Chromatin Condensation and Recruitment of PHD Finger Proteins
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

JB Review Biochemical and Structural Properties of Heterochromatin Protein 1: Understanding Its Role in Chromatin Assembly
J. Biochem. 2014;1Journal10 doi:10.1093/jb/mvu032 of Biochemistry Advance Access published June 4, 2014 JB Review Biochemical and structural properties of heterochromatin protein 1: understanding its role in chromatin assembly Received March 20, 2014; accepted April 9, 2014; published online May 13, 2014 Gohei Nishibuchi and Jun-ichi Nakayama* homologues and isoforms have been identified in eu- karyotic organisms, ranging from fission yeast to Graduate School of Natural Sciences, Nagoya City University, mammals, flies, and nematodes (69). In mammals, Nagoya 467-8501, Japan the three HP1 isoforms HP1a, HP1b and HP1g *Jun-ichi Nakayama, Graduate School of Natural Sciences, Nagoya (Fig. 1A) have different localization patterns within City University, 1 Yamanohata, Mizuho-cho, Mizuho-ku, Nagoya cells (11, 12). Mice deficient for each HP1 isoform ex- 467-8501, Japan. Tel: þ81-52-872-5838, Fax: þ81-52-872-5866, email: [email protected] hibit different phenotypes (1315), indicating that the isoforms have distinct functions in forming higher- Heterochromatin protein 1 (HP1) is an evolutionarily order chromatin structures. conserved chromosomal protein that binds lysine HP1-family proteins share a basic structure consist- Downloaded from 9-methylated histone H3 (H3K9me), a hallmark of het- ing of an amino-terminal chromodomain (CD) and a erochromatin, and plays a crucial role in forming carboxyl-terminal chromo shadow domain (CSD) higher-order chromatin structures. HP1 has an N-ter- linked by an unstructured hinge region (16). The CD minal chromodomain and a C-terminal chromo shadow functions as a binding module that targets lysine domain, linked by an unstructured hinge region. -
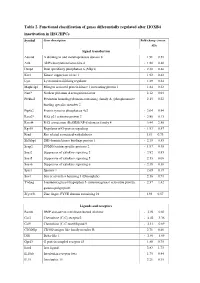
Table 2. Functional Classification of Genes Differentially Regulated After HOXB4 Inactivation in HSC/Hpcs
Table 2. Functional classification of genes differentially regulated after HOXB4 inactivation in HSC/HPCs Symbol Gene description Fold-change (mean ± SD) Signal transduction Adam8 A disintegrin and metalloprotease domain 8 1.91 ± 0.51 Arl4 ADP-ribosylation factor-like 4 - 1.80 ± 0.40 Dusp6 Dual specificity phosphatase 6 (Mkp3) - 2.30 ± 0.46 Ksr1 Kinase suppressor of ras 1 1.92 ± 0.42 Lyst Lysosomal trafficking regulator 1.89 ± 0.34 Mapk1ip1 Mitogen activated protein kinase 1 interacting protein 1 1.84 ± 0.22 Narf* Nuclear prelamin A recognition factor 2.12 ± 0.04 Plekha2 Pleckstrin homology domain-containing. family A. (phosphoinosite 2.15 ± 0.22 binding specific) member 2 Ptp4a2 Protein tyrosine phosphatase 4a2 - 2.04 ± 0.94 Rasa2* RAS p21 activator protein 2 - 2.80 ± 0.13 Rassf4 RAS association (RalGDS/AF-6) domain family 4 3.44 ± 2.56 Rgs18 Regulator of G-protein signaling - 1.93 ± 0.57 Rrad Ras-related associated with diabetes 1.81 ± 0.73 Sh3kbp1 SH3 domain kinase bindings protein 1 - 2.19 ± 0.53 Senp2 SUMO/sentrin specific protease 2 - 1.97 ± 0.49 Socs2 Suppressor of cytokine signaling 2 - 2.82 ± 0.85 Socs5 Suppressor of cytokine signaling 5 2.13 ± 0.08 Socs6 Suppressor of cytokine signaling 6 - 2.18 ± 0.38 Spry1 Sprouty 1 - 2.69 ± 0.19 Sos1 Son of sevenless homolog 1 (Drosophila) 2.16 ± 0.71 Ywhag 3-monooxygenase/tryptophan 5- monooxygenase activation protein. - 2.37 ± 1.42 gamma polypeptide Zfyve21 Zinc finger. FYVE domain containing 21 1.93 ± 0.57 Ligands and receptors Bambi BMP and activin membrane-bound inhibitor - 2.94 ± 0.62 -

Functional Roles of Bromodomain Proteins in Cancer
