This Thesis Has Been Submitted in Fulfilment of the Requirements for a Postgraduate Degree (E.G
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Supplemental Information to Mammadova-Bach Et Al., “Laminin Α1 Orchestrates VEGFA Functions in the Ecosystem of Colorectal Carcinogenesis”
Supplemental information to Mammadova-Bach et al., “Laminin α1 orchestrates VEGFA functions in the ecosystem of colorectal carcinogenesis” Supplemental material and methods Cloning of the villin-LMα1 vector The plasmid pBS-villin-promoter containing the 3.5 Kb of the murine villin promoter, the first non coding exon, 5.5 kb of the first intron and 15 nucleotides of the second villin exon, was generated by S. Robine (Institut Curie, Paris, France). The EcoRI site in the multi cloning site was destroyed by fill in ligation with T4 polymerase according to the manufacturer`s instructions (New England Biolabs, Ozyme, Saint Quentin en Yvelines, France). Site directed mutagenesis (GeneEditor in vitro Site-Directed Mutagenesis system, Promega, Charbonnières-les-Bains, France) was then used to introduce a BsiWI site before the start codon of the villin coding sequence using the 5’ phosphorylated primer: 5’CCTTCTCCTCTAGGCTCGCGTACGATGACGTCGGACTTGCGG3’. A double strand annealed oligonucleotide, 5’GGCCGGACGCGTGAATTCGTCGACGC3’ and 5’GGCCGCGTCGACGAATTCACGC GTCC3’ containing restriction site for MluI, EcoRI and SalI were inserted in the NotI site (present in the multi cloning site), generating the plasmid pBS-villin-promoter-MES. The SV40 polyA region of the pEGFP plasmid (Clontech, Ozyme, Saint Quentin Yvelines, France) was amplified by PCR using primers 5’GGCGCCTCTAGATCATAATCAGCCATA3’ and 5’GGCGCCCTTAAGATACATTGATGAGTT3’ before subcloning into the pGEMTeasy vector (Promega, Charbonnières-les-Bains, France). After EcoRI digestion, the SV40 polyA fragment was purified with the NucleoSpin Extract II kit (Machery-Nagel, Hoerdt, France) and then subcloned into the EcoRI site of the plasmid pBS-villin-promoter-MES. Site directed mutagenesis was used to introduce a BsiWI site (5’ phosphorylated AGCGCAGGGAGCGGCGGCCGTACGATGCGCGGCAGCGGCACG3’) before the initiation codon and a MluI site (5’ phosphorylated 1 CCCGGGCCTGAGCCCTAAACGCGTGCCAGCCTCTGCCCTTGG3’) after the stop codon in the full length cDNA coding for the mouse LMα1 in the pCIS vector (kindly provided by P. -

Transportin 1 Accumulates Specifically with FET Proteins but No Other
CORE Metadata, citation and similar papers at core.ac.uk Provided by RERO DOC Digital Library Acta Neuropathol DOI 10.1007/s00401-012-1020-6 ORIGINAL PAPER Transportin 1 accumulates specifically with FET proteins but no other transportin cargos in FTLD-FUS and is absent in FUS inclusions in ALS with FUS mutations Manuela Neumann • Chiara F. Valori • Olaf Ansorge • Hans A. Kretzschmar • David G. Munoz • Hirofumi Kusaka • Osamu Yokota • Kenji Ishihara • Lee-Cyn Ang • Juan M. Bilbao • Ian R. A. Mackenzie Received: 22 May 2012 / Accepted: 16 July 2012 Ó Springer-Verlag 2012 Abstract Accumulation of the DNA/RNA binding pro- in these conditions. While ALS-FUS showed only accu- tein fused in sarcoma (FUS) as inclusions in neurons and mulation of FUS, inclusions in FTLD-FUS revealed co- glia is the pathological hallmark of amyotrophic lateral accumulation of all members of the FET protein family, that sclerosis patients with mutations