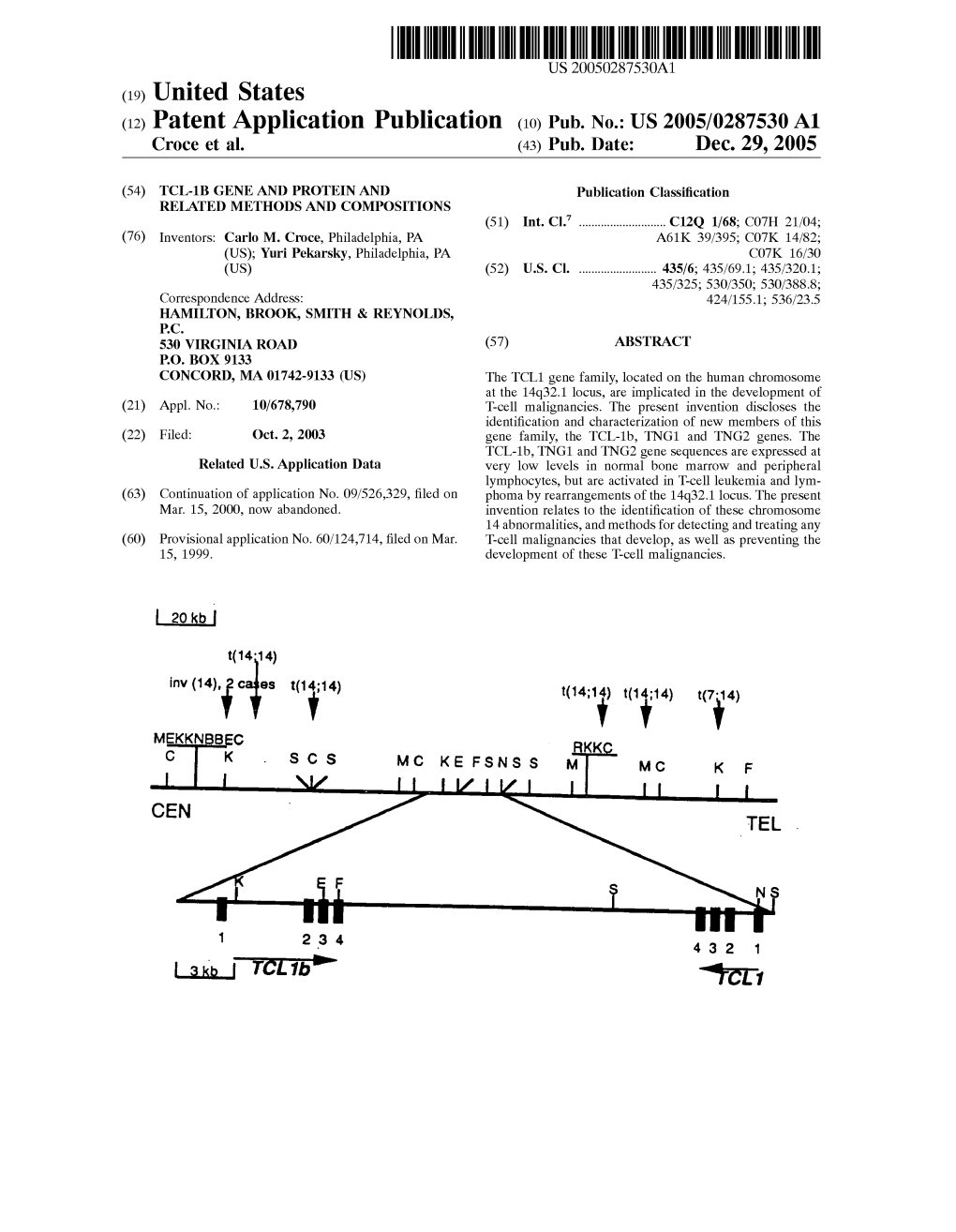
R" 9S "" T(14:14) T(14:14) To/14) MEKKNBBEC RKK S C S M C K E F S N S S M
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Molecular Dynamics and Evolutionary Aspects of the Transition from the Fully Grown Oocyte to Embryo
Downloaded from genesdev.cshlp.org on September 26, 2021 - Published by Cold Spring Harbor Laboratory Press Cracking the egg: molecular dynamics and evolutionary aspects of the transition from the fully grown oocyte to embryo Alexei V. Evsikov,1,5 Joel H. Graber,1 J. Michael Brockman,1,2 Aleš Hampl,3 Andrea E. Holbrook,1 Priyam Singh,1,2 John J. Eppig,1 Davor Solter,1,4 and Barbara B. Knowles1 1The Jackson Laboratory, Bar Harbor, Maine 04609, USA; 2 Bioinformatics Program, Boston University, Boston, Massachusetts 02215, USA; 3Masaryk University Brno and Institute of Experimental Medicine, 625 00 Brno, Czech Republic; 4Max Planck Institute of Immunobiology, 79108 Freiburg, Germany Fully grown oocytes (FGOs) contain all the necessary transcripts to activate molecular pathways underlying the oocyte-to-embryo transition (OET). To elucidate this critical period of development, an extensive survey of the FGO transcriptome was performed by analyzing 19,000 expressed sequence tags of the Mus musculus FGO cDNA library. Expression of 5400 genes and transposable elements is reported. For a majority of genes expressed in mouse FGOs, homologs transcribed in eggs of Xenopus laevis or Ciona intestinalis were found, pinpointing evolutionary conservation of most regulatory cascades underlying the OET in chordates. A large proportion of identified genes belongs to several gene families with oocyte-restricted expression, a likely result of lineage-specific genomic duplications. Gene loss by mutation and expression in female germline of retrotransposed genes specific to M. musculus is documented. These findings indicate rapid diversification of genes involved in female reproduction. Comparison of the FGO and two-cell embryo transcriptomes demarcated the processes important for oogenesis from those involved in OET and identified novel motifs in maternal mRNAs associated with transcript stability. -

The Cytogenetics of Hematologic Neoplasms 1 5
The Cytogenetics of Hematologic Neoplasms 1 5 Aurelia Meloni-Ehrig that errors during cell division were the basis for neoplastic Introduction growth was most likely the determining factor that inspired early researchers to take a better look at the genetics of the The knowledge that cancer is a malignant form of uncon- cell itself. Thus, the need to have cell preparations good trolled growth has existed for over a century. Several biologi- enough to be able to understand the mechanism of cell cal, chemical, and physical agents have been implicated in division became of critical importance. cancer causation. However, the mechanisms responsible for About 50 years after Boveri’s chromosome theory, the this uninhibited proliferation, following the initial insult(s), fi rst manuscripts on the chromosome makeup in normal are still object of intense investigation. human cells and in