Monoclonal Antibody to TCL1A / TCL1
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-
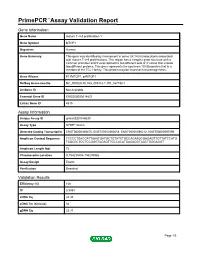
Primepcr™Assay Validation Report
PrimePCR™Assay Validation Report Gene Information Gene Name mature T-cell proliferation 1 Gene Symbol MTCP1 Organism Human Gene Summary This gene was identified by involvement in some t(X;14) translocations associated with mature T-cell proliferations. This region has a complex gene structure with a common promoter and 5' exon spliced to two different sets of 3' exons that encode two different proteins. This gene represents the upstream 13 kDa protein that is a member of the TCL1 family. This protein may be involved in leukemogenesis. Gene Aliases P13MTCP1, p8MTCP1 RefSeq Accession No. NC_000023.10, NG_005114.1, NT_167198.1 UniGene ID Not Available Ensembl Gene ID ENSG00000214827 Entrez Gene ID 4515 Assay Information Unique Assay ID qHsaCED0048630 Assay Type SYBR® Green Detected Coding Transcript(s) ENST00000369476, ENST00000362018, ENST00000596212, ENST00000597096 Amplicon Context Sequence TCCCCTGACCATTAAATGATGCTGTATCTGCCACAAGCGAGAGTTGTTATCCATG TAGCGCTCCTCCGGGTAGAGTTGCCACATGAGAGGTAGCTGGGAGGT Amplicon Length (bp) 72 Chromosome Location X:154293804-154293986 Assay Design Exonic Purification Desalted Validation Results Efficiency (%) 106 R2 0.9994 cDNA Cq 24.34 cDNA Tm (Celsius) 82 gDNA Cq 25.37 Page 1/5 PrimePCR™Assay Validation Report Specificity (%) 100 Information to assist with data interpretation is provided at the end of this report. Page 2/5 PrimePCR™Assay Validation Report MTCP1, Human Amplification Plot Amplification of cDNA generated from 25 ng of universal reference RNA Melt Peak Melt curve analysis of above amplification Standard -

Structural Studies of the Complex Between Akt-In and the Akt2-PH Domain Suggest That the Peptide Acts As an Allosteric Inhibitor of the Akt Kinase
The Open Spectroscopy Journal, 2009, 3, 65-76 65 Open Access Structural Studies of the Complex Between Akt-in and the Akt2-PH Domain Suggest that the Peptide Acts as an Allosteric Inhibitor of the Akt Kinase Virginie Ropars1,2,3, Philippe Barthe1,2,3, Chi-Shien Wang4, Wenlung Chen4, Der-Lii M. Tzou4,5, Anne Descours6, Loïc Martin6, Masayuki Noguchi7, Daniel Auguin1,2,3,,¶ and Christian Roumestand*,1,2,3 1CNRS UMR 5048, Centre de Biochimie Structurale, Montpellier, France 2INSERM U554, Montpellier, France 3Université Montpellier I et II, Montpellier, France 4Department of Applied Chemistry, National Chiayi University, Chiayi 60004, Taiwan, ROC 5Institute of Chemistry, Academia Sinica, Nankang, Taipei 11529, Taiwan, ROC 6CEA, iBiTecs, Service d’Ingénierie Moléculaire des Protéines, 91191 Gif sur Yvette, France 7Division of Cancer Biology, Institute for Genetic Medicine, Hokkaido University, N15 W7, Kita-ku, Sapporo 060-0815, Japan Abstract: Serine/threonine kinase Akt plays a central role in the regulation of cell survival and proliferation. Hence, the search for Akt specific inhibitors constitutes an attractive strategy for anticancer therapy. We have previously demonstrated that the proto-oncogene TCL1 coactivates Akt upon binding to its Plekstrin Homology Domain, and we proposed a model for the structure of the complex TCL1:Akt2-PHD. This model led to the rational design of Akt-in, a peptide inhibitor spanning the A ß-strand of human TCL1 that binds Akt2 PH domain and inhibits the kinase activation. In the present report, we used NMR spectroscopy to determine the 3D structure of the peptide free in solution and bound to Akt2-PHD. -

