A Fluorescent Sensor for the Real-Time Measurement of Extracellular Oxytocin Dynamics in the Brain
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

The Orphan Receptor GPR17 Is Unresponsive to Uracil Nucleotides and Cysteinyl Leukotrienes S
Supplemental material to this article can be found at: http://molpharm.aspetjournals.org/content/suppl/2017/03/02/mol.116.107904.DC1 1521-0111/91/5/518–532$25.00 https://doi.org/10.1124/mol.116.107904 MOLECULAR PHARMACOLOGY Mol Pharmacol 91:518–532, May 2017 Copyright ª 2017 by The American Society for Pharmacology and Experimental Therapeutics The Orphan Receptor GPR17 Is Unresponsive to Uracil Nucleotides and Cysteinyl Leukotrienes s Katharina Simon, Nicole Merten, Ralf Schröder, Stephanie Hennen, Philip Preis, Nina-Katharina Schmitt, Lucas Peters, Ramona Schrage,1 Celine Vermeiren, Michel Gillard, Klaus Mohr, Jesus Gomeza, and Evi Kostenis Molecular, Cellular and Pharmacobiology Section, Institute of Pharmaceutical Biology (K.S., N.M., Ral.S., S.H., P.P., N.-K.S, L.P., J.G., E.K.), Research Training Group 1873 (K.S., E.K.), Pharmacology and Toxicology Section, Institute of Pharmacy (Ram.S., K.M.), University of Bonn, Bonn, Germany; UCB Pharma, CNS Research, Braine l’Alleud, Belgium (C.V., M.G.). Downloaded from Received December 16, 2016; accepted March 1, 2017 ABSTRACT Pairing orphan G protein–coupled receptors (GPCRs) with their using eight distinct functional assay platforms based on label- cognate endogenous ligands is expected to have a major im- free pathway-unbiased biosensor technologies, as well as molpharm.aspetjournals.org pact on our understanding of GPCR biology. It follows that the canonical second-messenger or biochemical assays. Appraisal reproducibility of orphan receptor ligand pairs should be of of GPR17 activity can be accomplished with neither the coapplica- fundamental importance to guide meaningful investigations into tion of both ligand classes nor the exogenous transfection of partner the pharmacology and function of individual receptors. -

FTY720P Upregulates the Na+/K+ Atpase in Hepg2 Cells by Activating S1PR3 and Inducing PGE2 Release
Cellular Physiology Cell Physiol Biochem 2019;53:518-531 DOI: 10.33594/00000015510.33594/000000155 © 2019 The Author(s).© 2019 Published The Author(s) by and Biochemistry Published online: 11 SeptemberSeptember 20192019 Cell Physiol BiochemPublished Press GmbH&Co. by Cell Physiol KG Biochem 518 + + Press GmbH&Co. KG, Duesseldorf ChakkourAccepted: 5et September al.: FTY720-P 2019 Upregulates Na /K Atpase www.cellphysiolbiochem.com This article is licensed under the Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 Interna- tional License (CC BY-NC-ND). Usage and distribution for commercial purposes as well as any distribution of modified material requires written permission. Original Paper FTY720P Upregulates the Na+/K+ ATPase in HepG2 Cells by Activating S1PR3 and Inducing PGE2 Release Mohamed Chakkour Sawsan Kreydiyyeh Department of Biology, Faculty of Arts & Sciences, American University of Beirut, Beirut, Lebanon Key Words FTY720-P • Na+/K+ATPase • HepG2 • PKC • ERK • PGE2 Abstract Background/Aims: Liver regeneration is induced by S1P and accompanied with an increase in hepatic Na+/K+ ATPase activity, suggesting a potential modulatory role of the sphingolipid