genetic disorders started to appear, fol- The fi rst documented studies of cancer were performed lowed by those describing chromosome changes in neoplas- over a century ago on domestic animals. At that time, the tic cells. A milestone of this investigation occurred in 1960 lack of both theoretical and technological knowledge with the publication of the fi rst article by Nowell and impaired the formulations of conclusions about cancer, other Hungerford on the association of chronic myelogenous leu- than the visible presence of new growth, thus the term neo- kemia with a small size chromosome, known today as the plasm (from the Greek neo = new and plasma = growth). In Philadelphia (Ph) chromosome, to honor the city where it the early 1900s, the fundamental role of chromosomes in was discovered (see also Chap. -

Protooncogene Tcl1b Functions As an Akt Kinase Co-Activator That Exhibits Oncogenic Potency in Vivo
OPEN Citation: Oncogenesis (2013) 2, e70; doi:10.1038/oncsis.2013.30 & 2013 Macmillan Publishers Limited All rights reserved 2157-9024/13 www.nature.com/oncsis ORIGINAL ARTICLE Protooncogene TCL1b functions as an Akt kinase co-activator that exhibits oncogenic potency in vivo M Hashimoto1, F Suizu1, W Tokuyama2, H Noguchi3, N Hirata1, M Matsuda-Lennikov1, T Edamura1, M Masuzawa4, N Gotoh5, S Tanaka6 and M Noguchi1 Protooncogene T-cell leukemia 1 (TCL1), which is implicated in human T-cell prolymphocytic leukemia (T-PLL), interacts with Akt and enhances its kinase activity, functioning as an Akt kinase co-activator. Two major isoforms of TCL1 Protooncogenes (TCL1 and TCL1b) are present adjacent to each other on human chromosome 14q.32. In human T-PLL, both TCL1 and TCL1b are activated by chromosomal translocation. Moreover, TCL1b-transgenic mice have never been created. Therefore, it remains unclear whether TCL1b itself, independent of TCL1, exhibits oncogenicity. In co-immunoprecipitation assays, both ectopic and endogenous TCL1b interacted with Akt. In in vitro Akt kinase assays, TCL1b enhanced Akt kinase activity in dose- and time-dependent manners. Bioinformatics approaches utilizing multiregression analysis, cluster analysis, KEGG (Kyoto Encyclopedia of Genes and Genomes) pathway mapping, Venn diagrams and Gene Ontology (GO) demonstrated that TCL1b showed highly homologous gene-induction signatures similar to Myr-Akt or TCL1. TCL1b exhibited oncogenicity in in vitro colony-transformation assay. Further, two independent lines of b-actin promoter-driven TCL1b-transgenic mice developed angiosarcoma on the intestinal tract. Angiosarcoma is a rare form of cancer in humans with poor prognosis. Using immunohistochemistry, 11 out of 13 human angiosarcoma samples were positively stained with both anti-TCL1b and anti-phospho-Akt antibodies. -

Atlas Journal
