The Prevalence of Nucleic Acid Binding Among Reader Domains
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-
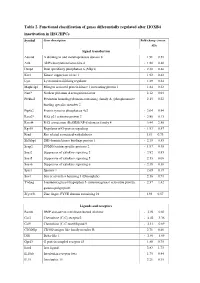
Table 2. Functional Classification of Genes Differentially Regulated After HOXB4 Inactivation in HSC/Hpcs
Table 2. Functional classification of genes differentially regulated after HOXB4 inactivation in HSC/HPCs Symbol Gene description Fold-change (mean ± SD) Signal transduction Adam8 A disintegrin and metalloprotease domain 8 1.91 ± 0.51 Arl4 ADP-ribosylation factor-like 4 - 1.80 ± 0.40 Dusp6 Dual specificity phosphatase 6 (Mkp3) - 2.30 ± 0.46 Ksr1 Kinase suppressor of ras 1 1.92 ± 0.42 Lyst Lysosomal trafficking regulator 1.89 ± 0.34 Mapk1ip1 Mitogen activated protein kinase 1 interacting protein 1 1.84 ± 0.22 Narf* Nuclear prelamin A recognition factor 2.12 ± 0.04 Plekha2 Pleckstrin homology domain-containing. family A. (phosphoinosite 2.15 ± 0.22 binding specific) member 2 Ptp4a2 Protein tyrosine phosphatase 4a2 - 2.04 ± 0.94 Rasa2* RAS p21 activator protein 2 - 2.80 ± 0.13 Rassf4 RAS association (RalGDS/AF-6) domain family 4 3.44 ± 2.56 Rgs18 Regulator of G-protein signaling - 1.93 ± 0.57 Rrad Ras-related associated with diabetes 1.81 ± 0.73 Sh3kbp1 SH3 domain kinase bindings protein 1 - 2.19 ± 0.53 Senp2 SUMO/sentrin specific protease 2 - 1.97 ± 0.49 Socs2 Suppressor of cytokine signaling 2 - 2.82 ± 0.85 Socs5 Suppressor of cytokine signaling 5 2.13 ± 0.08 Socs6 Suppressor of cytokine signaling 6 - 2.18 ± 0.38 Spry1 Sprouty 1 - 2.69 ± 0.19 Sos1 Son of sevenless homolog 1 (Drosophila) 2.16 ± 0.71 Ywhag 3-monooxygenase/tryptophan 5- monooxygenase activation protein. - 2.37 ± 1.42 gamma polypeptide Zfyve21 Zinc finger. FYVE domain containing 21 1.93 ± 0.57 Ligands and receptors Bambi BMP and activin membrane-bound inhibitor - 2.94 ± 0.62 -

Functional Roles of Bromodomain Proteins in Cancer
cancers Review Functional Roles of Bromodomain Proteins in Cancer Samuel P. Boyson 1,2, Cong Gao 3, Kathleen Quinn 2,3, Joseph Boyd 3, Hana Paculova 3 , Seth Frietze 3,4,* and Karen C. Glass 1,2,4,* 1 Department of Pharmaceutical Sciences, Albany College of Pharmacy and Health Sciences, Colchester, VT 05446, USA; [email protected] 2 Department of Pharmacology, Larner College of Medicine, University of Vermont, Burlington, VT 05405, USA; [email protected] 3 Department of Biomedical and Health Sciences, University of Vermont, Burlington, VT 05405, USA; [email protected] (C.G.); [email protected] (J.B.); [email protected] (H.P.) 4 University of Vermont Cancer Center, Burlington, VT 05405, USA * Correspondence: [email protected] (S.F.); [email protected] (K.C.G.) Simple Summary: This review provides an in depth analysis of the role of bromodomain-containing proteins in cancer development. As readers of acetylated lysine on nucleosomal histones, bromod- omain proteins are poised to activate gene expression, and often promote cancer progression. We examined changes in gene expression patterns that are observed in bromodomain-containing proteins and associated with specific cancer types. We also mapped the protein–protein interaction network for the human bromodomain-containing proteins, discuss the cellular roles of these epigenetic regu- lators as part of nine different functional groups, and identify bromodomain-specific mechanisms in cancer development. Lastly, we summarize emerging strategies to target bromodomain proteins in cancer therapy, including those that may be essential for overcoming resistance. Overall, this review provides a timely discussion of the different mechanisms of bromodomain-containing pro- Citation: Boyson, S.P.; Gao, C.; teins in cancer, and an updated assessment of their utility as a therapeutic target for a variety of Quinn, K.; Boyd, J.; Paculova, H.; cancer subtypes. -