cancers Review Functional Roles of Bromodomain Proteins in Cancer Samuel P. Boyson 1,2, Cong Gao 3, Kathleen Quinn 2,3, Joseph Boyd 3, Hana Paculova 3 , Seth Frietze 3,4,* and Karen C. Glass 1,2,4,* 1 Department of Pharmaceutical Sciences, Albany College of Pharmacy and Health Sciences, Colchester, VT 05446, USA; [email protected] 2 Department of Pharmacology, Larner College of Medicine, University of Vermont, Burlington, VT 05405, USA; [email protected] 3 Department of Biomedical and Health Sciences, University of Vermont, Burlington, VT 05405, USA; [email protected] (C.G.); [email protected] (J.B.); [email protected] (H.P.) 4 University of Vermont Cancer Center, Burlington, VT 05405, USA * Correspondence: [email protected] (S.F.); [email protected] (K.C.G.) Simple Summary: This review provides an in depth analysis of the role of bromodomain-containing proteins in cancer development. As readers of acetylated lysine on nucleosomal histones, bromod- omain proteins are poised to activate gene expression, and often promote cancer progression. We examined changes in gene expression patterns that are observed in bromodomain-containing proteins and associated with specific cancer types. We also mapped the protein–protein interaction network for the human bromodomain-containing proteins, discuss the cellular roles of these epigenetic regu- lators as part of nine different functional groups, and identify bromodomain-specific mechanisms in cancer development. Lastly, we summarize emerging strategies to target bromodomain proteins in cancer therapy, including those that may be essential for overcoming resistance. Overall, this review provides a timely discussion of the different mechanisms of bromodomain-containing pro- Citation: Boyson, S.P.; Gao, C.; teins in cancer, and an updated assessment of their utility as a therapeutic target for a variety of Quinn, K.; Boyd, J.; Paculova, H.; cancer subtypes. -

Modes of Interaction of KMT2 Histone H3 Lysine 4 Methyltransferase/COMPASS Complexes with Chromatin
cells Review Modes of Interaction of KMT2 Histone H3 Lysine 4 Methyltransferase/COMPASS Complexes with Chromatin Agnieszka Bochy ´nska,Juliane Lüscher-Firzlaff and Bernhard Lüscher * ID Institute of Biochemistry and Molecular Biology, Medical School, RWTH Aachen University, Pauwelsstrasse 30, 52057 Aachen, Germany; [email protected] (A.B.); jluescher-fi[email protected] (J.L.-F.) * Correspondence: [email protected]; Tel.: +49-241-8088850; Fax: +49-241-8082427 Received: 18 January 2018; Accepted: 27 February 2018; Published: 2 March 2018 Abstract: Regulation of gene expression is achieved by sequence-specific transcriptional regulators, which convey the information that is contained in the sequence of DNA into RNA polymerase activity. This is achieved by the recruitment of transcriptional co-factors. One of the consequences of co-factor recruitment is the control of specific properties of nucleosomes, the basic units of chromatin, and their protein components, the core histones. The main principles are to regulate the position and the characteristics of nucleosomes. The latter includes modulating the composition of core histones and their variants that are integrated into nucleosomes, and the post-translational modification of these histones referred to as histone marks. One of these marks is the methylation of lysine 4 of the core histone H3 (H3K4). While mono-methylation of H3K4 (H3K4me1) is located preferentially at active enhancers, tri-methylation (H3K4me3) is a mark found at open and potentially active promoters. Thus, H3K4 methylation is typically associated with gene transcription. The class 2 lysine methyltransferases (KMTs) are the main enzymes that methylate H3K4. KMT2 enzymes function in complexes that contain a necessary core complex composed of WDR5, RBBP5, ASH2L, and DPY30, the so-called WRAD complex. -