Atlas of Genetics and Cytogenetics in Oncology and Haematology Home Genes Leukemias Solid Tumours Cancer-Prone Deep Insight Portal Teaching X Y 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 NA Atlas Journal Atlas Journal versus Atlas Database: the accumulation of the issues of the Journal constitutes the body of the Database/Text-Book. TABLE OF CONTENTS Volume 12, Number 5, Sep-Oct 2008 Previous Issue / Next Issue Genes XAF1 (XIAP associated factor-1) (17p13.2). Stéphanie Plenchette, Wai Gin Fong, Robert G Korneluk. Atlas Genet Cytogenet Oncol Haematol 2008; 12 (5): 668-673. [Full Text] [PDF] URL : http://atlasgeneticsoncology.org/Genes/XAF1ID44095ch17p13.html WWP1 (WW domain containing E3 ubiquitin protein ligase 1) (8q21.3). Ceshi Chen. Atlas Genet Cytogenet Oncol Haematol 2008; 12 (5): 674-680. [Full Text] [PDF] URL : http://atlasgeneticsoncology.org/Genes/WWP1ID42993ch8q21.html TSPAN1 (tetraspanin 1) (1p34.1). David Murray, Peter Doran. Atlas Genet Cytogenet Oncol Haematol 2008; 12 (5): 681-683. [Full Text] [PDF] URL : http://atlasgeneticsoncology.org/Genes/TSPAN1ID44178ch1p34.html TCL1B (T-cell leukemia/lymphoma 1B) (14q32.13). Herbert Eradat, Michael A Teitell. Atlas Genet Cytogenet Oncol Haematol 2008; 12 (5): 684-686. [Full Text] [PDF] URL : http://atlasgeneticsoncology.org/Genes/TCL1BID354ch14q32.html PVRL4 (poliovirus receptor-related 4) (1q23.3). Marc Lopez. Atlas Genet Cytogenet Oncol Haematol 2008; 12 (5): 687-690. [Full Text] [PDF] URL : http://atlasgeneticsoncology.org/Genes/PVRL4ID44141ch1q23.html PTTG1IP (pituitary tumor-transforming 1 interacting protein) (21q22.3). Vicki Smith, Chris McCabe. Atlas Genet Cytogenet Oncol Haematol 2008; 12 (5): 691-694. -

DNA Methylation Abnormalities in Congenital Heart Disease
Epigenetics ISSN: 1559-2294 (Print) 1559-2308 (Online) Journal homepage: https://www.tandfonline.com/loi/kepi20 DNA methylation abnormalities in congenital heart disease Clara Serra-Juhé, Ivon Cuscó, Aïda Homs, Raquel Flores, Núria Torán & Luis A Pérez-Jurado To cite this article: Clara Serra-Juhé, Ivon Cuscó, Aïda Homs, Raquel Flores, Núria Torán & Luis A Pérez-Jurado (2015) DNA methylation abnormalities in congenital heart disease, Epigenetics, 10:2, 167-177, DOI: 10.1080/15592294.2014.998536 To link to this article: https://doi.org/10.1080/15592294.2014.998536 © 2015 The Author(s). Published with View supplementary material license by Taylor & Francis Group, LLC© Clara Serra-Juhé, Ivon Cuscó, Aïda Homs, Raquel Flores, Núria Torán, and Luis A Pérez-Jurado Accepted author version posted online: 14 Submit your article to this journal Jan 2015. Published online: 17 Mar 2015. Article views: 2326 View related articles View Crossmark data Citing articles: 42 View citing articles Full Terms & Conditions of access and use can be found at https://www.tandfonline.com/action/journalInformation?journalCode=kepi20 RESEARCH PAPER Epigenetics 10:2, 167--177; February 2015; Published with license by Taylor & Francis Group, LLC DNA methylation abnormalities in congenital heart disease Clara Serra-Juhe1,2,3, Ivon Cusco1,2,3,A€ıda Homs1,2,3, Raquel Flores1,2,3,Nuria Toran4,2, and Luis A Perez-Jurado 1,2,3,* 1Department of Experimental and Health Sciences; Universitat Pompeu Fabra; Barcelona, Spain; 2Centro de Investigacion Biomedica en Red de Enfermedades -

Application of the High-Throughput TAB-Array for the Discovery of Novel 5-Hydroxymethylcytosine Biomarkers in Pancreatic Ductal Adenocarcinoma