Modes of Interaction of KMT2 Histone H3 Lysine 4 Methyltransferase/COMPASS Complexes with Chromatin
cells Review Modes of Interaction of KMT2 Histone H3 Lysine 4 Methyltransferase/COMPASS Complexes with Chromatin Agnieszka Bochy ´nska,Juliane Lüscher-Firzlaff and Bernhard Lüscher * ID Institute of Biochemistry and Molecular Biology, Medical School, RWTH Aachen University, Pauwelsstrasse 30, 52057 Aachen, Germany; [email protected] (A.B.); jluescher-fi[email protected] (J.L.-F.) * Correspondence: [email protected]; Tel.: +49-241-8088850; Fax: +49-241-8082427 Received: 18 January 2018; Accepted: 27 February 2018; Published: 2 March 2018 Abstract: Regulation of gene expression is achieved by sequence-specific transcriptional regulators, which convey the information that is contained in the sequence of DNA into RNA polymerase activity. This is achieved by the recruitment of transcriptional co-factors. One of the consequences of co-factor recruitment is the control of specific properties of nucleosomes, the basic units of chromatin, and their protein components, the core histones. The main principles are to regulate the position and the characteristics of nucleosomes. The latter includes modulating the composition of core histones and their variants that are integrated into nucleosomes, and the post-translational modification of these histones referred to as histone marks. One of these marks is the methylation of lysine 4 of the core histone H3 (H3K4). While mono-methylation of H3K4 (H3K4me1) is located preferentially at active enhancers, tri-methylation (H3K4me3) is a mark found at open and potentially active promoters. Thus, H3K4 methylation is typically associated with gene transcription. The class 2 lysine methyltransferases (KMTs) are the main enzymes that methylate H3K4. KMT2 enzymes function in complexes that contain a necessary core complex composed of WDR5, RBBP5, ASH2L, and DPY30, the so-called WRAD complex. -

Histone-Binding of DPF2 Mediates Its Repressive Role in Myeloid Differentiation
Histone-binding of DPF2 mediates its repressive role in myeloid differentiation Ferdinand M. Hubera,1, Sarah M. Greenblattb,1, Andrew M. Davenporta,1, Concepcion Martinezb,YeXub,LyP.Vuc, Stephen D. Nimerb,2, and André Hoelza,2 aDivision of Chemistry and Chemical Engineering, California Institute of Technology, Pasadena, CA 91125; bSylvester Comprehensive Cancer Center, University of Miami Miller School of Medicine, Miami, FL 33136; and cMolecular Pharmacology Program, Memorial Sloan Kettering Cancer Center, New York, NY 10065 Edited by Douglas C. Rees, Howard Hughes Medical Institute, California Institute of Technology, Pasadena, CA, and approved April 26, 2017 (received for review January 6, 2017) Double plant homeodomain finger 2 (DPF2) is a highly evolution- RUNX1 form a methylation-dependent repressive complex in arily conserved member of the d4 protein family that is ubiqui- AML, although it remains unclear whether the two proteins bind tously expressed in human tissues and was recently shown to each other directly or act concertedly as part of a larger complex. inhibit the myeloid differentiation of hematopoietic stem/progen- Here, we present the crystal structure of the human DPF2 itor and acute myelogenous leukemia cells. Here, we present the tandem PHD finger domain at a 1.6-Å resolution. We demon- crystal structure of the tandem plant homeodomain finger domain strate that the DPF2 tandem PHD finger domain binds acetylated of human DPF2 at 1.6-Å resolution. We show that DPF2 interacts H3 and H4 histone tails, identify the primary determinants of with the acetylated tails of both histones 3 and 4 via bipartite histone recognition, and confirm these interactions in vivo. -