on the ATPase activity. The ability of S1P to alter the ATPase activity was confirmed in a previous work which showed a time dependent effect, with an inhibition appearing at 15min and a stimulation at two hours. The aim of this work was to investigate if FTY720-P, an analogue of S1P used in the treatment of multiple sclerosis, exerts a similar effect at 2 hours. Methods: HepG2 cells were treated with FTY720-P for two hours and the activity of the Na+/K+ ATPase was assayed by measuring the amount of inorganic phosphate liberated in presence and absence of ouabain. -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

Targeting Lysophosphatidic Acid in Cancer: the Issues in Moving from Bench to Bedside
View metadata, citation and similar papers at core.ac.uk brought to you by CORE provided by IUPUIScholarWorks cancers Review Targeting Lysophosphatidic Acid in Cancer: The Issues in Moving from Bench to Bedside Yan Xu Department of Obstetrics and Gynecology, Indiana University School of Medicine, 950 W. Walnut Street R2-E380, Indianapolis, IN 46202, USA; [email protected]; Tel.: +1-317-274-3972 Received: 28 August 2019; Accepted: 8 October 2019; Published: 10 October 2019 Abstract: Since the clear demonstration of lysophosphatidic acid (LPA)’s pathological roles in cancer in the mid-1990s, more than 1000 papers relating LPA to various types of cancer were published. Through these studies, LPA was established as a target for cancer. Although LPA-related inhibitors entered clinical trials for fibrosis, the concept of targeting LPA is yet to be moved to clinical cancer treatment. The major challenges that we are facing in moving LPA application from bench to bedside include the intrinsic and complicated metabolic, functional, and signaling properties of LPA, as well as technical issues, which are discussed in this review. Potential strategies and perspectives to improve the translational progress are suggested. Despite these challenges, we are optimistic that LPA blockage, particularly in combination with other agents, is on the horizon to be incorporated into clinical applications. Keywords: Autotaxin (ATX); ovarian cancer (OC); cancer stem cell (CSC); electrospray ionization tandem mass spectrometry (ESI-MS/MS); G-protein coupled receptor (GPCR); lipid phosphate phosphatase enzymes (LPPs); lysophosphatidic acid (LPA); phospholipase A2 enzymes (PLA2s); nuclear receptor peroxisome proliferator-activated receptor (PPAR); sphingosine-1 phosphate (S1P) 1. -

A Benchmarking Study on Virtual Ligand Screening Against Homology Models of Human Gpcrs
bioRxiv preprint doi: https://doi.org/10.1101/284075; this version posted March 19, 2018. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. A benchmarking study on virtual ligand screening against homology models of human GPCRs Victor Jun Yu Lima,b, Weina Dua, Yu Zong Chenc, Hao Fana,d aBioinformatics Institute (BII), Agency for Science, Technology and Research (A*STAR), 30 Biopolis Street, Matrix No. 07-01, 138671, Singapore; bSaw Swee Hock School of Public Health, National University of Singapore, 12 Science Drive 2, 117549, Singapore; cDepartment of Pharmacy, National University of Singapore, 18 Science Drive 4, 117543, Singapore; dDepartment of Biological Sciences, National University of Singapore, 16 Science Drive 4, Singapore 117558 To whom correspondence should be addressed: Dr. Hao Fan, Bioinformatics Institute (BII), Agency for Science, Technology and Research (A*STAR), 30 Biopolis Street, Matrix No. 07-01, 138671, Singapore; Telephone: +65 64788500; Email: [email protected] Keywords: G-protein-coupled receptor, GPCR, X-ray crystallography, virtual screening, homology modelling, consensus enrichment 1 bioRxiv preprint doi: https://doi.org/10.1101/284075; this version posted March 19, 2018. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Abstract G-protein-coupled receptor (GPCR) is an important target class of proteins for drug discovery, with over 27% of FDA-approved drugs targeting GPCRs. However, being a membrane protein, it is difficult to obtain the 3D crystal structures of GPCRs for virtual screening of ligands by molecular docking. -

G-Protein-Coupled Receptor Gpr17 Regulates Oligodendrocyte
www.nature.com/scientificreports OPEN G-Protein-Coupled Receptor Gpr17 Regulates Oligodendrocyte Diferentiation in Response Received: 11 May 2017 Accepted: 2 October 2017 to Lysolecithin-Induced Published: xx xx xxxx Demyelination Changqing Lu1,2, Lihua Dong2, Hui Zhou3, Qianmei Li3, Guojiao Huang3, Shu jun Bai3 & Linchuan Liao1 Oligodendrocytes are the myelin-producing cells of the central nervous system (CNS). A variety of brain disorders from “classical” demyelinating diseases, such as multiple sclerosis, stroke, schizophrenia, depression, Down syndrome and autism, are shown myelination defects. Oligodendrocyte myelination is regulated by a complex interplay of intrinsic, epigenetic and extrinsic factors. Gpr17 (G protein- coupled receptor 17) is a G protein-coupled receptor, and has been identifed to be a regulator for oligodendrocyte development. Here, we demonstrate that the absence of Gpr17 enhances remyelination in vivo with a toxin-induced model whereby focal demyelinated lesions are generated in spinal cord white matter of adult mice by localized injection of LPC(L-a-lysophosphatidylcholine). The increased expression of the activated form of Erk1/2 (phospho-Erk1/2) in lesion areas suggested the potential role of Erk1/2 activity on the Gpr17-dependent modulation of myelination. The absence of Gpr17 enhances remyelination is correlate with the activated Erk1/2 (phospho-Erk1/2).Being a membrane receptor, Gpr17 represents an ideal druggable target to be exploited for innovative regenerative approaches to acute and chronic CNS diseases. Oligodendrocytes are the myelin-producing cells of the central nervous system (CNS), and as such, wrap layers of lipid-dense insulating myelin around axons1. Mature oligodendrocytes have also been shown to provide met- abolic support to axons through transport systems within myelin, which may help prevent neurodegeneration2. -

Supplementary Table S5. Differentially Expressed Gene Lists of PD-1High CD39+ CD8 Tils According to 4-1BB Expression Compared to PD-1+ CD39- CD8 Tils