epigenomes Article Application of the High-Throughput TAB-Array for the Discovery of Novel 5-Hydroxymethylcytosine Biomarkers in Pancreatic Ductal Adenocarcinoma 1,2, 1, 1 3 1,4, Chang Zeng y , Zhou Zhang y, Jun Wang , Brian C-H Chiu , Lifang Hou z and 1,4, , Wei Zhang * z 1 Department of Preventive Medicine, Northwestern University Feinberg School of Medicine, Chicago, IL 60611, USA 2 Driskill Graduate Program in Life Sciences, Northwestern University Feinberg School of Medicine, Chicago, IL 60611, USA 3 Department of Public Health Sciences, the University of Chicago, Chicago, IL 60637, USA 4 The Robert H. Lurie Comprehensive Cancer Center, Northwestern University Feinberg School of Medicine, Chicago, IL 60611, USA * Correspondence: [email protected]; Tel.: +1-312-503-1040 These authors contributed equally to this work. y These authors contributed equally as senior authors. z Received: 5 July 2019; Accepted: 8 August 2019; Published: 10 August 2019 Abstract: The clinical outcomes of pancreatic ductal adenocarcinoma (PDAC) remain dismal, with an estimated five-year survival rate of less than 5%. Early detection and prognostic approaches, including robust biomarkers for PDAC, are critical for improving patient survival. Our goal was to explore the biomarker potential of 5-hydroxymethylcytosines (5hmC), an emerging epigenetic marker with a distinct role in cancer pathobiology, yet under-investigated, due largely to technical constraints relating to PDAC. The TET-assisted bisulfite (TAB)-Array assay represents state-of-the-art technology and was used to directly profile 5hmC at single-base resolution with the Illumina EPIC array (~850,000 cytosine modification sites) in 17 pairs of tumor/adjacent tissue samples from US patients collected at the University of Chicago Medical Center. -

Identification of the TCL1/MTCP1-Like 1 (TML1) Gene from the Region Next to the TCL1 Locus1
[CANCER RESEARCH 59, 2313–2317, May 15, 1999] Advances in Brief Identification of the TCL1/MTCP1-like 1 (TML1) Gene from the Region Next to the TCL1 Locus1 Jun Sugimoto, Toyomasa Hatakeyama, Maria Grazia Narducci, Giandomenico Russo, and Masaharu Isobe2 Department of Materials and Biosystem Engineering, Faculty of Engineering, Toyama University, Toyama City, 930-8555 Japan [J. S., T. H., M. I.]; and IDI-IRCCS, Research Laboratories, 00167 Rome, Italy [M. G. N., G. R.] Abstract malignant clonal expansion of T cells of patients with AT. We and others have cloned numerous breakpoints at 14q32.1 involved in The region on chromosome 14q32.1 is frequently involved in chromo- T-cell neoplasms (7–11). By placing these breakpoints on the map of somal translocations and inversions with one of the T-cell receptor loci in the region, we found that the breakpoints involve a chromosomal human T-cell leukemias and lymphomas. The breakpoints of the different ; rearrangements segregate into two clusters: inversion on the centromeric segment of 400 kb and cluster in two regions (12). The centromeric side and simple balanced translocations on the telomeric side. If the