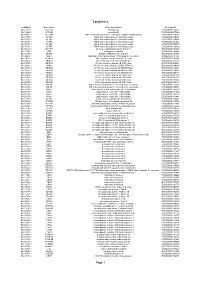
Epigenetics Page 1
Epigenetics esiRNA ID Gene Name Gene Description Ensembl ID HU-13237-1 ACTL6A actin-like 6A ENSG00000136518 HU-13925-1 ACTL6B actin-like 6B ENSG00000077080 HU-14457-1 ACTR1A ARP1 actin-related protein 1 homolog A, centractin alpha (yeast) ENSG00000138107 HU-10579-1 ACTR2 ARP2 actin-related protein 2 homolog (yeast) ENSG00000138071 HU-10837-1 ACTR3 ARP3 actin-related protein 3 homolog (yeast) ENSG00000115091 HU-09776-1 ACTR5 ARP5 actin-related protein 5 homolog (yeast) ENSG00000101442 HU-00773-1 ACTR6 ARP6 actin-related protein 6 homolog (yeast) ENSG00000075089 HU-07176-1 ACTR8 ARP8 actin-related protein 8 homolog (yeast) ENSG00000113812 HU-09411-1 AHCTF1 AT hook containing transcription factor 1 ENSG00000153207 HU-15150-1 AIRE autoimmune regulator ENSG00000160224 HU-12332-1 AKAP1 A kinase (PRKA) anchor protein 1 ENSG00000121057 HU-04065-1 ALG13 asparagine-linked glycosylation 13 homolog (S. cerevisiae) ENSG00000101901 HU-13552-1 ALKBH1 alkB, alkylation repair homolog 1 (E. coli) ENSG00000100601 HU-06662-1 ARID1A AT rich interactive domain 1A (SWI-like) ENSG00000117713 HU-12790-1 ARID1B AT rich interactive domain 1B (SWI1-like) ENSG00000049618 HU-09415-1 ARID2 AT rich interactive domain 2 (ARID, RFX-like) ENSG00000189079 HU-03890-1 ARID3A AT rich interactive domain 3A (BRIGHT-like) ENSG00000116017 HU-14677-1 ARID3B AT rich interactive domain 3B (BRIGHT-like) ENSG00000179361 HU-14203-1 ARID3C AT rich interactive domain 3C (BRIGHT-like) ENSG00000205143 HU-09104-1 ARID4A AT rich interactive domain 4A (RBP1-like) ENSG00000032219 HU-12512-1 ARID4B AT rich interactive domain 4B (RBP1-like) ENSG00000054267 HU-12520-1 ARID5A AT rich interactive domain 5A (MRF1-like) ENSG00000196843 HU-06595-1 ARID5B AT rich interactive domain 5B (MRF1-like) ENSG00000150347 HU-00556-1 ASF1A ASF1 anti-silencing function 1 homolog A (S. -

Aberrant Activity of Histone–Lysine N-Methyltransferase 2 (KMT2) Complexes in Oncogenesis
International Journal of Molecular Sciences Review Aberrant Activity of Histone–Lysine N-Methyltransferase 2 (KMT2) Complexes in Oncogenesis Elzbieta Poreba 1,* , Krzysztof Lesniewicz 2 and Julia Durzynska 1,* 1 Institute of Experimental Biology, Faculty of Biology, Adam Mickiewicz University, ul. Uniwersytetu Pozna´nskiego6, 61-614 Pozna´n,Poland 2 Department of Molecular and Cellular Biology, Institute of Molecular Biology and Biotechnology, Faculty of Biology, Adam Mickiewicz University, ul. Uniwersytetu Pozna´nskiego6, 61-614 Pozna´n,Poland; [email protected] * Correspondence: [email protected] (E.P.); [email protected] (J.D.); Tel.: +48-61-829-5857 (E.P.) Received: 19 November 2020; Accepted: 6 December 2020; Published: 8 December 2020 Abstract: KMT2 (histone-lysine N-methyltransferase subclass 2) complexes methylate lysine 4 on the histone H3 tail at gene promoters and gene enhancers and, thus, control the process of gene transcription. These complexes not only play an essential role in normal development but have also been described as involved in the aberrant growth of tissues. KMT2 mutations resulting from the rearrangements of the KMT2A (MLL1) gene at 11q23 are associated with pediatric mixed-lineage leukemias, and recent studies demonstrate that KMT2 genes are frequently mutated in many types of human cancers. Moreover, other components of the KMT2 complexes have been reported to contribute to oncogenesis. This review summarizes the recent advances in our knowledge of the role of KMT2 complexes in cell transformation. In addition, it discusses the therapeutic targeting of different components of the KMT2 complexes. Keywords: histone–lysine N-methyltransferase 2; COMPASS; COMPASS-like; H3K4 methylation; oncogenesis; cancer; epigenetics; chromatin 1. -