BMJ Publishing Group Limited (BMJ) disclaims all liability and responsibility arising from any reliance Supplemental material placed on this supplemental material which has been supplied by the author(s) J Immunother Cancer Supplementary Table S5. Differentially expressed gene lists of PD-1high CD39+ CD8 TILs according to 4-1BB expression compared to PD-1+ CD39- CD8 TILs Up- or down- regulated genes in Up- or down- regulated genes Up- or down- regulated genes only PD-1high CD39+ CD8 TILs only in 4-1BBneg PD-1high CD39+ in 4-1BBpos PD-1high CD39+ CD8 compared to PD-1+ CD39- CD8 CD8 TILs compared to PD-1+ TILs compared to PD-1+ CD39- TILs CD39- CD8 TILs CD8 TILs IL7R KLRG1 TNFSF4 ENTPD1 DHRS3 LEF1 ITGA5 MKI67 PZP KLF3 RYR2 SIK1B ANK3 LYST PPP1R3B ETV1 ADAM28 H2AC13 CCR7 GFOD1 RASGRP2 ITGAX MAST4 RAD51AP1 MYO1E CLCF1 NEBL S1PR5 VCL MPP7 MS4A6A PHLDB1 GFPT2 TNF RPL3 SPRY4 VCAM1 B4GALT5 TIPARP TNS3 PDCD1 POLQ AKAP5 IL6ST LY9 PLXND1 PLEKHA1 NEU1 DGKH SPRY2 PLEKHG3 IKZF4 MTX3 PARK7 ATP8B4 SYT11 PTGER4 SORL1 RAB11FIP5 BRCA1 MAP4K3 NCR1 CCR4 S1PR1 PDE8A IFIT2 EPHA4 ARHGEF12 PAICS PELI2 LAT2 GPRASP1 TTN RPLP0 IL4I1 AUTS2 RPS3 CDCA3 NHS LONRF2 CDC42EP3 SLCO3A1 RRM2 ADAMTSL4 INPP5F ARHGAP31 ESCO2 ADRB2 CSF1 WDHD1 GOLIM4 CDK5RAP1 CD69 GLUL HJURP SHC4 GNLY TTC9 HELLS DPP4 IL23A PITPNC1 TOX ARHGEF9 EXO1 SLC4A4 CKAP4 CARMIL3 NHSL2 DZIP3 GINS1 FUT8 UBASH3B CDCA5 PDE7B SOGA1 CDC45 NR3C2 TRIB1 KIF14 TRAF5 LIMS1 PPP1R2C TNFRSF9 KLRC2 POLA1 CD80 ATP10D CDCA8 SETD7 IER2 PATL2 CCDC141 CD84 HSPA6 CYB561 MPHOSPH9 CLSPN KLRC1 PTMS SCML4 ZBTB10 CCL3 CA5B PIP5K1B WNT9A CCNH GEM IL18RAP GGH SARDH B3GNT7 C13orf46 SBF2 IKZF3 ZMAT1 TCF7 NECTIN1 H3C7 FOS PAG1 HECA SLC4A10 SLC35G2 PER1 P2RY1 NFKBIA WDR76 PLAUR KDM1A H1-5 TSHZ2 FAM102B HMMR GPR132 CCRL2 PARP8 A2M ST8SIA1 NUF2 IL5RA RBPMS UBE2T USP53 EEF1A1 PLAC8 LGR6 TMEM123 NEK2 SNAP47 PTGIS SH2B3 P2RY8 S100PBP PLEKHA7 CLNK CRIM1 MGAT5 YBX3 TP53INP1 DTL CFH FEZ1 MYB FRMD4B TSPAN5 STIL ITGA2 GOLGA6L10 MYBL2 AHI1 CAND2 GZMB RBPJ PELI1 HSPA1B KCNK5 GOLGA6L9 TICRR TPRG1 UBE2C AURKA Leem G, et al. -

Guthrie Cdna Resource Center
cDNA Resource Center cDNA Resource Center Catalog cDNA Resource Center Missouri University of Science and Technology 400 W 11th Rolla, MO 65409 TEL: (573) 341-7610 FAX: (573) 341-7609 EMAIL: [email protected] www.cdna.org September, 2008 1 cDNA Resource Center Visit our web site for product updates 2 cDNA Resource Center The cDNA Resource Center The cDNA Resource Center is a service provided by the faculty of the Department of Biological Sciences of Missouri University of Science and Technology. The purpose of the cDNA Resource Center is to further scientific investigation by providing cDNA clones of human proteins involved in signal transduction processes. This is achieved by providing high quality clones for important signaling proteins in a timely manner. By high quality, we mean that the clones are • Sequence verified • Propagated in a versatile vector useful in bacterial and mammalian systems • Free of extraneous 3' and 5' untranslated regions • Expression verified (in most cases) by coupled in vitro transcription/translation assays • Available in wild-type, epitope-tagged and common mutant forms (e.g., constitutively- active or dominant negative) By timely, we mean that the clones are • Usually shipped within a day from when you place your order. Clones can be ordered from our web pages, by FAX or by phone. Within the United States, clones are shipped by overnight courier (FedEx); international orders are shipped International Priority (FedEx). The clones are supplied for research purposes only. Details on use of the material are included on the Material Transfer Agreement (page 3). Clones are distributed by agreement in Invitrogen's pcDNA3.1+ vector. -