target region is mainly involved in inversions, whereas the telomeric region gene activated by these different types of chromosomal rearrangements is is involved in simple translocations. These two regions enclose a the same, the gene must reside between the two clusters of breakpoints in segment of ;160 kb. We postulated that, if the oncogene activated by a region of ;160 kb. By screening of a placenta cDNA library using these different rearrangements is the same, it must reside between genomic probes derived from the vicinity of TCL1 locus, we have identi- these two clusters of breakpoints. -

Transcriptome Analyses Reveal Differential Transcriptional Profiles
G C A T T A C G G C A T genes Article Transcriptome Analyses Reveal Differential Transcriptional Profiles in Early- and Late-Dividing Porcine Somatic Cell Nuclear Transfer Embryos Zhiguo Liu , Guangming Xiang, Kui Xu, Jingjing Che, Changjiang Xu, Kui Li, Bingyuan Wang * and Yulian Mu Institute of Animal Sciences, Chinese Academy of Agricultural Sciences, Beijing 100193, China; [email protected] (Z.L.); [email protected] (G.X.); [email protected] (K.X.); [email protected] (J.C.); [email protected] (C.X.); [email protected] (K.L.); [email protected] (Y.M.) * Correspondence: [email protected] Received: 20 October 2020; Accepted: 7 December 2020; Published: 12 December 2020 Abstract: Somatic cell nuclear transfer (SCNT) is not only a valuable tool for understanding nuclear reprogramming, but it also facilitates the generation of genetically modified animals. However, the development of SCNT embryos has remained an uncontrollable process. It was reported that the SCNT embryos that complete the first cell division sooner are more likely to develop to the blastocyst stage, suggesting their better developmental competence. Therefore, to better understand the underlying molecular mechanisms, RNA-seq of pig SCNT embryos that were early-dividing (24 h postactivation) and late-dividing (36 h postactivation) was performed. Our analysis revealed that early- and late-dividing embryos have distinct RNA profiles, and, in all, 3077 genes were differentially expressed. Gene ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) analyses revealed that early-dividing embryos exhibited higher expression in genes that participated in the meiotic cell cycle, while enrichment of RNA processing- and translation-related genes was found in late-dividing embryos. -

Table S1. 103 Ferroptosis-Related Genes Retrieved from the Genecards
Table S1. 103 ferroptosis-related genes retrieved from the GeneCards. Gene Symbol Description Category GPX4 Glutathione Peroxidase 4 Protein Coding AIFM2 Apoptosis Inducing Factor Mitochondria Associated 2 Protein Coding TP53 Tumor Protein P53 Protein Coding ACSL4 Acyl-CoA Synthetase Long Chain Family Member 4 Protein Coding SLC7A11 Solute Carrier Family 7 Member 11 Protein Coding VDAC2 Voltage Dependent Anion Channel 2 Protein Coding VDAC3 Voltage Dependent Anion Channel 3 Protein Coding ATG5 Autophagy Related 5 Protein Coding ATG7 Autophagy Related 7 Protein Coding NCOA4 Nuclear Receptor Coactivator 