Pointing the Finger
RESEARCH HIGHLIGHTS Nature Reviews Molecular Cell Biology | AOP, published online 14 June 2006; doi:10.1038/nrm1966 DOI: 10.1038/nrm1966 URLs BPTF http://ca.expasy.org/uniprot/ Q9UIG2 ING2 http://ca.expasy.org/uniprot/ Q9ESK4 GENE EXPRESSION Pointing the finger Mechanistic links between specific histone-tail tumour suppressors, bound specifically to modifications and their effects on gene H3K4me3. ING2 is a subunit of a SIN3a–HDAC1 expression have been difficult to establish. histone-deacetylase complex. The authors However, four papers in Nature now identify the showed that in response to DNA damage, binding plant homeodomain (PHD) finger as an important of the ING2 PHD finger to H3K4me3 that is effector domain that binds to the trimethylated present at the promoters of actively transcribed K4 residue of histone H3 (H3K4me3) and couples proliferation genes enhanced the association of it to gene activation in one case and to gene the ING2–HDAC1 complex at these genes. This repression in another. resulted in increased histone-deacetylase activity Wysocka et al. affinity purified the BPTF and hence acute repression of the cognate (bromodomain and PHD finger transcription transcript. These findings, together with those of factor) subunit of NURF — an ATP-dependent Wysocka et al., indicate that PHD fingers have a chromatin remodelling complex — using an general role as effector domains that link H3K4me3-containing peptide. Mutational analysis H3K4me3 to diverse biological outcomes. narrowed the H3K4me3-interaction region to the The molecular mechanism that underlies the second, conserved bromodomain-proximal PHD recognition of H3K4me3 by the PHD finger of finger of BPTF. Knockdown of the histone ING2 was reported in a fourth linked paper. -

Chromatin Condensation and Recruitment of PHD Finger Proteins
6102–6112 Nucleic Acids Research, 2016, Vol. 44, No. 13 Published online 25 March 2016 doi: 10.1093/nar/gkw193 Chromatin condensation and recruitment of PHD finger proteins to histone H3K4me3 are mutually exclusive Jovylyn Gatchalian1,†, Carmen Mora Gallardo2,†, Stephen A. Shinsky3, Ruben Rosas Ospina1, Andrea Mansilla Liendo2, Krzysztof Krajewski3, Brianna J. Klein1, Forest H. Andrews1, Brian D. Strahl3, Karel H. M. van Wely2,* and Tatiana G. Kutateladze1,* 1Department of Pharmacology, University of Colorado School of Medicine, Aurora, CO 80045, USA, 2Department of Immunology and Oncology, Centro Nacional de Biotecnolog´ıa/CSIC, 28049 Madrid, Spain and 3Department of Biochemistry & Biophysics, The University of North Carolina School of Medicine, Chapel Hill, NC 27599, USA Received January 06, 2016; Revised March 12, 2016; Accepted March 15, 2016 ABSTRACT modifications (PTMs), including acetylation and methyla- tion of lysine, methylation of arginine, and phosphorylation Histone post-translational modifications, and spe- of serine and threonine. Over 500 histone PTMs have been cific combinations they create, mediate a wide range identified by mass spectrometry analysis, and a number of of nuclear events. However, the mechanistic bases these modifications, or epigenetic marks, have been shown for recognition of these combinations have not been to modulate chromatin structure and cell cycle specific pro- elucidated. Here, we characterize crosstalk between cesses (1). H3T3 and H3T6 phosphorylation, occurring in mito- Methylation of histone H3 at lysine 4 is one of the canon- sis, and H3K4me3, a mark associated with active ical PTMs (2–4). The trimethylated species (H3K4me3), transcription. We detail the molecular mechanisms which is found primarily in active promoters, is impli- by which H3T3ph/K4me3/T6ph switches mediate ac- cated in transcriptional regulation (5). -