in FUS (ALS-FUS) as well include FUS, Ewing’s sarcoma (EWS) and TATA-binding as in several subtypes of frontotemporal lobar degeneration protein-associated factor 15 (TAF15) suggesting a more (FTLD-FUS), which are not associated with FUS muta- complex disturbance of transportin-mediated nuclear tions. Despite some overlap in the phenotype and import of proteins in FTLD-FUS compared to ALS-FUS. neuropathology of FTLD-FUS and ALS-FUS, significant To gain more insight into the mechanisms of inclusion body differences of potential pathomechanistic relevance were formation, we investigated the role of Transportin 1 (Trn1) recently identified in the protein composition of inclusions as well as 13 additional cargo proteins of Transportin in the spectrum of FUS-opathies by immunohistochemistry and biochemically. -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

FUS-NLS/Transportin 1 Complex Structure Provides
University of Kentucky UKnowledge Molecular and Cellular Biochemistry Faculty Molecular and Cellular Biochemistry Publications 10-8-2012 FUS-NLS/Transportin 1 complex structure provides insights into the nuclear targeting mechanism of FUS and the implications in ALS Chunyan Niu Chinese Academy of Sciences, China Jiayu Zhang University of Kentucky, [email protected] Feng Gao Chinese Academy of Sciences, China Liuqing Yang University of Kentucky, [email protected] Minze Jia Chinese Academy of Sciences, China See next page for additional authors Right click to open a feedback form in a new tab to let us know how this document benefits oy u. Follow this and additional works at: https://uknowledge.uky.edu/biochem_facpub Part of the Biochemistry, Biophysics, and Structural Biology Commons Repository Citation Niu, Chunyan; Zhang, Jiayu; Gao, Feng; Yang, Liuqing; Jia, Minze; Zhu, Haining; and Gong, Weimin, "FUS-NLS/Transportin 1 complex structure provides insights into the nuclear targeting mechanism of FUS and the implications in ALS" (2012). Molecular and Cellular Biochemistry Faculty Publications. 9. https://uknowledge.uky.edu/biochem_facpub/9 This Article is brought to you for free and open access by the Molecular and Cellular Biochemistry at UKnowledge. It has been accepted for inclusion in Molecular and Cellular Biochemistry Faculty Publications by an authorized administrator of UKnowledge. For more information, please contact [email protected]. Authors Chunyan Niu, Jiayu Zhang, Feng Gao, Liuqing Yang, Minze Jia, Haining Zhu, and Weimin Gong FUS-NLS/Transportin 1 complex structure provides insights into the nuclear targeting mechanism of FUS and the implications in ALS Notes/Citation Information Published in PLoS ONE, v. -

Cyclin K Interacts with Β-Catenin to Induce Cyclin D1 Expression And
Theranostics 2020, Vol. 10, Issue 24 11144 Ivyspring International Publisher Theranostics 2020; 10(24): 11144-11158. doi: 10.7150/thno.42578 Research Paper Cyclin K interacts with β-catenin to induce Cyclin D1 expression and facilitates tumorigenesis and radioresistance in lung cancer Guojun Yao*, Jing Tang*, Xijie Yang, Ye Zhao, Rui Zhou, Rui Meng, Sheng Zhang, Xiaorong Dong, Tao Zhang, Kunyu Yang, Gang Wu and Shuangbing Xu Cancer Center, Union Hospital, Tongji Medical College, Huazhong University of Science and Technology, Wuhan 430022, China. *These authors contributed equally to this work. Corresponding author: Shuangbing