A Merged Microarray Meta-Dataset for Transcriptionally Profiling Colorectal Neoplasm Formation and Progression
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Histone Isoform H2A1H Promotes Attainment of Distinct Physiological
Bhattacharya et al. Epigenetics & Chromatin (2017) 10:48 DOI 10.1186/s13072-017-0155-z Epigenetics & Chromatin RESEARCH Open Access Histone isoform H2A1H promotes attainment of distinct physiological states by altering chromatin dynamics Saikat Bhattacharya1,4,6, Divya Reddy1,4, Vinod Jani5†, Nikhil Gadewal3†, Sanket Shah1,4, Raja Reddy2,4, Kakoli Bose2,4, Uddhavesh Sonavane5, Rajendra Joshi5 and Sanjay Gupta1,4* Abstract Background: The distinct functional efects of the replication-dependent histone H2A isoforms have been dem- onstrated; however, the mechanistic basis of the non-redundancy remains unclear. Here, we have investigated the specifc functional contribution of the histone H2A isoform H2A1H, which difers from another isoform H2A2A3 in the identity of only three amino acids. Results: H2A1H exhibits varied expression levels in diferent normal tissues and human cancer cell lines (H2A1C in humans). It also promotes cell proliferation in a context-dependent manner when exogenously overexpressed. To uncover the molecular basis of the non-redundancy, equilibrium unfolding of recombinant H2A1H-H2B dimer was performed. We found that the M51L alteration at the H2A–H2B dimer interface decreases the temperature of melting of H2A1H-H2B by ~ 3 °C as compared to the H2A2A3-H2B dimer. This diference in the dimer stability is also refected in the chromatin dynamics as H2A1H-containing nucleosomes are more stable owing to M51L and K99R substitu- tions. Molecular dynamic simulations suggest that these substitutions increase the number of hydrogen bonds and hydrophobic interactions of H2A1H, enabling it to form more stable nucleosomes. Conclusion: We show that the M51L and K99R substitutions, besides altering the stability of histone–histone and histone–DNA complexes, have the most prominent efect on cell proliferation, suggesting that the nucleosome sta- bility is intimately linked with the physiological efects observed. -

Differences for Ectopic Versus Eutopic Cells
556 RBMO VOLUME 39 ISSUE 4 2019 ARTICLE Chemosensitivity and chemoresistance in endometriosis – differences for ectopic versus eutopic cells BIOGRAPHY Andres Salumets is Professor of Reproductive Medicine at the University of Tartu, and Scientific Head at the Competence Centre on Health Technologies, Tartu, Estonia. He has been involved in assisted reproduction for 20 years, first as an embryologist and later as a researcher. His major interests are endometriosis, endometrial biology and implantation. Darja Lavogina1,2,*, Külli Samuel1, Arina Lavrits1,3, Alvin Meltsov1, Deniss Sõritsa1,4,5, Ülle Kadastik6, Maire Peters1,4, Ago Rinken2, Andres Salumets1,4,7, 8 KEY MESSAGE Akt/PKB inhibitor GSK690693, CK2 inhibitor ARC-775, MAPK pathway inhibitor sorafenib, proteasome inhibitor bortezomib, and microtubule-depolymerizing toxin MMAE showed higher cytotoxicity in eutopic cells. In contrast, 10 µmol/l of the anthracycline toxin doxorubicin caused cellular death in ectopic cells more effectively than in eutopic cells, underlining the potential of doxorubicin in endometriosis research. ABSTRACT Research question: Endometriosis is a common gynaecological disease defined by the presence of endometrium-like tissue outside the uterus. This complex disease, often accompanied by severe pain and infertility, causes a significant medical and socioeconomic burden; hence, novel strategies are being sought for the treatment of endometriosis. Here, we set out to explore the cytotoxic effects of a panel of compounds to find toxins with different efficiency in eutopic versus ectopic cells, thus highlighting alterations in the corresponding molecular pathways. Design: The effect on cellular viability of 14 compounds was established in a cohort of paired eutopic and ectopic endometrial stromal cell samples from 11 patients. -