Histone-Binding of DPF2 Mediates Its Repressive Role in Myeloid Differentiation
Histone-binding of DPF2 mediates its repressive role in myeloid differentiation Ferdinand M. Hubera,1, Sarah M. Greenblattb,1, Andrew M. Davenporta,1, Concepcion Martinezb,YeXub,LyP.Vuc, Stephen D. Nimerb,2, and André Hoelza,2 aDivision of Chemistry and Chemical Engineering, California Institute of Technology, Pasadena, CA 91125; bSylvester Comprehensive Cancer Center, University of Miami Miller School of Medicine, Miami, FL 33136; and cMolecular Pharmacology Program, Memorial Sloan Kettering Cancer Center, New York, NY 10065 Edited by Douglas C. Rees, Howard Hughes Medical Institute, California Institute of Technology, Pasadena, CA, and approved April 26, 2017 (received for review January 6, 2017) Double plant homeodomain finger 2 (DPF2) is a highly evolution- RUNX1 form a methylation-dependent repressive complex in arily conserved member of the d4 protein family that is ubiqui- AML, although it remains unclear whether the two proteins bind tously expressed in human tissues and was recently shown to each other directly or act concertedly as part of a larger complex. inhibit the myeloid differentiation of hematopoietic stem/progen- Here, we present the crystal structure of the human DPF2 itor and acute myelogenous leukemia cells. Here, we present the tandem PHD finger domain at a 1.6-Å resolution. We demon- crystal structure of the tandem plant homeodomain finger domain strate that the DPF2 tandem PHD finger domain binds acetylated of human DPF2 at 1.6-Å resolution. We show that DPF2 interacts H3 and H4 histone tails, identify the primary determinants of with the acetylated tails of both histones 3 and 4 via bipartite histone recognition, and confirm these interactions in vivo. -
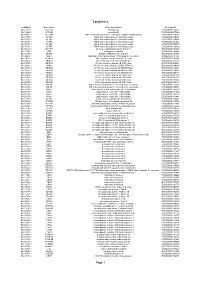
Epigenetics Page 1
Epigenetics esiRNA ID Gene Name Gene Description Ensembl ID HU-13237-1 ACTL6A actin-like 6A ENSG00000136518 HU-13925-1 ACTL6B actin-like 6B ENSG00000077080 HU-14457-1 ACTR1A ARP1 actin-related protein 1 homolog A, centractin alpha (yeast) ENSG00000138107 HU-10579-1 ACTR2 ARP2 actin-related protein 2 homolog (yeast) ENSG00000138071 HU-10837-1 ACTR3 ARP3 actin-related protein 3 homolog (yeast) ENSG00000115091 HU-09776-1 ACTR5 ARP5 actin-related protein 5 homolog (yeast) ENSG00000101442 HU-00773-1 ACTR6 ARP6 actin-related protein 6 homolog (yeast) ENSG00000075089 HU-07176-1 ACTR8 ARP8 actin-related protein 8 homolog (yeast) ENSG00000113812 HU-09411-1 AHCTF1 AT hook containing transcription factor 1 ENSG00000153207 HU-15150-1 AIRE autoimmune regulator ENSG00000160224 HU-12332-1 AKAP1 A kinase (PRKA) anchor protein 1 ENSG00000121057 HU-04065-1 ALG13 asparagine-linked glycosylation 13 homolog (S. cerevisiae) ENSG00000101901 HU-13552-1 ALKBH1 alkB, alkylation repair homolog 1 (E. coli) ENSG00000100601 HU-06662-1 ARID1A AT rich interactive domain 1A (SWI-like) ENSG00000117713 HU-12790-1 ARID1B AT rich interactive domain 1B (SWI1-like) ENSG00000049618 HU-09415-1 ARID2 AT rich interactive domain 2 (ARID, RFX-like) ENSG00000189079 HU-03890-1 ARID3A AT rich interactive domain 3A (BRIGHT-like) ENSG00000116017 HU-14677-1 ARID3B AT rich interactive domain 3B (BRIGHT-like) ENSG00000179361 HU-14203-1 ARID3C AT rich interactive domain 3C (BRIGHT-like) ENSG00000205143 HU-09104-1 ARID4A AT rich interactive domain 4A (RBP1-like) ENSG00000032219 HU-12512-1 ARID4B AT rich interactive domain 4B (RBP1-like) ENSG00000054267 HU-12520-1 ARID5A AT rich interactive domain 5A (MRF1-like) ENSG00000196843 HU-06595-1 ARID5B AT rich interactive domain 5B (MRF1-like) ENSG00000150347 HU-00556-1 ASF1A ASF1 anti-silencing function 1 homolog A (S. -