Molecular Dynamics and Evolutionary Aspects of the Transition from the Fully Grown Oocyte to Embryo
Downloaded from genesdev.cshlp.org on September 26, 2021 - Published by Cold Spring Harbor Laboratory Press Cracking the egg: molecular dynamics and evolutionary aspects of the transition from the fully grown oocyte to embryo Alexei V. Evsikov,1,5 Joel H. Graber,1 J. Michael Brockman,1,2 Aleš Hampl,3 Andrea E. Holbrook,1 Priyam Singh,1,2 John J. Eppig,1 Davor Solter,1,4 and Barbara B. Knowles1 1The Jackson Laboratory, Bar Harbor, Maine 04609, USA; 2 Bioinformatics Program, Boston University, Boston, Massachusetts 02215, USA; 3Masaryk University Brno and Institute of Experimental Medicine, 625 00 Brno, Czech Republic; 4Max Planck Institute of Immunobiology, 79108 Freiburg, Germany Fully grown oocytes (FGOs) contain all the necessary transcripts to activate molecular pathways underlying the oocyte-to-embryo transition (OET). To elucidate this critical period of development, an extensive survey of the FGO transcriptome was performed by analyzing 19,000 expressed sequence tags of the Mus musculus FGO cDNA library. Expression of 5400 genes and transposable elements is reported. For a majority of genes expressed in mouse FGOs, homologs transcribed in eggs of Xenopus laevis or Ciona intestinalis were found, pinpointing evolutionary conservation of most regulatory cascades underlying the OET in chordates. A large proportion of identified genes belongs to several gene families with oocyte-restricted expression, a likely result of lineage-specific genomic duplications. Gene loss by mutation and expression in female germline of retrotransposed genes specific to M. musculus is documented. These findings indicate rapid diversification of genes involved in female reproduction. Comparison of the FGO and two-cell embryo transcriptomes demarcated the processes important for oogenesis from those involved in OET and identified novel motifs in maternal mRNAs associated with transcript stability. -

Leukaemia Section
Atlas of Genetics and Cytogenetics in Oncology and Haematology OPEN ACCESS JOURNAL INIST-CNRS Leukaemia Section Short Communication t(7;14)(q35;q32.1) TRB/TCL1A inv(14)(q11q32.1) TRA-TRD/TCL1A t(14;14)(q11;q32.1) TRA-TRD/TCL1A Tatiana Gindina Raisa Gorbacheva Memorial Institute of Children's Oncology, Hematology and Transplantation at First Pavlov St. Petersburg State Medical University, Saint-Petersburg, Russia Published in Atlas Database: June 2018 Online updated version : http://AtlasGeneticsOncology.org/Anomalies/inv14ID2049.html Printable original version : http://documents.irevues.inist.fr/bitstream/handle/2042/70184/06-2018-inv14ID2049.pdf DOI: 10.4267/2042/70184 This article is an update of : Boyer J. t(7;14)(q35;q32.1) TRB@/TCL1A. Atlas Genet Cytogenet Oncol Haematol 2001;5(3) This work is licensed under a Creative Commons Attribution-Noncommercial-No Derivative Works 2.0 France Licence. © 2019 Atlas of Genetics and Cytogenetics in Oncology and Haematology TRD; B lymphoblastic leukaemia/lymphoma; T Abstract lymphoblastic leukaemia/lymphoma; Adult T-cell Review on t(7;14)(q35;q32), inv(14)(q11q32) and leukemia/lymphoma; T-cell prolymphocytic t(14;14)(q11;q32), with data on clinics, and the leukemia; Angioimmunoblastic T-cell lymphoma; genes involved. Chronic lymphocytic leukemia; syndrome; Mycosis Keywords fungoides; Hepatosplenic T-cell lymphoma; Acute Chromosome 7; Chromosome 14; TCL1A; TRA; myeloid leukaemia; Ataxia telangiectasia. Left: inv(14)(q11q32)and i(8q), G- banding - Courtesy Jean Luc Lai; right: inv(14)(q11q32) and t(14)(14) with i(7q), G- banding - Courtesy Tatiana Gindina. Atlas Genet Cytogenet Oncol Haematol. 2019; 23(4) 90 t(7;14)(q35;q32.1) TRB/TCL1A Gindina T inv(14)(q11q32.1) TRA-TRD/TCL1A t(14;14)(q11;q32.1) TRA-TRD/TCL1A Cytogenetics Clinics and pathology Chromosomal abnormalities are detected in most T- Disease PLL after culture with mitogens like PHA. -