Reconstructing Cell Cycle Pseudo Time-Series Via Single-Cell Transcriptome Data—Supplement
School of Natural Sciences and Mathematics Reconstructing Cell Cycle Pseudo Time-Series Via Single-Cell Transcriptome Data—Supplement UT Dallas Author(s): Michael Q. Zhang Rights: CC BY 4.0 (Attribution) ©2017 The Authors Citation: Liu, Zehua, Huazhe Lou, Kaikun Xie, Hao Wang, et al. 2017. "Reconstructing cell cycle pseudo time-series via single-cell transcriptome data." Nature Communications 8, doi:10.1038/s41467-017-00039-z This document is being made freely available by the Eugene McDermott Library of the University of Texas at Dallas with permission of the copyright owner. All rights are reserved under United States copyright law unless specified otherwise. File name: Supplementary Information Description: Supplementary figures, supplementary tables, supplementary notes, supplementary methods and supplementary references. CCNE1 CCNE1 CCNE1 CCNE1 36 40 32 34 32 35 30 32 28 30 30 28 28 26 24 25 Normalized Expression Normalized Expression Normalized Expression Normalized Expression 26 G1 S G2/M G1 S G2/M G1 S G2/M G1 S G2/M Cell Cycle Stage Cell Cycle Stage Cell Cycle Stage Cell Cycle Stage CCNE1 CCNE1 CCNE1 CCNE1 40 32 40 40 35 30 38 30 30 28 36 25 26 20 20 34 Normalized Expression Normalized Expression Normalized Expression 24 Normalized Expression G1 S G2/M G1 S G2/M G1 S G2/M G1 S G2/M Cell Cycle Stage Cell Cycle Stage Cell Cycle Stage Cell Cycle Stage Supplementary Figure 1 | High stochasticity of single-cell gene expression means, as demonstrated by relative expression levels of gene Ccne1 using the mESC-SMARTer data. For every panel, 20 sample cells were randomly selected for each of the three stages, followed by plotting the mean expression levels at each stage. -

(12) United States Patent (10) Patent No.: US 7,799,528 B2 Civin Et Al
US007799528B2 (12) United States Patent (10) Patent No.: US 7,799,528 B2 Civin et al. (45) Date of Patent: Sep. 21, 2010 (54) THERAPEUTIC AND DIAGNOSTIC Al-Hajj et al., “Prospective identification of tumorigenic breast can APPLICATIONS OF GENES cer cells.” Proc. Natl. Acad. Sci. U.S.A., 100(7):3983-3988 (2003). DIFFERENTIALLY EXPRESSED IN Bhatia et al., “A newly discovered class of human hematopoietic cells LYMPHO-HEMATOPOETC STEM CELLS with SCID-repopulating activity.” Nat. Med. 4(9)1038-45 (1998). (75) Inventors: Curt I. Civin, Baltimore, MD (US); Bonnet, D., “Normal and leukemic CD35-negative human Robert W. Georgantas, III, Towson, hematopoletic stem cells.” Rev. Clin. Exp. Hematol. 5:42-61 (2001). Cambot et al., “Human Immune Associated Nucleotide 1: a member MD (US) of a new guanosine triphosphatase family expressed in resting T and (73) Assignee: The Johns Hopkins University, B cells.” Blood, 99(9):3293-3301 (2002). Baltimore, MD (US) Chen et al., “Kruppel-like Factor 4 (Gut-enriched Kruppel-like Fac tor) Inhibits Cell Proliferation by Blocking G1/S Progression of the *) NotOt1Ce: Subjubject to anyy d1Sclaimer,disclai theh term off thisthi Cell Cycle.” J. Biol. Chem., 276(32):30423-30428 (2001). patent is extended or adjusted under 35 Chen et al., “Transcriptional profiling of Kurppel-like factor 4 reveals U.S.C. 154(b) by 168 days. a function in cell cycle regulation and epithelial differentiation.” J. Mol. Biol. 326(3):665-677 (2003). (21) Appl. No.: 11/199.665 Civin et al., “Highly purified CD34-positive cells reconstitute (22) Filed: Aug. 9, 2005 hematopoiesis,” J. -