Xu or Gang Wu, Cancer Center, Union Hospital, Tongji Medical College, Huazhong University of Science and Technology, Wuhan 430022, China. E-mail: [email protected] or [email protected]. © The author(s). This is an open access article distributed under the terms of the Creative Commons Attribution License (https://creativecommons.org/licenses/by/4.0/). See http://ivyspring.com/terms for full terms and conditions. Received: 2019.11.29; Accepted: 2020.08.24; Published: 2020.09.11 Abstract Rationale: Radioresistance remains the major cause of local relapse and distant metastasis in lung cancer. However, the underlying molecular mechanisms remain poorly defined. This study aimed to investigate the role and regulatory mechanism of Cyclin K in lung cancer radioresistance. Methods: Expression levels of Cyclin K were measured by immunohistochemistry in human lung cancer tissues and adjacent normal lung tissues. Cell growth and proliferation, neutral comet and foci formation assays, G2/M checkpoint and a xenograft mouse model were used for functional analyses. Gene expression was examined by RNA sequencing and quantitative real-time PCR. -

Protein Identities in Evs Isolated from U87-MG GBM Cells As Determined by NG LC-MS/MS
Protein identities in EVs isolated from U87-MG GBM cells as determined by NG LC-MS/MS. No. Accession Description Σ Coverage Σ# Proteins Σ# Unique Peptides Σ# Peptides Σ# PSMs # AAs MW [kDa] calc. pI 1 A8MS94 Putative golgin subfamily A member 2-like protein 5 OS=Homo sapiens PE=5 SV=2 - [GG2L5_HUMAN] 100 1 1 7 88 110 12,03704523 5,681152344 2 P60660 Myosin light polypeptide 6 OS=Homo sapiens GN=MYL6 PE=1 SV=2 - [MYL6_HUMAN] 100 3 5 17 173 151 16,91913397 4,652832031 3 Q6ZYL4 General transcription factor IIH subunit 5 OS=Homo sapiens GN=GTF2H5 PE=1 SV=1 - [TF2H5_HUMAN] 98,59 1 1 4 13 71 8,048185945 4,652832031 4 P60709 Actin, cytoplasmic 1 OS=Homo sapiens GN=ACTB PE=1 SV=1 - [ACTB_HUMAN] 97,6 5 5 35 917 375 41,70973209 5,478027344 5 P13489 Ribonuclease inhibitor OS=Homo sapiens GN=RNH1 PE=1 SV=2 - [RINI_HUMAN] 96,75 1 12 37 173 461 49,94108966 4,817871094 6 P09382 Galectin-1 OS=Homo sapiens GN=LGALS1 PE=1 SV=2 - [LEG1_HUMAN] 96,3 1 7 14 283 135 14,70620005 5,503417969 7 P60174 Triosephosphate isomerase OS=Homo sapiens GN=TPI1 PE=1 SV=3 - [TPIS_HUMAN] 95,1 3 16 25 375 286 30,77169764 5,922363281 8 P04406 Glyceraldehyde-3-phosphate dehydrogenase OS=Homo sapiens GN=GAPDH PE=1 SV=3 - [G3P_HUMAN] 94,63 2 13 31 509 335 36,03039959 8,455566406 9 Q15185 Prostaglandin E synthase 3 OS=Homo sapiens GN=PTGES3 PE=1 SV=1 - [TEBP_HUMAN] 93,13 1 5 12 74 160 18,68541938 4,538574219 10 P09417 Dihydropteridine reductase OS=Homo sapiens GN=QDPR PE=1 SV=2 - [DHPR_HUMAN] 93,03 1 1 17 69 244 25,77302971 7,371582031 11 P01911 HLA class II histocompatibility antigen, -
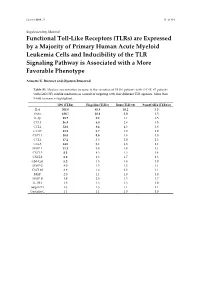
Are Expressed by a Majority of Primary Human Acute Myeloid Leukemia Cells and Inducibility of the TLR Signaling Pathway Is Associated with a More Favorable Phenotype