Aberrant Activity of Histone–Lysine N-Methyltransferase 2 (KMT2) Complexes in Oncogenesis
International Journal of Molecular Sciences Review Aberrant Activity of Histone–Lysine N-Methyltransferase 2 (KMT2) Complexes in Oncogenesis Elzbieta Poreba 1,* , Krzysztof Lesniewicz 2 and Julia Durzynska 1,* 1 Institute of Experimental Biology, Faculty of Biology, Adam Mickiewicz University, ul. Uniwersytetu Pozna´nskiego6, 61-614 Pozna´n,Poland 2 Department of Molecular and Cellular Biology, Institute of Molecular Biology and Biotechnology, Faculty of Biology, Adam Mickiewicz University, ul. Uniwersytetu Pozna´nskiego6, 61-614 Pozna´n,Poland; [email protected] * Correspondence: [email protected] (E.P.); [email protected] (J.D.); Tel.: +48-61-829-5857 (E.P.) Received: 19 November 2020; Accepted: 6 December 2020; Published: 8 December 2020 Abstract: KMT2 (histone-lysine N-methyltransferase subclass 2) complexes methylate lysine 4 on the histone H3 tail at gene promoters and gene enhancers and, thus, control the process of gene transcription. These complexes not only play an essential role in normal development but have also been described as involved in the aberrant growth of tissues. KMT2 mutations resulting from the rearrangements of the KMT2A (MLL1) gene at 11q23 are associated with pediatric mixed-lineage leukemias, and recent studies demonstrate that KMT2 genes are frequently mutated in many types of human cancers. Moreover, other components of the KMT2 complexes have been reported to contribute to oncogenesis. This review summarizes the recent advances in our knowledge of the role of KMT2 complexes in cell transformation. In addition, it discusses the therapeutic targeting of different components of the KMT2 complexes. Keywords: histone–lysine N-methyltransferase 2; COMPASS; COMPASS-like; H3K4 methylation; oncogenesis; cancer; epigenetics; chromatin 1. -

Pointing the Finger
RESEARCH HIGHLIGHTS Nature Reviews Molecular Cell Biology | AOP, published online 14 June 2006; doi:10.1038/nrm1966 DOI: 10.1038/nrm1966 URLs BPTF http://ca.expasy.org/uniprot/ Q9UIG2 ING2 http://ca.expasy.org/uniprot/ Q9ESK4 GENE EXPRESSION Pointing the finger Mechanistic links between specific histone-tail tumour suppressors, bound specifically to modifications and their effects on gene H3K4me3. ING2 is a subunit of a SIN3a–HDAC1 expression have been difficult to establish. histone-deacetylase complex. The authors However, four papers in Nature now identify the showed that in response to DNA damage, binding plant homeodomain (PHD) finger as an important of the ING2 PHD finger to H3K4me3 that is effector domain that binds to the trimethylated present at the promoters of actively transcribed K4 residue of histone H3 (H3K4me3) and couples proliferation genes enhanced the association of it to gene activation in one case and to gene the ING2–HDAC1 complex at these genes. This repression in another. resulted in increased histone-deacetylase activity Wysocka et al. affinity purified the BPTF and hence acute repression of the cognate (bromodomain and PHD finger transcription transcript. These findings, together with those of factor) subunit of NURF — an ATP-dependent Wysocka et al., indicate that PHD fingers have a chromatin remodelling complex — using an general role as effector domains that link H3K4me3-containing peptide. Mutational analysis H3K4me3 to diverse biological outcomes. narrowed the H3K4me3-interaction region to the The molecular mechanism that underlies the second, conserved bromodomain-proximal PHD recognition of H3K4me3 by the PHD finger of finger of BPTF. Knockdown of the histone ING2 was reported in a fourth linked paper. -