4 Protein Coding HMOX1 Heme Oxygenase 1 Protein Coding SLC3A2 Solute Carrier Family 3 Member 2 Protein Coding ALOX15 Arachidonate 15-Lipoxygenase Protein Coding BECN1 Beclin 1 Protein Coding PRKAA1 Protein Kinase AMP-Activated Catalytic Subunit Alpha 1 Protein Coding SAT1 Spermidine/Spermine N1-Acetyltransferase 1 Protein Coding NF2 Neurofibromin 2 Protein Coding YAP1 Yes1 Associated Transcriptional Regulator Protein Coding FTH1 Ferritin Heavy Chain 1 Protein Coding TF Transferrin Protein Coding TFRC Transferrin Receptor Protein Coding FTL Ferritin Light Chain Protein Coding CYBB Cytochrome B-245 Beta Chain Protein Coding GSS Glutathione Synthetase Protein Coding CP Ceruloplasmin Protein Coding PRNP Prion Protein Protein Coding SLC11A2 Solute Carrier Family 11 Member 2 Protein Coding SLC40A1 Solute Carrier Family 40 Member 1 Protein Coding STEAP3 STEAP3 Metalloreductase Protein Coding ACSL1 Acyl-CoA Synthetase Long Chain Family Member 1 Protein -

Monoclonal Antibody to TCL1A / TCL1
AM26558BT-N OriGene Technologies Inc. OriGene EU Acris Antibodies GmbH 9620 Medical Center Drive, Ste 200 Schillerstr. 5 Rockville, MD 20850 32052 Herford UNITED STATES GERMANY Phone: +1-888-267-4436 Phone: +49-5221-34606-0 Fax: +1-301-340-8606 Fax: +49-5221-34606-11 [email protected] [email protected] Monoclonal Antibody to TCL1A / TCL1 - Biotin Alternate names: T-cell leukemia/lymphoma protein 1A, TCL-1 Catalog No.: AM26558BT-N Quantity: 0.1 ml Concentration: 0.5 mg/ml Background: TCL1 (T cell leukemia/lymphoma 1), MTCP1 (mature T cell proliferation 1) and TCL1b belong to the TCL1 proto-oncogene family. The TCL1 gene at chromosome 14q32.1 is co mmonly activated in T cell neoplasms by chromosome r earrangements such as inversions inv(14)(q11;q32) and translocations t(14;14)(q11;q32) or t(7;14)(q35;q32). TCL1 expression is limited to immature thymocytes in early T-cell progenitors and from pre-B to mature B cells in the B-cell lineage. The TCL1/MTCP1/TCL1b proto-oncogene activation is the hallmark of human T-cell prolymphocytic leukemia (T-PLL), a form of adult leukemia. In addition to T- PLL, TCL1 is overexpressed in Burkitt's lymphoma cell lines, the majority of AIDS-related non-Hodgkin’s lymphoma-designated immunoblastic lymphoma plasmacytoids, lymphoblastic lymphoma, chronic lymphocytic leukemia, mantle cell lymphoma, follicular lymphoma, diffuse large B-cell lymphoma, and primary cutaneous B-cell lymphoma. It is also implicated in the development of hematopoietic abnormalities in patients with ataxia- telangiectasia. The members of the TCL1 proto-oncogene family bind to the serine/threonine kinase Akt (PKB), increasing its phosphorylation status and kinase activity. -
TCL1 (1-21) Mouse Mab A