Cancers 2019, 11 S1 of S19 Supplementary Material Functional Toll-Like Receptors (TLRs) are Expressed by a Majority of Primary Human Acute Myeloid Leukemia Cells and Inducibility of the TLR Signaling Pathway is Associated with a More Favorable Phenotype Annette K. Brenner and Øystein Bruserud Table S1. Median concentration increase in the secretion of 19 (16 patients with G-CSF, 67 patients with GM-CSF) soluble mediators as a result of targeting with four different TLR agonists. More than 5-fold increase is highlighted. LPS (TLR4) Flagellin (TLR5) R848 (TLR7/8) Pam3CSK4 (TLR1/2) IL-6 301.0 45.9 10.2 5.3 TNFα 188.7 10.6 5.0 1.5 IL-1β 89.7 9.2 1.1 1.5 CCL3 56.4 6.0 2.4 1.5 CCL2 52.6 9.4 4.3 2.8 G-CSF 51.9 2.7 1.0 1.0 CXCL1 38.0 5.8 1.4 1.5 CCL4 17.2 3.3 2.0 1.3 CCL5 14.5 2.1 1.3 1.1 MMP-1 11.1 3.4 1.4 1.1 CXCL5 8.3 4.3 1.3 1.4 CXCL8 6.0 4.3 1.7 1.3 GM-CSF 5.2 1.5 1.4 1.0 MMP-2 4.0 1.5 1.2 1.1 CXCL10 3.9 1.6 2.2 1.1 HGF 2.0 1.1 1.0 1.0 MMP-9 1.9 2.5 1.3 1.7 IL-1RA 1.8 1.3 1.3 1.0 Serpin E1 1.3 1.3 1.1 1.1 Cystatin C 1.1 1.1 1.0 1.0 Cancers 2019, 11 S2 of S19 Table S2. -

Cytotoxic Effects and Changes in Gene Expression Profile
Toxicology in Vitro 34 (2016) 309–320 Contents lists available at ScienceDirect Toxicology in Vitro journal homepage: www.elsevier.com/locate/toxinvit Fusarium mycotoxin enniatin B: Cytotoxic effects and changes in gene expression profile Martina Jonsson a,⁎,MarikaJestoib, Minna Anthoni a, Annikki Welling a, Iida Loivamaa a, Ville Hallikainen c, Matti Kankainen d, Erik Lysøe e, Pertti Koivisto a, Kimmo Peltonen a,f a Chemistry and Toxicology Research Unit, Finnish Food Safety Authority (Evira), Mustialankatu 3, FI-00790 Helsinki, Finland b Product Safety Unit, Finnish Food Safety Authority (Evira), Mustialankatu 3, FI-00790 Helsinki, c The Finnish Forest Research Institute, Rovaniemi Unit, P.O. Box 16, FI-96301 Rovaniemi, Finland d Institute for Molecular Medicine Finland (FIMM), University of Helsinki, P.O. Box 20, FI-00014, Finland e Plant Health and Biotechnology, Norwegian Institute of Bioeconomy, Høyskoleveien 7, NO -1430 Ås, Norway f Finnish Safety and Chemicals Agency (Tukes), Opastinsilta 12 B, FI-00521 Helsinki, Finland article info abstract Article history: The mycotoxin enniatin B, a cyclic hexadepsipeptide produced by the plant pathogen Fusarium,isprevalentin Received 3 December 2015 grains and grain-based products in different geographical areas. Although enniatins have not been associated Received in revised form 5 April 2016 with toxic outbreaks, they have caused toxicity in vitro in several cell lines. In this study, the cytotoxic effects Accepted 28 April 2016 of enniatin B were assessed in relation to cellular energy metabolism, cell proliferation, and the induction of ap- Available online 6 May 2016 optosis in Balb 3T3 and HepG2 cells. The mechanism of toxicity was examined by means of whole genome ex- fi Keywords: pression pro ling of exposed rat primary hepatocytes. -

Cellular and Molecular Signatures in the Disease Tissue of Early
Cellular and Molecular Signatures in the Disease Tissue of Early Rheumatoid Arthritis Stratify Clinical Response to csDMARD-Therapy and Predict Radiographic Progression Frances Humby1,* Myles Lewis1,* Nandhini Ramamoorthi2, Jason Hackney3, Michael Barnes1, Michele Bombardieri1, Francesca Setiadi2, Stephen Kelly1, Fabiola Bene1, Maria di Cicco1, Sudeh Riahi1, Vidalba Rocher-Ros1, Nora Ng1, Ilias Lazorou1, Rebecca E. Hands1, Desiree van der Heijde4, Robert Landewé5, Annette van der Helm-van Mil4, Alberto Cauli6, Iain B. McInnes7, Christopher D. Buckley8, Ernest Choy9, Peter Taylor10, Michael J. Townsend2 & Costantino Pitzalis1 1Centre for Experimental Medicine and Rheumatology, William Harvey Research