Pharmacological Unmasking Microarray Approach-Based Discovery of Novel DNA Methylation Markers for Hepatocellular Carcinoma
ORIGINAL ARTICLE Oncology & Hematology http://dx.doi.org/10.3346/jkms.2012.27.6.594 • J Korean Med Sci 2012; 27: 594-604 Pharmacological Unmasking Microarray Approach-Based Discovery of Novel DNA Methylation Markers for Hepatocellular Carcinoma Namhee Jung1, Jae Kyung Won2, DNA methylation is one of the main epigenetic mechanisms and hypermethylation of CpG Baek-Hui Kim3, Kyung Suk Suh4, islands at tumor suppressor genes switches off these genes. To find novel DNA methylation Ja-June Jang2, and Gyeong Hoon Kang1,2 markers in hepatocellular carcinoma (HCC), we performed pharmacological unmasking (treatment with 5-aza-2´-deoxycytidine or trichostatin A) followed by microarray analysis 1Laboratory of Epigenetics, Cancer Research Institute, 2Departments of Pathology, 3General in HCC cell lines. Of the 239 promoter CpG island loci hypermethylated in HCC cell lines Surgery, Seoul National University College of (as revealed by methylation-specific PCR), 221 loci were found to be hypermethylated in Medicine, Seoul; 4Department of Pathology, Korea HCC or nonneoplastic liver tissues. Thirty-three loci showed a 20% higher methylation University School of Medicine, Seoul, Korea frequency in tumors than in adjacent nonneoplastic tissues. Correlation of individual Received: 21 November 2011 cancer-related methylation markers with clinicopathological features of HCC patients Accepted: 23 February 2012 (n = 95) revealed that the number of hypermethylated genes in HCC tumors was higher in older than in younger patients. Univariate and multivariate survival analysis revealed that Address for Correspondence: the HIST1H2AE methylation status is closely correlated with the patient’s overall survival Gyeong Hoon Kang, MD Department of Pathology, Seoul National University College of (P = 0.022 and P = 0.010, respectively). -

A Yeast Phenomic Model for the Influence of Warburg Metabolism on Genetic Buffering of Doxorubicin Sean M
Santos and Hartman Cancer & Metabolism (2019) 7:9 https://doi.org/10.1186/s40170-019-0201-3 RESEARCH Open Access A yeast phenomic model for the influence of Warburg metabolism on genetic buffering of doxorubicin Sean M. Santos and John L. Hartman IV* Abstract Background: The influence of the Warburg phenomenon on chemotherapy response is unknown. Saccharomyces cerevisiae mimics the Warburg effect, repressing respiration in the presence of adequate glucose. Yeast phenomic experiments were conducted to assess potential influences of Warburg metabolism on gene-drug interaction underlying the cellular response to doxorubicin. Homologous genes from yeast phenomic and cancer pharmacogenomics data were analyzed to infer evolutionary conservation of gene-drug interaction and predict therapeutic relevance. Methods: Cell proliferation phenotypes (CPPs) of the yeast gene knockout/knockdown library were measured by quantitative high-throughput cell array phenotyping (Q-HTCP), treating with escalating doxorubicin concentrations under conditions of respiratory or glycolytic metabolism. Doxorubicin-gene interaction was quantified by departure of CPPs observed for the doxorubicin-treated mutant strain from that expected based on an interaction model. Recursive expectation-maximization clustering (REMc) and Gene Ontology (GO)-based analyses of interactions identified functional biological modules that differentially buffer or promote doxorubicin cytotoxicity with respect to Warburg metabolism. Yeast phenomic and cancer pharmacogenomics data were integrated to predict differential gene expression causally influencing doxorubicin anti-tumor efficacy. Results: Yeast compromised for genes functioning in chromatin organization, and several other cellular processes are more resistant to doxorubicin under glycolytic conditions. Thus, the Warburg transition appears to alleviate requirements for cellular functions that buffer doxorubicin cytotoxicity in a respiratory context. -