Revision 1 C 0 2 - t TCL1 (1-21) Mouse mAb a e r o t S Orders: 877-616-CELL (2355) [email protected] Support: 877-678-TECH (8324) 6 1 Web: [email protected] 0 www.cellsignal.com 4 # 3 Trask Lane Danvers Massachusetts 01923 USA For Research Use Only. Not For Use In Diagnostic Procedures. Applications: Reactivity: Sensitivity: MW (kDa): Source/Isotype: UniProt ID: Entrez-Gene Id: WB, IP H Endogenous 14 Mouse IgG2b P56279 8115 Product Usage Information Application Dilution Western Blotting 1:1000 Immunoprecipitation 1:50 Storage Supplied in 10 mM sodium HEPES (pH 7.5), 150 mM NaCl, 100 µg/ml BSA, 50% glycerol and less than 0.02% sodium azide. Store at –20°C. Do not aliquot the antibody. Specificity / Sensitivity TCL1 (1-21) Mouse mAb detects endogenous levels of total TCL1 protein. Species Reactivity: Human Species predicted to react based on 100% sequence homology: Monkey Source / Purification Monoclonal antibody is produced by immunizing animals with TCL1. Background TCL1 (T cell leukemia 1), MTCP1 and TCL1b belong to the TCL1 proto-oncogene family, and their products are involved in Akt activation during embryonic development, T cell leukemias, prolymphocytic leukemias and B cell lymphomas (1-3). The Akt association domain of TCL1 binds with the PH domain of Akt. The formation of an oligomeric TCL- Akt complex is required for TCL1 coactivator function and results in phosphorylation and activation of Akt . Furthermore, functional analysis indicates that the interaction between TCL1 and Akt promotes translocation of Akt to the nucleus (4-6). These findings are supported by the crystal structure of TCL1, which suggests that TCL1 may participate in molecular transport (7). -

Abstracts from the 53Rd European Society of Human Genetics (ESHG) Conference: E-Posters
European Journal of Human Genetics (2020) 28:798–1016 https://doi.org/10.1038/s41431-020-00741-5 ABSTRACTS COLLECTION Abstracts from the 53rd European Society of Human Genetics (ESHG) Conference: e-Posters © European Society of Human Genetics 2020. Modified from the conference website and published with permission 2020 Volume 28 | Supplement 1 Virtual Conference June 6-9, 2020 Sponsorship: Publication of this supplement was sponsored by the European Society of Human Genetics. All content was reviewed and approved by the ESHG Scientific Programme Committee, which held full responsibility for the abstract selections. 1234567890();,: 1234567890();,: Disclosure Information: In order to help readers form their own judgments of potential bias in published abstracts, authors are asked to declare any competing financial interests. Contributions of up to EUR 10 000.- (Ten thousand Euros, or equivalent value in kind) per year per company are considered “Modest”. Contributions above EUR 10 000.- per year are considered “Significant”. Presenting authors are indicated with bold typeface in the contributor lists. e-Posters improve the prevention and treatment of congenital malformations. E-P01 Reproductive Genetics/Prenatal Genetics Material and methods: The data from Medical Genetic Center Register was analyzing by the MedCalc 15.8 E-P01.07 program. Epidemiological monitoring of congenital malforma- Results: The incidence rate of congenital malformations tions in Yakutia from 2007 to 2018 over a 12-year period averaged 29.4 cases per 1000 newborns with a standard deviation of 2.9. It’s high A. I. Fedorov1, A. L. Sukhomyasova1,2, A. N. Sleptsov2,1,N. indicator In Russia (>20 ‰). At this period, the average R.