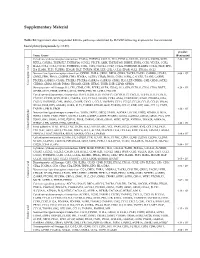
Supplementary Material
Supplementary Material Table S1: Significant downregulated KEGGs pathways identified by DAVID following exposure to five cinnamon- based phenylpropanoids (p < 0.05). p-value Term: Genes (Benjamini) Cytokine-cytokine receptor interaction: FASLG, TNFSF14, CXCL11, IL11, FLT3LG, CCL3L1, CCL3L3, CXCR6, XCR1, 2.43 × 105 RTEL1, CSF2RA, TNFRSF17, TNFRSF14, CCNL2, VEGFB, AMH, TNFRSF10B, INHBE, IFNB1, CCR3, VEGFA, CCR2, IL12A, CCL1, CCL3, CXCL5, TNFRSF25, CCR1, CSF1, CX3CL1, CCL7, CCL24, TNFRSF1B, IL12RB1, CCL21, FIGF, EPO, IL4, IL18R1, FLT1, TGFBR1, EDA2R, HGF, TNFSF8, KDR, LEP, GH2, CCL13, EPOR, XCL1, IFNA16, XCL2 Neuroactive ligand-receptor interaction: OPRM1, THRA, GRIK1, DRD2, GRIK2, TACR2, TACR1, GABRB1, LPAR4, 9.68 × 105 GRIK5, FPR1, PRSS1, GNRHR, FPR2, EDNRA, AGTR2, LTB4R, PRSS2, CNR1, S1PR4, CALCRL, TAAR5, GABRE, PTGER1, GABRG3, C5AR1, PTGER3, PTGER4, GABRA6, GABRA5, GRM1, PLG, LEP, CRHR1, GH2, GRM3, SSTR2, Chlorogenic acid Chlorogenic CHRM3, GRIA1, MC2R, P2RX2, TBXA2R, GHSR, HTR2C, TSHR, LHB, GLP1R, OPRD1 Hematopoietic cell lineage: IL4, CR1, CD8B, CSF1, FCER2, GYPA, ITGA2, IL11, GP9, FLT3LG, CD38, CD19, DNTT, 9.29 × 104 GP1BB, CD22, EPOR, CSF2RA, CD14, THPO, EPO, HLA-DRA, ITGA2B Cytokine-cytokine receptor interaction: IL6ST, IL21R, IL19, TNFSF15, CXCR3, IL15, CXCL11, TGFB1, IL11, FLT3LG, CXCL10, CCR10, XCR1, RTEL1, CSF2RA, IL21, CCNL2, VEGFB, CCR8, AMH, TNFRSF10C, IFNB1, PDGFRA, EDA, CXCL5, TNFRSF25, CSF1, IFNW1, CNTFR, CX3CL1, CCL5, TNFRSF4, CCL4, CCL27, CCL24, CCL25, CCL23, IFNA6, IFNA5, FIGF, EPO, AMHR2, IL2RA, FLT4, TGFBR2, EDA2R, -

LJMU Research Online
LJMU Research Online Fergani, C, Routly, JE, Jones, DN, Pickavance, LC, Smith, RF and Dobson, H KNDy neurone activation prior to the LH surge of the ewe is disrupted by LPS http://researchonline.ljmu.ac.uk/id/eprint/8794/ Article Citation (please note it is advisable to refer to the publisher’s version if you intend to cite from this work) Fergani, C, Routly, JE, Jones, DN, Pickavance, LC, Smith, RF and Dobson, H (2017) KNDy neurone activation prior to the LH surge of the ewe is disrupted by LPS. Reproduction, 154 (3). pp. 281-292. ISSN 1470-1626 LJMU has developed LJMU Research Online for users to access the research output of the University more effectively. Copyright © and Moral Rights for the papers on this site are retained by the individual authors and/or other copyright owners. Users may download and/or print one copy of any article(s) in LJMU Research Online to facilitate their private study or for non-commercial research. You may not engage in further distribution of the material or use it for any profit-making activities or any commercial gain. The version presented here may differ from the published version or from the version of the record. Please see the repository URL above for details on accessing the published version and note that access may require a subscription. For more information please contact [email protected] http://researchonline.ljmu.ac.uk/ 1 1 KNDy neurone activation prior to the LH surge of the ewe is disrupted by LPS 2 C. Fergania, J.E. -

The Determination of Activated Neurokinin B