The Prevalence of Nucleic Acid Binding Among Reader Domains
molecules Review Reading More than Histones: The Prevalence of Nucleic Acid Binding among Reader Domains Tyler M. Weaver †, Emma A. Morrison † and Catherine A. Musselman * Department of Biochemistry, Carver College of Medicine, University of Iowa, Iowa City, IA 52242, USA; [email protected] (T.M.W.); [email protected] (E.A.M.) * Correspondence: [email protected] † These authors contributed equally to this work. Received: 1 September 2018; Accepted: 7 October 2018; Published: 12 October 2018 Abstract: The eukaryotic genome is packaged into the cell nucleus in the form of chromatin, a complex of genomic DNA and histone proteins. Chromatin structure regulation is critical for all DNA templated processes and involves, among many things, extensive post-translational modification of the histone proteins. These modifications can be “read out” by histone binding subdomains known as histone reader domains. A large number of reader domains have been identified and found to selectively recognize an array of histone post-translational modifications in order to target, retain, or regulate chromatin-modifying and remodeling complexes at their substrates. Interestingly, an increasing number of these histone reader domains are being identified as also harboring nucleic acid binding activity. In this review, we present a summary of the histone reader domains currently known to bind nucleic acids, with a focus on the molecular mechanisms of binding and the interplay between DNA and histone recognition. Additionally, we highlight the functional implications of nucleic acid binding in chromatin association and regulation. We propose that nucleic acid binding is as functionally important as histone binding, and that a significant portion of the as yet untested reader domains will emerge to have nucleic acid binding capabilities. -

The Arabidopsis LHP1 Protein Is a Component of Euchromatin
Planta (2005) 222: 910–925 DOI 10.1007/s00425-005-0129-4 ORIGINAL ARTICLE Marc Libault Æ Federico Tessadori Æ Sophie Germann Berend Snijder Æ Paul Fransz Æ Vale´rie Gaudin The Arabidopsis LHP1 protein is a component of euchromatin Received: 11 June 2005 / Accepted: 29 August 2005 / Published online: 22 October 2005 Ó Springer-Verlag 2005 Abstract The HP1 family proteins are involved in several A. thaliana HP1 homolog with the mammal HP1c iso- aspects of chromatin function and regulation in Dro- form, besides specific plant properties. sophila, mammals and the fission yeast. Here we inves- tigate the localization of LHP1, the unique Arabidopsis Keywords Arabidopsis Æ Chromatin Æ Heterochromatin thaliana HP1 homolog known at present time, to ap- protein 1 Æ Mitosis Æ Nucleolus proach its function. A functional LHP1–GFP fusion protein, able to restore the wild-type phenotype in the Abbreviations PI: Propidium iodide Æ DAPI: lhp1 mutant, was used to analyze the subnuclear distri- 4¢,6-diamidino-2-phenylindole Æ HP1: Heterochromatin bution of LHP1 in both A. thaliana and Nicotiana protein 1 Æ CD: Chromo domain Æ CSD: Chromo tabacum.InA. thaliana interphase nuclei, LHP1 was shadow domain Æ HR: Hinge region Æ RHF: Relative predominantly located outside the heterochromatic heterochromatin fraction Æ FISH: Fluorescent in-situ chromocenters. No major aberrations were observed in hybridization Æ NLS: Nuclear localization heterochromatin content or chromocenter organization signal Æ NoLS: Nucleolar localization signal in lhp1 plants. These data indicate that LHP1 is mainly involved in euchromatin organization in A. thaliana.In tobacco BY-2 cells, the LHP1 distribution, although in foci, slightly differed suggesting that LHP1 localization Introduction is determined by the underlying genome organization of plant species. -