Rap1-Mediated Chromatin and Gene Expression Changes at Senescence
University of Pennsylvania ScholarlyCommons Publicly Accessible Penn Dissertations 2019 Rap1-Mediated Chromatin And Gene Expression Changes At Senescence Shufei Song University of Pennsylvania Follow this and additional works at: https://repository.upenn.edu/edissertations Part of the Biochemistry Commons, and the Cell Biology Commons Recommended Citation Song, Shufei, "Rap1-Mediated Chromatin And Gene Expression Changes At Senescence" (2019). Publicly Accessible Penn Dissertations. 3557. https://repository.upenn.edu/edissertations/3557 This paper is posted at ScholarlyCommons. https://repository.upenn.edu/edissertations/3557 For more information, please contact [email protected]. Rap1-Mediated Chromatin And Gene Expression Changes At Senescence Abstract ABSTRACT RAP1-MEDIATED CHROMATIN AND GENE EXPRESSION CHANGES AT SENESCENCE The telomeric protein Rap1 has been extensively studied for its roles as a transcriptional activator and repressor. Indeed, in both yeast and mammals, Rap1 is known to bind throughout the genome to reorganize chromatin and regulate gene transcription. Previously, our lab published evidence that Rap1 plays important roles in cellular senescence. In telomerase-deficient S. cerevisiae, Rap1 relocalizes from telomeres and subtelomeres to new Rap1 target at senescence (NRTS). This leads to two types of histone loss: Rap1 lowers global histone levels by repressing histone gene transcription and it also results in local nucleosome displacement at the promoters of the activated NRTS. Here, I examine mechanisms of site-specific histone loss by presenting evidence that Rap1 can directly interact with histone tetramers H3/H4, and map this interaction to a three-amino-acid-patch within the DNA binding domain. Functional studies are performed in vivo using a mutant form of Rap1 with weakened histone interactions, and deficient promoter clearance as well as blunted gene activation is observed, indicating that direct Rap1-H3/H4 interactions are involved in nucleosome displacement. -

Early During Myelomagenesis Alterations in DNA Methylation That
Myeloma Is Characterized by Stage-Specific Alterations in DNA Methylation That Occur Early during Myelomagenesis This information is current as Christoph J. Heuck, Jayesh Mehta, Tushar Bhagat, Krishna of September 23, 2021. Gundabolu, Yiting Yu, Shahper Khan, Grigoris Chrysofakis, Carolina Schinke, Joseph Tariman, Eric Vickrey, Natalie Pulliam, Sangeeta Nischal, Li Zhou, Sanchari Bhattacharyya, Richard Meagher, Caroline Hu, Shahina Maqbool, Masako Suzuki, Samir Parekh, Frederic Reu, Ulrich Steidl, John Greally, Amit Verma and Seema B. Downloaded from Singhal J Immunol 2013; 190:2966-2975; Prepublished online 13 February 2013; doi: 10.4049/jimmunol.1202493 http://www.jimmunol.org/content/190/6/2966 http://www.jimmunol.org/ Supplementary http://www.jimmunol.org/content/suppl/2013/02/13/jimmunol.120249 Material 3.DC1 References This article cites 38 articles, 15 of which you can access for free at: http://www.jimmunol.org/content/190/6/2966.full#ref-list-1 by guest on September 23, 2021 Why The JI? Submit online. • Rapid Reviews! 30 days* from submission to initial decision • No Triage! Every submission reviewed by practicing scientists • Fast Publication! 4 weeks from acceptance to publication *average Subscription Information about subscribing to The Journal of Immunology is online at: http://jimmunol.org/subscription Permissions Submit copyright permission requests at: http://www.aai.org/About/Publications/JI/copyright.html Email Alerts Receive free email-alerts when new articles cite this article. Sign up at: http://jimmunol.org/alerts The Journal of Immunology is published twice each month by The American Association of Immunologists, Inc., 1451 Rockville Pike, Suite 650, Rockville, MD 20852 Copyright © 2013 by The American Association of Immunologists, Inc. -