Supp Table 6.Pdf
Supplementary Table 6. Processes associated to the 2037 SCL candidate target genes ID Symbol Entrez Gene Name Process NM_178114 AMIGO2 adhesion molecule with Ig-like domain 2 adhesion NM_033474 ARVCF armadillo repeat gene deletes in velocardiofacial syndrome adhesion NM_027060 BTBD9 BTB (POZ) domain containing 9 adhesion NM_001039149 CD226 CD226 molecule adhesion NM_010581 CD47 CD47 molecule adhesion NM_023370 CDH23 cadherin-like 23 adhesion NM_207298 CERCAM cerebral endothelial cell adhesion molecule adhesion NM_021719 CLDN15 claudin 15 adhesion NM_009902 CLDN3 claudin 3 adhesion NM_008779 CNTN3 contactin 3 (plasmacytoma associated) adhesion NM_015734 COL5A1 collagen, type V, alpha 1 adhesion NM_007803 CTTN cortactin adhesion NM_009142 CX3CL1 chemokine (C-X3-C motif) ligand 1 adhesion NM_031174 DSCAM Down syndrome cell adhesion molecule adhesion NM_145158 EMILIN2 elastin microfibril interfacer 2 adhesion NM_001081286 FAT1 FAT tumor suppressor homolog 1 (Drosophila) adhesion NM_001080814 FAT3 FAT tumor suppressor homolog 3 (Drosophila) adhesion NM_153795 FERMT3 fermitin family homolog 3 (Drosophila) adhesion NM_010494 ICAM2 intercellular adhesion molecule 2 adhesion NM_023892 ICAM4 (includes EG:3386) intercellular adhesion molecule 4 (Landsteiner-Wiener blood group)adhesion NM_001001979 MEGF10 multiple EGF-like-domains 10 adhesion NM_172522 MEGF11 multiple EGF-like-domains 11 adhesion NM_010739 MUC13 mucin 13, cell surface associated adhesion NM_013610 NINJ1 ninjurin 1 adhesion NM_016718 NINJ2 ninjurin 2 adhesion NM_172932 NLGN3 neuroligin -

Application of Microrna Database Mining in Biomarker Discovery and Identification of Therapeutic Targets for Complex Disease
Article Application of microRNA Database Mining in Biomarker Discovery and Identification of Therapeutic Targets for Complex Disease Jennifer L. Major, Rushita A. Bagchi * and Julie Pires da Silva * Department of Medicine, Division of Cardiology, University of Colorado Anschutz Medical Campus, Aurora, CO 80045, USA; [email protected] * Correspondence: [email protected] (R.A.B.); [email protected] (J.P.d.S.) Supplementary Tables Methods Protoc. 2021, 4, 5. https://doi.org/10.3390/mps4010005 www.mdpi.com/journal/mps Methods Protoc. 2021, 4, 5. https://doi.org/10.3390/mps4010005 2 of 25 Table 1. List of all hsa-miRs identified by Human microRNA Disease Database (HMDD; v3.2) analysis. hsa-miRs were identified using the term “genetics” and “circulating” as input in HMDD. Targets CAD hsa-miR-1 Targets IR injury hsa-miR-423 Targets Obesity hsa-miR-499 hsa-miR-146a Circulating Obesity Genetics CAD hsa-miR-423 hsa-miR-146a Circulating CAD hsa-miR-149 hsa-miR-499 Circulating IR Injury hsa-miR-146a Circulating Obesity hsa-miR-122 Genetics Stroke Circulating CAD hsa-miR-122 Circulating Stroke hsa-miR-122 Genetics Obesity Circulating Stroke hsa-miR-26b hsa-miR-17 hsa-miR-223 Targets CAD hsa-miR-340 hsa-miR-34a hsa-miR-92a hsa-miR-126 Circulating Obesity Targets IR injury hsa-miR-21 hsa-miR-423 hsa-miR-126 hsa-miR-143 Targets Obesity hsa-miR-21 hsa-miR-223 hsa-miR-34a hsa-miR-17 Targets CAD hsa-miR-223 hsa-miR-92a hsa-miR-126 Targets IR injury hsa-miR-155 hsa-miR-21 Circulating CAD hsa-miR-126 hsa-miR-145 hsa-miR-21 Targets Obesity hsa-mir-223 hsa-mir-499 hsa-mir-574 Targets IR injury hsa-mir-21 Circulating IR injury Targets Obesity hsa-mir-21 Targets CAD hsa-mir-22 hsa-mir-133a Targets IR injury hsa-mir-155 hsa-mir-21 Circulating Stroke hsa-mir-145 hsa-mir-146b Targets Obesity hsa-mir-21 hsa-mir-29b Methods Protoc. -