Intact Nucleosomal Context Enables Chromodomain Reader
bioRxiv preprint doi: https://doi.org/10.1101/2020.04.30.070136; this version posted April 30, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Intact nucleosomal context enables chromodomain reader MRG15 to distinguish H3K36me3 from -me2 Sarah Faulkner1,2, Antony M. Couturier2, Brian Josephson1, Tom Watts1, Benjamin G. Davis1,3# and Fumiko Esashi2# 1 Department of Chemistry, University of Oxford, 12 Mansfield Rd, Oxford OX1 3TA, United Kingdom 2 Sir William Dunn School of Pathology, University of Oxford, S Parks Rd, Oxford OX1 3RE, United Kingdom 3 The Rosalind Franklin Institute, Oxfordshire, OX11 0FA, United Kingdom #To whom correspondence should be addressed: [email protected] and [email protected] Keywords: chromodomain, H3K36 methylation, MRG15, nucleosome, SETD2 1 bioRxiv preprint doi: https://doi.org/10.1101/2020.04.30.070136; this version posted April 30, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Abstract A wealth of in vivo evidence demonstrates the physiological importance of histone H3 trimethylation at lysine 36 (H3K36me3), to which chromodomain-containing proteins, such as MRG15, bind preferentially compared to their dimethyl (H3K36me2) counterparts. However, in vitro studies using isolated H3 peptides have failed to recapitulate a causal interaction. Here, we show that MRG15 can clearly discriminate between synthetic, fully intact model nucleosomes containing H3K36me2 and H3K36me3. MRG15 docking studies, along with experimental observations and nucleosome structure analysis suggest a model where the H3K36 side chain is sequestered in intact nucleosomes via a hydrogen bonding interaction with the DNA backbone, which is abrogated when the third methyl group is added to form H3K36me3. -

Supp Table 6.Pdf
Supplementary Table 6. Processes associated to the 2037 SCL candidate target genes ID Symbol Entrez Gene Name Process NM_178114 AMIGO2 adhesion molecule with Ig-like domain 2 adhesion NM_033474 ARVCF armadillo repeat gene deletes in velocardiofacial syndrome adhesion NM_027060 BTBD9 BTB (POZ) domain containing 9 adhesion NM_001039149 CD226 CD226 molecule adhesion NM_010581 CD47 CD47 molecule adhesion NM_023370 CDH23 cadherin-like 23 adhesion NM_207298 CERCAM cerebral endothelial cell adhesion molecule adhesion NM_021719 CLDN15 claudin 15 adhesion NM_009902 CLDN3 claudin 3 adhesion NM_008779 CNTN3 contactin 3 (plasmacytoma associated) adhesion NM_015734 COL5A1 collagen, type V, alpha 1 adhesion NM_007803 CTTN cortactin adhesion NM_009142 CX3CL1 chemokine (C-X3-C motif) ligand 1 adhesion NM_031174 DSCAM Down syndrome cell adhesion molecule adhesion NM_145158 EMILIN2 elastin microfibril interfacer 2 adhesion NM_001081286 FAT1 FAT tumor suppressor homolog 1 (Drosophila) adhesion NM_001080814 FAT3 FAT tumor suppressor homolog 3 (Drosophila) adhesion NM_153795 FERMT3 fermitin family homolog 3 (Drosophila) adhesion NM_010494 ICAM2 intercellular adhesion molecule 2 adhesion NM_023892 ICAM4 (includes EG:3386) intercellular adhesion molecule 4 (Landsteiner-Wiener blood group)adhesion NM_001001979 MEGF10 multiple EGF-like-domains 10 adhesion NM_172522 MEGF11 multiple EGF-like-domains 11 adhesion NM_010739 MUC13 mucin 13, cell surface associated adhesion NM_013610 NINJ1 ninjurin 1 adhesion NM_016718 NINJ2 ninjurin 2 adhesion NM_172932 NLGN3 neuroligin