Association of Gene Ontology Categories with Decay Rate for Hepg2 Experiments These Tables Show Details for All Gene Ontology Categories
Supplementary Table 1: Association of Gene Ontology Categories with Decay Rate for HepG2 Experiments These tables show details for all Gene Ontology categories. Inferences for manual classification scheme shown at the bottom. Those categories used in Figure 1A are highlighted in bold. Standard Deviations are shown in parentheses. P-values less than 1E-20 are indicated with a "0". Rate r (hour^-1) Half-life < 2hr. Decay % GO Number Category Name Probe Sets Group Non-Group Distribution p-value In-Group Non-Group Representation p-value GO:0006350 transcription 1523 0.221 (0.009) 0.127 (0.002) FASTER 0 13.1 (0.4) 4.5 (0.1) OVER 0 GO:0006351 transcription, DNA-dependent 1498 0.220 (0.009) 0.127 (0.002) FASTER 0 13.0 (0.4) 4.5 (0.1) OVER 0 GO:0006355 regulation of transcription, DNA-dependent 1163 0.230 (0.011) 0.128 (0.002) FASTER 5.00E-21 14.2 (0.5) 4.6 (0.1) OVER 0 GO:0006366 transcription from Pol II promoter 845 0.225 (0.012) 0.130 (0.002) FASTER 1.88E-14 13.0 (0.5) 4.8 (0.1) OVER 0 GO:0006139 nucleobase, nucleoside, nucleotide and nucleic acid metabolism3004 0.173 (0.006) 0.127 (0.002) FASTER 1.28E-12 8.4 (0.2) 4.5 (0.1) OVER 0 GO:0006357 regulation of transcription from Pol II promoter 487 0.231 (0.016) 0.132 (0.002) FASTER 6.05E-10 13.5 (0.6) 4.9 (0.1) OVER 0 GO:0008283 cell proliferation 625 0.189 (0.014) 0.132 (0.002) FASTER 1.95E-05 10.1 (0.6) 5.0 (0.1) OVER 1.50E-20 GO:0006513 monoubiquitination 36 0.305 (0.049) 0.134 (0.002) FASTER 2.69E-04 25.4 (4.4) 5.1 (0.1) OVER 2.04E-06 GO:0007050 cell cycle arrest 57 0.311 (0.054) 0.133 (0.002) -