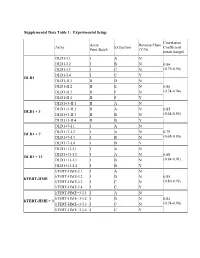
Table 3: Average Gene Expression Profiles by Chromosome
Supplemental Data Table 1: Experimental Setup Correlation Array Reverse Fluor Array Extraction Coefficient Print Batch (Y/N) mean (range) DLD1-I.1 I A N DLD1-I.2 I B N 0.86 DLD1-I.3 I C N (0.79-0.90) DLD1-I.4 I C Y DLD1 DLD1-II.1 II D N DLD1-II.2 II E N 0.86 DLD1-II.3 II F N (0.74-0.94) DLD1-II.4 II F Y DLD1+3-II.1 II A N DLD1+3-II.2 II A N 0.85 DLD1 + 3 DLD1+3-II.3 II B N (0.64-0.95) DLD1+3-II.4 II B Y DLD1+7-I.1 I A N DLD1+7-I.2 I A N 0.79 DLD1 + 7 DLD1+7-I.3 I B N (0.68-0.90) DLD1+7-I.4 I B Y DLD1+13-I.1 I A N DLD1+13-I.2 I A N 0.88 DLD1 + 13 DLD1+13-I.3 I B N (0.84-0.91) DLD1+13-I.4 I B Y hTERT-HME-I.1 I A N hTERT-HME-I.2 I B N 0.85 hTERT-HME hTERT-HME-I.3 I C N (0.80-0.92) hTERT-HME-I.4 I C Y hTERT-HME+3-I.1 I A N hTERT-HME+3-I.2 I B N 0.84 hTERT-HME + 3 hTERT-HME+3-I.3 I C N (0.74-0.90) hTERT-HME+3-I.4 I C Y Supplemental Data Table 2: Average gene expression profiles by chromosome arm DLD1 hTERT-HME Ratio.7 Ratio.1 Ratio.3 Ratio.3 Chrom. -

Hidden Markov Models Predict Epigenetic Chromatin Domains
Hidden Markov Models Predict Epigenetic Chromatin Domains The Harvard community has made this article openly available. Please share how this access benefits you. Your story matters Citation Larson, Jessica. 2012. Hidden Markov Models Predict Epigenetic Chromatin Domains. Doctoral dissertation, Harvard University. Citable link http://nrs.harvard.edu/urn-3:HUL.InstRepos:10087396 Terms of Use This article was downloaded from Harvard University’s DASH repository, and is made available under the terms and conditions applicable to Other Posted Material, as set forth at http:// nrs.harvard.edu/urn-3:HUL.InstRepos:dash.current.terms-of- use#LAA ©2012- Jessica Lynn Larson All rights reserved Dissertation Advisor: Professor Guo-Cheng Yuan Jessica Lynn Larson Hidden Markov Models Predict Epigenetic Chromatin Domains Abstract Epigenetics is an important layer of transcriptional control necessary for cell-type specific gene regulation. We developed computational methods to analyze the combinatorial effect and large-scale organizations of genome-wide distributions of epigenetic marks. Throughout this dissertation, we show that regions containing multiple genes with similar epigenetic patterns are found throughout the genome, suggesting the presence of several chromatin domains. In Chapter 1, we develop a hidden Markov model (HMM) for detecting the types and locations of epigenetic domains from multiple histone modifications. We use this method to analyze a published ChIP-seq dataset of five histone modification marks in mouse embryonic stem cells. We successfully detect domains of consistent epigenetic patterns from ChIP-seq data, providing new insights into the role of epigenetics in long- range gene regulation. In Chapter 2, we expand our model to investigate the genome-wide patterns of histone modifications in multiple human cell lines. -

Characterization of Chromatin Interaction in Mammalian Cells Jufen Zhu University of Connecticut - Storrs, [email protected]