Examining High Salt Diet, Puberty, and Interactions of Kisspeptin, Neurokinin B, and the Vasopressin Receptor Department of Zoology and Physiology, University of Wyoming Donal Skinner Dori Pitynski Brooke Fallon Background • Early puberty in females • Copenhagen Puberty Study- 2,095 girls • In 1991, mean age: 10.88 years • In 2006, mean age: 9.86 years • Adverse effects (Aksglaede, Pediatrics, 2009) Innovation (Centers for Disease Control and Prevention, 2009) KNDy cells, GnRH, and the reproductive axis • Kisspeptin, Neurokinin B, Dynorphin • GnRH: Gonadotropin releasing hormone (preoptic area) OVERVIEW Specific Aim of Research: 1. Do NKB/Kiss neurons have vasopressin receptors in rat brains? 2. Does salt increase the expression of NKB in rat brains around the time of puberty? Vasopressin • Arcuate nucleus- site of initiation of puberty • Kisspeptin neurons have V1aR- AVPV • Could the same be occurring in arcuate? • Salt = increased release of vasopressin • Link between salt and puberty via kiss/NKB (Shinji, 2013) Methods Brain tissue • Slicing on the cryostat • 20 micron slices • Fixed to slides and labeled for neurotransmitters via immunohistochemistry Immunohistochemistry • Fluorescence, double label (Dr. Mouktahr, Suez Canal University) Primary Antibody Stain 1% Serum Primary Antibody • 48 hours Rinse: 1% PBS + NaN3 Secondary Antibody Stain • Cover-slipped with Vectashield with DAPI Secondary Antibody Results from Kisspeptin/ V1aR double label DAPI V1aR Kisspeptin Merged Neurokinin B/V1aR double label • Antibodies raised in the same -

G Protein-Coupled Receptors
S.P.H. Alexander et al. The Concise Guide to PHARMACOLOGY 2015/16: G protein-coupled receptors. British Journal of Pharmacology (2015) 172, 5744–5869 THE CONCISE GUIDE TO PHARMACOLOGY 2015/16: G protein-coupled receptors Stephen PH Alexander1, Anthony P Davenport2, Eamonn Kelly3, Neil Marrion3, John A Peters4, Helen E Benson5, Elena Faccenda5, Adam J Pawson5, Joanna L Sharman5, Christopher Southan5, Jamie A Davies5 and CGTP Collaborators 1School of Biomedical Sciences, University of Nottingham Medical School, Nottingham, NG7 2UH, UK, 2Clinical Pharmacology Unit, University of Cambridge, Cambridge, CB2 0QQ, UK, 3School of Physiology and Pharmacology, University of Bristol, Bristol, BS8 1TD, UK, 4Neuroscience Division, Medical Education Institute, Ninewells Hospital and Medical School, University of Dundee, Dundee, DD1 9SY, UK, 5Centre for Integrative Physiology, University of Edinburgh, Edinburgh, EH8 9XD, UK Abstract The Concise Guide to PHARMACOLOGY 2015/16 provides concise overviews of the key properties of over 1750 human drug targets with their pharmacology, plus links to an open access knowledgebase of drug targets and their ligands (www.guidetopharmacology.org), which provides more detailed views of target and ligand properties. The full contents can be found at http://onlinelibrary.wiley.com/doi/ 10.1111/bph.13348/full. G protein-coupled receptors are one of the eight major pharmacological targets into which the Guide is divided, with the others being: ligand-gated ion channels, voltage-gated ion channels, other ion channels, nuclear hormone receptors, catalytic receptors, enzymes and transporters. These are presented with nomenclature guidance and summary information on the best available pharmacological tools, alongside key references and suggestions for further reading.