TCL1A Expression Delineates Biological and Clinical Variability in B-Cell Lymphoma
Modern Pathology (2009) 22, 206–215 & 2009 USCAP, Inc All rights reserved 0893-3952/09 $32.00 www.modernpathology.org TCL1A expression delineates biological and clinical variability in B-cell lymphoma Mohit Aggarwal1,4, Raquel Villuendas1, Gonzalo Gomez2, Socorro M Rodriguez-Pinilla1, Margarita Sanchez-Beato1, David Alvarez1, Nerea Martinez1, Antonia Rodriguez1, Maria E Castillo1, Francisca I Camacho1, Santiago Montes-Moreno1, Jose A Garcia-Marco3, Eva Kimby4, David G Pisano2 and Miguel A Piris1 1Molecular Pathology Programme, Spanish National Cancer Research Centre (CNIO), Madrid, Spain; 2Bioinformatics Unit, Structural Biology and Biocomputing Programme, CNIO, Madrid, Spain; 3Department of Haematology, Hospital Universitario Puerta de Hierro, Madrid, Spain and 4Division of Hematology, Department of Internal Medicine at Huddinge, Karolinska Institutet, Stockholm, Sweden The assembly of a collection of gene-expression signatures of the major types of B-cell non-Hodgkin’s lymphoma has identified increased T-cell leukemia/lymphoma 1A (TCL1) expression in multiple lymphoma types and cases, and has enabled the investigation of the functional and clinical importance of TCL1 expression. Specifically, Burkitt’s lymphoma cases show a homogeneously strong expression of TCL1, whereas diffuse large B-cell lymphoma, follicular lymphoma, mantle cell lymphoma, chronic lymphocytic leukemia, nodal marginal zone lymphoma, and splenic marginal zone lymphoma display a striking variability in the intensity of TCL1 staining. This was validated in two independent series. A Gene-Set Enrichment Analysis of the genes correlated with TCL1A expression found that variation in the level of expression of TCL1A was significantly associated with some of the most important gene signatures recognizing B-cell lymphoma pathogenesis and heterogeneity, such as germinal center, B-cell receptor, NF-jB (and its target genes), death, MAP kinases, TNFR1, TOLL, and IL1R. -

Regulation of the Akt Kinase by Interacting Proteins
Oncogene (2005) 24, 7401–7409 & 2005 Nature Publishing Group All rights reserved 0950-9232/05 $30.00 www.nature.com/onc Regulation of the Akt kinase by interacting proteins Keyong Du1 and Philip N Tsichlis*,1 1Molecular Oncology Research Institute, Tufts-New England Medical Center, Boston, MA 02111, USA Ten years ago, it was observed that the Akt kinase revolutionized our understanding of cell function at is activated by phosphorylation via a phosphoinositide the molecular level.These technologies also allowed 3-kinase (PI-3K)-dependent process. This discovery gene- the identification of a large array of protein–protein rated enormous interest because it provided a link between interaction motifs that facilitate our understanding of PI-3K, an enzyme known to play a critical role in cellular signal transduction by providing clues on the potential physiology, and its downstream targets. Subsequently, it function of novel signaling proteins (Barnouin, 2004; was shown that the activity of the core components of the Miller and Stagljar, 2004). ‘PI-3K/Akt pathway’ is modulated by a complex network Akt1, also known as protein kinase Ba (PKBa) of regulatory proteins and pathways. Some of the Akt- (Bellacosa et al., 1991; Coffer and Woodgett, 1991; binding partners modulate its activation by external Jones et al., 1991), is the founding member of a protein signals by interacting with different domains of the Akt kinase family composed of three members, Akt1, Akt2, protein. This review focuses on the Akt interacting and Akt3.Akt family members regulate a diverse proteins and the mechanisms by which they regulate Akt array of cellular functions, including apoptosis, cellular activation. -

The Cytogenetics of Hematologic Neoplasms 1 5
The Cytogenetics of Hematologic Neoplasms 1 5 Aurelia Meloni-Ehrig that errors during cell division were the basis for neoplastic Introduction growth was most likely the determining factor that inspired early researchers to take a better look at the genetics of the The knowledge that cancer is a malignant form of uncon- cell itself. Thus, the need to have cell preparations good trolled growth has existed for over a century. Several biologi- enough to be able to understand the mechanism of cell cal, chemical, and physical agents have been implicated in division became of critical importance. cancer causation. However, the mechanisms responsible for About 50 years after Boveri’s chromosome theory, the this uninhibited proliferation, following the initial insult(s), fi rst manuscripts on the chromosome makeup in normal are still object of intense investigation. human cells and in genetic disorders started to appear, fol- The fi rst documented studies of cancer were performed lowed by those describing chromosome changes in neoplas- over a century ago on domestic animals. At that time, the tic cells. A milestone of this investigation occurred in 1960 lack of both theoretical and technological knowledge with the publication of the fi rst article by Nowell and impaired the formulations of conclusions about cancer, other Hungerford on the association of chronic myelogenous leu- than the visible presence of new growth, thus the term neo- kemia with a small size chromosome, known today as the plasm (from the Greek neo = new and plasma = growth). In Philadelphia (Ph) chromosome, to honor the city where it the early 1900s, the fundamental role of chromosomes in was discovered (see also Chap. -

