of the American Thoracic Society (ATS) and European Respiratory Society (ERS), Japanese Respiratory Society (JRS), and the Latin American Thoracic Association (ALAT) (ref. 5 in main text). Non-diseased normal lung tissues were procured from lungs not used by the Northern California Transplant Donor Network. -

Sized Neuropeptides
M ETHODS IN MOLECULAR BIOLOGY™ Series Editor John M. Walker School of Life Sciences University of Hertfordshire Hatfield, Hertfordshire, AL10 9AB, UK For further volumes: http://www.springer.com/series/7651 Neuropeptides Methods and Protocols Edited by Adalberto Merighi Dipartimento di Morfofisiologia Veterinaria, Università degli Studi di Torino, Grugliasco, TO, Italy; Istituto Nazionale di Neuroscienze (INN), Università degli Studi di Torino, Grugliasco, TO, Italy Editor Adalberto Merighi Dipartimento di Morfofisiologia Veterinaria Università degli Studi di Torino and Istituto Nazionale di Neuroscienze (INN) Università degli Studi di Torino Grugliasco, TO, Italy [email protected] Please note that additional material for this book can be downloaded from http://extras.springer.com ISSN 1064-3745 e-ISSN 1940-6029 ISBN 978-1-61779-309-7 e-ISBN 978-1-61779-310-3 DOI 10.1007/978-1-61779-310-3 Springer New York Dordrecht Heidelberg London Library of Congress Control Number: 2011936011 © Springer Science+Business Media, LLC 2011 All rights reserved. This work may not be translated or copied in whole or in part without the written permission of the publisher (Humana Press, c/o Springer Science+Business Media, LLC, 233 Spring Street, New York, NY 10013, USA), except for brief excerpts in connection with reviews or scholarly analysis. Use in connection with any form of information storage and retrieval, electronic adaptation, computer software, or by similar or dissimilar methodology now known or hereafter developed is forbidden. The use in this publication of trade names, trademarks, service marks, and similar terms, even if they are not identified as such, is not to be taken as an expression of opinion as to whether or not they are subject to proprietary rights. -

Sensory and Signaling Mechanisms of Bradykinin, Eicosanoids, Platelet-Activating Factor, and Nitric Oxide in Peripheral Nociceptors
Physiol Rev 92: 1699–1775, 2012 doi:10.1152/physrev.00048.2010 SENSORY AND SIGNALING MECHANISMS OF BRADYKININ, EICOSANOIDS, PLATELET-ACTIVATING FACTOR, AND NITRIC OXIDE IN PERIPHERAL NOCICEPTORS Gábor Peth˝o and Peter W. Reeh Pharmacodynamics Unit, Department of Pharmacology and Pharmacotherapy, Faculty of Medicine, University of Pécs, Pécs, Hungary; and Institute of Physiology and Pathophysiology, University of Erlangen/Nürnberg, Erlangen, Germany Peth˝o G, Reeh PW. Sensory and Signaling Mechanisms of Bradykinin, Eicosanoids, Platelet-Activating Factor, and Nitric Oxide in Peripheral Nociceptors. Physiol Rev 92: 1699–1775, 2012; doi:10.1152/physrev.00048.2010.—Peripheral mediators can contribute to the development and maintenance of inflammatory and neuropathic pain and its concomitants (hyperalgesia and allodynia) via two mechanisms. Activation Lor excitation by these substances of nociceptive nerve endings or fibers implicates generation of action potentials which then travel to the central nervous system and may induce pain sensation. Sensitization of nociceptors refers to their increased responsiveness to either thermal, mechani- cal, or chemical stimuli that may be translated to corresponding hyperalgesias. This review aims to give an account of the excitatory and sensitizing actions of inflammatory mediators including bradykinin, prostaglandins, thromboxanes, leukotrienes, platelet-activating factor, and nitric oxide on nociceptive primary afferent neurons. Manifestations, receptor molecules, and intracellular signaling mechanisms -

RT² Profiler PCR Array (Rotor-Gene® Format) Rat Pain: Neuropathic & Inflammatory
RT² Profiler PCR Array (Rotor-Gene® Format) Rat Pain: Neuropathic & Inflammatory Cat. no. 330231 PARN-162ZR For pathway expression analysis Format For use with the following real-time cyclers RT² Profiler PCR Array, Rotor-Gene Q, other Rotor-Gene cyclers Format R Description The Rat Pain: Neuropathic & Inflammatory RT² Profiler PCR Array profiles the expression of 84 genes involved in the transduction, maintenance, and modulation of pain responses. Noxious environmental stimuli, tissue damage, and disease all evoke pain. Since it afflicts up to 20% of the population at any given time, pain provides both a massive therapeutic target and a route to understanding the molecular mechanisms of nervous system function. While neuropathic pain often results from damage to the peripheral (PNS) or central nervous system (CNS), peripheral tissue damage and/or inflammation generally initiates inflammatory pain. Neuropathic and inflammatory pain both cause activation of damage-sensing neurons (nociceptors) that innervate the skin, muscle and viscera and terminate in the laminae of the spinal cord dorsal horn. Nociceptors conduct information to the CNS via neurotransmission and action potentials generated by ion channel and purinergic, opioid, and cannabinoid receptors leading to second order neuron activation. Synaptic transmission via glutamate, serotonin, and dopamine systems then follows. The transduction by nociceptors can be modulated by mediators of inflammation released by infiltrating immune cells and damaged neurons. Excitability of spinal neurons is also modulated by activation of resident microglia that release growth factors (such as BDNF), chemokines, and cytokines. Endogenous opioid peptides and arachidonic acid metabolites acting through G-protein coupled receptors also modulate neuronal excitability. -