Arabidopsis Locus ID Gene Model Description TF Family Name I
SI Appendix Table S6.1 List of RBR-regulated transcription factors, and stress and stimulus-responsive transcripts in the egg cell, subtracted from the ovule transcriptomes (A) Transcription Factors Arabidopsis Gene Model Description TF Family Name Locus ID I. Enriched in rbr-3 AT1G08970 nuclear factor Y%2C subunit C9 CCAAT-HAP5 AT1G14410 ssDNA-binding transcriptional regulator Whirly AT1G14510 alfin-like 7 Alfin AT1G22070 transcription factor TGA3 bZIP AT1G32700 PLATZ transcription factor family protein PLATZ AT1G34370 C2H2 and C2HC zinc fingers superfamily protein C2H2 AT1G42990 basic region/leucine zipper motif 60 bZIP AT1G63490 transcription factor jumonji (jmjC) domain-containing protein JUMONJI AT2G02160 CCCH-type zinc finger family protein C3H AT2G17410 ARID/BRIGHT DNA-binding domain-containing protein ARID AT2G24430 NAC domain containing protein 38 NAC AT2G27050 ETHYLENE-INSENSITIVE3-like 1 EIL AT2G34720 nuclear factor Y%2C subunit A4 CCAAT-HAP2 AT2G35430 Zinc finger C-x8-C-x5-C-x3-H type family protein C3H AT2G36010 E2F transcription factor 3 E2F-DP AT2G39830 DA1-related protein 2 LIM AT3G07670 Rubisco methyltransferase family protein PcG AT3G10500 NAC domain containing protein 53 NAC AT3G10760 Homeodomain-like superfamily protein GARP-G2-like AT3G12130 KH domain-containing protein / zinc finger (CCCH type) family protein C3H AT3G59580 Plant regulator RWP-RK family protein Nin-like AT4G17750 heat shock factor 1 HSF AT4G24470 GATA-type zinc finger protein with TIFY domain-containing protein ZIM AT4G30935 WRKY DNA-binding protein -

Supplementary Table 1. List of Genes Up-Regulated in Abiraterone-Resistant Vcap Xenograft Samples PIK3IP1 Phosphoinositide-3-Kin
Supplementary Table 1. List of genes up-regulated in abiraterone-resistant VCaP xenograft samples PIK3IP1 phosphoinositide-3-kinase interacting protein 1 TMEM45A transmembrane protein 45A THBS1 thrombospondin 1 C7orf63 chromosome 7 open reading frame 63 OPTN optineurin FAM49A family with sequence similarity 49, member A APOL4 apolipoprotein L, 4 C17orf108|LOC201229 chromosome 17 open reading frame 108 | hypothetical protein LOC201229 SNORD94 small nucleolar RNA, C/D box 94 PCDHB11 protocadherin beta 11 RBM11 RNA binding motif protein 11 C6orf225 chromosome 6 open reading frame 225 KIAA1984|C9orf86|TMEM14 1 KIAA1984 | chromosome 9 open reading frame 86 | transmembrane protein 141 KIAA1107 TLR3 toll-like receptor 3 LPAR6 lysophosphatidic acid receptor 6 KIAA1683 GRB10 growth factor receptor-bound protein 10 TIMP2 TIMP metallopeptidase inhibitor 2 CCDC28A coiled-coil domain containing 28A FBXL2 F-box and leucine-rich repeat protein 2 NOV nephroblastoma overexpressed gene TSPAN31 tetraspanin 31 NR3C2 nuclear receptor subfamily 3, group C, member 2 DYNC2LI1 dynein, cytoplasmic 2, light intermediate chain 1 C15orf51 dynamin 1 pseudogene SAMD13 sterile alpha motif domain containing 13 RASSF6 Ras association (RalGDS/AF-6) domain family member 6 ZNF167 zinc finger protein 167 GATA2 GATA binding protein 2 NUDT7 nudix (nucleoside diphosphate linked moiety X)-type motif 7 DNAJC18 DnaJ (Hsp40) homolog, subfamily C, member 18 SNORA57 small nucleolar RNA, H/ACA box 57 CALCOCO1 calcium binding and coiled-coil domain 1 RLN2 relaxin 2 ING4 inhibitor of