Biological Pathways, Candidate Genes, and Molecular Markers Associated with Quality-Of-Life Domains: an Update
Qual Life Res (2014) 23:1997–2013 DOI 10.1007/s11136-014-0656-1 REVIEW Biological pathways, candidate genes, and molecular markers associated with quality-of-life domains: an update Mirjam A. G. Sprangers • Melissa S. Y. Thong • Meike Bartels • Andrea Barsevick • Juan Ordon˜ana • Qiuling Shi • Xin Shelley Wang • Pa˚l Klepstad • Eddy A. Wierenga • Jasvinder A. Singh • Jeff A. Sloan Accepted: 19 February 2014 / Published online: 7 March 2014 Ó Springer International Publishing Switzerland 2014 Abstract (depressed mood) and positive (well-being/happiness) Background There is compelling evidence of a genetic emotional functioning, social functioning, and overall foundation of patient-reported quality of life (QOL). Given QOL. the rapid development of substantial scientific advances in Methods We followed a purposeful search algorithm of this area of research, the current paper updates and extends existing literature to capture empirical papers investigating reviews published in 2010. the relationship between biological pathways and molecu- Objectives The objective was to provide an updated lar markers and the identified QOL domains. overview of the biological pathways, candidate genes, and Results Multiple major pathways are involved in each molecular markers involved in fatigue, pain, negative QOL domain. The inflammatory pathway has the strongest evidence as a controlling mechanism underlying fatigue. Inflammation and neurotransmission are key processes On behalf of the GeneQol Consortium. involved in pain perception, and the catechol-O-methyl- transferase (COMT) gene is associated with multiple sorts Electronic supplementary material The online version of this article (doi:10.1007/s11136-014-0656-1) contains supplementary of pain. The neurotransmitter and neuroplasticity theories material, which is available to authorized users. -

Tcl1 Enhances Akt Kinase Activity and Mediates Its Nuclear Translocation
Tcl1 enhances Akt kinase activity and mediates its nuclear translocation Yuri Pekarsky*, Anatoliy Koval*, Cora Hallas*, Roberta Bichi*, Maria Tresini*, Scott Malstrom*, Giandomenico Russo†, Philip Tsichlis*, and Carlo M. Croce*‡ *Kimmel Cancer Center, Jefferson Medical College, Philadelphia, PA 19107; and †Istituto Dermopatico dell’Immacolata, Laboratory of Vascular Pathology, 00167 Rome, Italy Contributed by Carlo M. Croce, December 20, 1999 The TCL1 oncogene at 14q32.1 is involved in the development of wortmannin, a (PI-3K)-kinase inhibitor, completely inhibits the human mature T-cell leukemia. The mechanism of action of Tcl1 is activation of Akt (reviewed in ref. 9). Recent studies showed that unknown. Because the virus containing the v-akt oncogene causes Akt is a key player in the transduction of antiapoptotic and T-cell lymphoma in mice and Akt is a key player in transduction of proliferative signals in T-cells (refs. 10 and 11; reviewed in ref. antiapoptotic and proliferative signals in T-cells, we investigated 9). Activated Akt enhances both cell cycle progression and IL2 whether Akt and Tcl1 function in the same pathway. Coimmuno- production through the inhibition by phosphorylation of the precipitation experiments showed that endogenous Akt1 and Tcl1 proapoptotic factor Bad (11). Introduction of a constitutively physically interact in the T-cell leukemia cell line SupT11; both activated AKT1 transgene under the control of the proximal lck proteins also interact when cotransfected into 293 cells. Using promoter causes T-cell lymphomas in mice (S.M. and P.T., several AKT1 constructs in cotransfection experiments, we deter- unpublished data). In cultured cells, Akt1 can be localized in mined that this interaction occurs through the pleckstrin homology both the nucleus and cytoplasm (12). -