University of Connecticut OpenCommons@UConn Doctoral Dissertations University of Connecticut Graduate School 1-8-2019 Characterization of Chromatin Interaction in Mammalian Cells Jufen Zhu University of Connecticut - Storrs, [email protected] Follow this and additional works at: https://opencommons.uconn.edu/dissertations Recommended Citation Zhu, Jufen, "Characterization of Chromatin Interaction in Mammalian Cells" (2019). Doctoral Dissertations. 2053. https://opencommons.uconn.edu/dissertations/2053 Characterization of Chromatin Interaction in Mammalian Cells Jacqueline Jufen Zhu, Ph.D. University of Connecticut, 2019 Abstract Higher-order chromatin organization in cell nucleus is mysterious. Microscopy and high- throughput sequencing technologies have been applied to reveal the connections between genome structure and gene transcription, which is the main topic of my thesis. Higher-order chromatin organization differs across cell types and species, giving an insight into disease genesis and tumorigenesis. Despite an enormous progress has been made recently in epigenomic study, lots of things remain unknown. My dissertation reviews the current understanding of epigenomics, characterizes chromatin interaction in mammalian cells using different technologies, investigates chromatin reorganization in breast cancer cells, analyzes integrated genomic and epigenomic data in breast cancer metastasis and discusses results and future directions. Dissertation Jacqueline Jufen Zhu Characterization of Chromatin Interaction in Mammalian Cells Jacqueline Jufen Zhu B.E., Nanjing Agricultural University, 2010 M.S., Chinese Academy of Sciences, 2013 A Dissertation Submitted in Partial Fulfillment of the Requirements for the Degree of Doctor of Philosophy at the University of Connecticut 2019 i Dissertation Jacqueline Jufen Zhu Copyright by Jacqueline Jufen Zhu 2019 ii Dissertation Jacqueline Jufen Zhu Approval Page Doctoral of Philosophy Dissertation Characterization of Chromatin Interaction in Mammalian Cells Presented by Jacqueline Jufen Zhu, M.S. -

The in Vitro and in Vivo Effects of Re-Expressing Methylated Von Hippel-Lindau Tumor Suppressor Gene in Clear Cell Renal Carcinoma with 5-Aza-2-Deoxycytidine
Vol. 10, 7011–7021, October 15, 2004 Clinical Cancer Research 7011 The In vitro and In vivo Effects of Re-Expressing Methylated von Hippel-Lindau Tumor Suppressor Gene in Clear Cell Renal Carcinoma with 5-Aza-2-deoxycytidine Wade G. Alleman,1,2 Ray L. Tabios,2 Well described phenotypic changes of VHL expression in- Gadisetti V. R. Chandramouli,3 cluding decreased invasiveness into Matrigel, and decreased Olga N. Aprelikova,3 Carlos Torres-Cabala,2 vascular endothelial growth factor and glucose transport- 4 5 er-1 expression were observed in the treated lines. VHL Arnulfo Mendoza, Craig Rodgers, methylated ccRCC xenografted tumors were significantly 2 2 Nikolai A. Sopko, W. Marston Linehan, and reduced in size in mice treated with 5-aza-dCyd. Mice bear- James R. Vasselli2 ing nonmethylated but VHL-mutated tumors showed no 1Howard Hughes Medical Institute, Chevy Chase, MD; 2Urology tumor shrinkage with 5-aza-dCyd treatment. Branch, 3Laboratory of Biosystems and Cancer, and 4Pediatric Conclusion: Hypo-methylating agents may be useful in Oncology Branch, Center for Cancer Research, National Cancer 5 the treatment of patients having ccRCC tumors consisting of Institute, Bethesda, Maryland; The Brady Urologic Institute, The cells with methylated VHL. Johns Hopkins Hospital, Baltimore, Maryland INTRODUCTION ABSTRACT An estimated 31,900 people are diagnosed annually with Purpose: Clear cell renal carcinoma (ccRCC) is cancer of the kidney in the United States, of which the majority strongly associated with loss of the von Hippel-Lindau (VHL) are clear cell type. In sporadic clear cell renal carcinoma tumor suppressor gene. The VHL gene is functionally lost (ccRCC), 50–85% of patients are found to have biallelic loss of through hypermethylation in up to 19% of sporadic ccRCC the von Hippel-Lindau (VHL) tumor suppressor gene (1–3), cases.