Involvement of the Sigma-1 Receptor in Methamphetamine-Mediated Changes to Astrocyte Structure and Function" (2020)
University of Kentucky UKnowledge Theses and Dissertations--Medical Sciences Medical Sciences 2020 Involvement of the Sigma-1 Receptor in Methamphetamine- Mediated Changes to Astrocyte Structure and Function Richik Neogi University of Kentucky, [email protected] Author ORCID Identifier: https://orcid.org/0000-0002-8716-8812 Digital Object Identifier: https://doi.org/10.13023/etd.2020.363 Right click to open a feedback form in a new tab to let us know how this document benefits ou.y Recommended Citation Neogi, Richik, "Involvement of the Sigma-1 Receptor in Methamphetamine-Mediated Changes to Astrocyte Structure and Function" (2020). Theses and Dissertations--Medical Sciences. 12. https://uknowledge.uky.edu/medsci_etds/12 This Master's Thesis is brought to you for free and open access by the Medical Sciences at UKnowledge. It has been accepted for inclusion in Theses and Dissertations--Medical Sciences by an authorized administrator of UKnowledge. For more information, please contact [email protected]. STUDENT AGREEMENT: I represent that my thesis or dissertation and abstract are my original work. Proper attribution has been given to all outside sources. I understand that I am solely responsible for obtaining any needed copyright permissions. I have obtained needed written permission statement(s) from the owner(s) of each third-party copyrighted matter to be included in my work, allowing electronic distribution (if such use is not permitted by the fair use doctrine) which will be submitted to UKnowledge as Additional File. I hereby grant to The University of Kentucky and its agents the irrevocable, non-exclusive, and royalty-free license to archive and make accessible my work in whole or in part in all forms of media, now or hereafter known. -

Inflammatory Modulation of Hematopoietic Stem Cells by Magnetic Resonance Imaging
Electronic Supplementary Material (ESI) for RSC Advances. This journal is © The Royal Society of Chemistry 2014 Inflammatory modulation of hematopoietic stem cells by Magnetic Resonance Imaging (MRI)-detectable nanoparticles Sezin Aday1,2*, Jose Paiva1,2*, Susana Sousa2, Renata S.M. Gomes3, Susana Pedreiro4, Po-Wah So5, Carolyn Ann Carr6, Lowri Cochlin7, Ana Catarina Gomes2, Artur Paiva4, Lino Ferreira1,2 1CNC-Center for Neurosciences and Cell Biology, University of Coimbra, Coimbra, Portugal, 2Biocant, Biotechnology Innovation Center, Cantanhede, Portugal, 3King’s BHF Centre of Excellence, Cardiovascular Proteomics, King’s College London, London, UK, 4Centro de Histocompatibilidade do Centro, Coimbra, Portugal, 5Department of Neuroimaging, Institute of Psychiatry, King's College London, London, UK, 6Cardiac Metabolism Research Group, Department of Physiology, Anatomy & Genetics, University of Oxford, UK, 7PulseTeq Limited, Chobham, Surrey, UK. *These authors contributed equally to this work. #Correspondence to Lino Ferreira ([email protected]). Experimental Section Preparation and characterization of NP210-PFCE. PLGA (Resomers 502 H; 50:50 lactic acid: glycolic acid) (Boehringer Ingelheim) was covalently conjugated to fluoresceinamine (Sigma- Aldrich) according to a protocol reported elsewhere1. NPs were prepared by dissolving PLGA (100 mg) in a solution of propylene carbonate (5 mL, Sigma). PLGA solution was mixed with perfluoro- 15-crown-5-ether (PFCE) (178 mg) (Fluorochem, UK) dissolved in trifluoroethanol (1 mL, Sigma). This solution was then added to a PVA solution (10 mL, 1% w/v in water) dropwise and stirred for 3 h. The NPs were then transferred to a dialysis membrane and dialysed (MWCO of 50 kDa, Spectrum Labs) against distilled water before freeze-drying. Then, NPs were coated with protamine sulfate (PS).