Goat Anti-TCL1A Antibody Peptide-Affinity Purified Goat Antibody Catalog # Af2075a
10320 Camino Santa Fe, Suite G San Diego, CA 92121 Tel: 858.875.1900 Fax: 858.622.0609 Goat Anti-TCL1A Antibody Peptide-affinity purified goat antibody Catalog # AF2075a Specification Goat Anti-TCL1A Antibody - Product Information Application WB Primary Accession P56279 Other Accession NP_068801, 8115 Reactivity Human Host Goat Clonality Polyclonal Concentration 100ug/200ul Isotype IgG Calculated MW 13460 Goat Anti-TCL1A Antibody - Additional Information AF2075a (0.1 µg/ml) staining of Jurkat Cell Gene ID 8115 lysate (35 µg protein in RIPA buffer). Primary incubation was 1 hour. Detected by Other Names chemiluminescence. T-cell leukemia/lymphoma protein 1A, Oncogene TCL-1, Oncogene TCL1, Protein p14 TCL1, TCL1A, TCL1 Goat Anti-TCL1A Antibody - Background Format Overexpression of the TCL1 gene in humans 0.5 mg IgG/ml in Tris saline (20mM Tris has been implicated in the development of pH7.3, 150mM NaCl), 0.02% sodium azide, mature T cell leukemia, in which chromosomal with 0.5% bovine serum albumin rearrangements bring the TCL1 gene in close proximity to the T-cell antigen receptor Storage (TCR)-alpha (MIM 186880) or TCR-beta (MIM Maintain refrigerated at 2-8°C for up to 6 186930) regulatory elements (summarized by months. For long term storage store at -20°C in small aliquots to prevent Virgilio et al., 1998 [PubMed 9520462]). In freeze-thaw cycles. normal T cells TCL1 is expressed in CD4-/CD8- cells, but not in cells at later stages of Precautions differentiation. TCL1 functions as a coactivator Goat Anti-TCL1A Antibody is for research of the cell survival kinase AKT (MIM 164730) use only and not for use in diagnostic or (Laine et al., 2000 [PubMed 10983986]). -

Highly Dampened Blood Transcriptome Response in HIV Patients After Respiratory Infection Subhashini A
www.nature.com/scientificreports OPEN Highly dampened blood transcriptome response in HIV patients after respiratory infection Subhashini A. Sellers1, William A. Fischer II1, Mark T. Heise2,3,4 & Klaus Schughart 5,6,7* Respiratory viral (RV) infections represent a major threat for human health worldwide. Persons with HIV (PWH) have a compromised immune response and are thought to be at higher risk for severe RV disease. However, very little is known about the host immune response to RV infection in PWH. Here, we investigated gene expression changes in the peripheral blood of PWH co-infected with RV. Only very few diferentially expressed genes could be detected between PWH with and without RV infection, suggesting that the immune response to RV in PWH is strongly dampened. Our data provides important insights into the host response to RV infections in HIV patients. Respiratory viral (RV) infections, including infuenza and presently SARS-CoV-2 pose major threats to pub- lic health as they are responsible for epidemics and pandemics resulting in high morbidity and mortality worldwide1,2. Te most severe infuenza human pandemic in 1918 resulted in about 30 million fatal casualties 3 and the current coronavirus pandemic has already resulted in over 700,000 deaths (WHO situation report 2020). Seasonal RV infections also represent a major health hazard causing morbidity, mortality, and enormous loss of the work force yearly1,4,5. Very little is known about the immune response in persons with HIV (PWH) afer RV infection. Symptoms of RV infection appear to be similar in PWH and non-HIV patients 6.