Institute, Barts and The London School of Medicine and Dentistry, Queen Mary University of London, Charterhouse Square, London EC1M 6BQ, UK. Departments of 2Biomarker Discovery OMNI, 3Bioinformatics and Computational Biology, Genentech Research and Early Development, South San Francisco, California 94080 USA 4Department of Rheumatology, Leiden University Medical Center, The Netherlands 5Department of Clinical Immunology & Rheumatology, Amsterdam Rheumatology & Immunology Center, Amsterdam, The Netherlands 6Rheumatology Unit, Department of Medical Sciences, Policlinico of the University of Cagliari, Cagliari, Italy 7Institute of Infection, Immunity and Inflammation, University of Glasgow, Glasgow G12 8TA, UK 8Rheumatology Research Group, Institute of Inflammation and Ageing (IIA), University of Birmingham, Birmingham B15 2WB, UK 9Institute of -

A Universal Transportin Protein Drives Stochastic Choice of Olfactory Neurons Via Specific Nuclear Import of a Sox-2-Activating Factor
A universal transportin protein drives stochastic choice of olfactory neurons via specific nuclear import of a sox-2-activating factor Amel Alqadaha, Yi-Wen Hsieha,1, Rui Xionga,1, Bluma J. Leschb, Chieh Changa, and Chiou-Fen Chuanga,2 aDepartment of Biological Sciences, University of Illinois at Chicago, IL 60607; and bDepartment of Genetics, Yale University School of Medicine, New Haven, CT 06510 Edited by Iva Greenwald, Columbia University, New York, NY, and approved October 31, 2019 (received for review June 25, 2019) Stochastic neuronal cell fate choice involving notch-independent mechanisms act downstream of the BK potassium channels to mechanisms is a poorly understood biological process. The induce the AWCON identity. Caenorhabditis elegans AWC olfactory neuron pair asymmetrically Here, we identify a role of the karyopherin imb-2/transportin 1 differentiates into the default AWCOFF and induced AWCON subtypes downstream of the SLO BK potassium channels in promoting in a stochastic manner. Stochastic choice of the AWCON subtype is the AWCON subtype from an unbiased forward genetic screen. established using gap junctions and SLO BK potassium channels to We show that asymmetrical expression of imb-2 in AWCON cells, repress a calcium-activated protein kinase pathway. However, it is which is dependent on nsy-5 (gap junction) and slo-1 (BK po- unknown how the potassium channel-repressed calcium signaling is tassium channel), is necessary and sufficient for AWC asymmetry. translated into the induction of the AWCON subtype. Here, we iden- In addition, IMB-2 localizes in close proximity to the homeo- tify a detailed working mechanism of how the homeodomain-like domain-like transcription factor NSY-7 and mediates nuclear transcription factor NSY-7, previously described as a repressor in the transport of NSY-7 to specify the AWCON subtype. -

Human Cyclin-Dependent Kinase 12 (CDK12), Kinase Domain a Target Enabling Package (TEP)
Human Cyclin-Dependent Kinase 12 (CDK12), Kinase Domain A Target Enabling Package (TEP) Gene ID / UniProt ID / EC CDK12, 51755 / Q9NYV4/ 2.7.11.22, 2.7.11.23 CCNK, 8812 / O75909/ - Target Nominator Gregg Morin (UBC, Canada), Nathanael Gray (Harvard) SGC Authors Sarah E. Dixon-Clarke, Jonathan M. Elkins, Nathanael S. Gray, and Alex N. Bullock Collaborating Authors S.-W. Grace Cheng1, Tinghu Zhang2, Nicholas Kwiatkowski3, Calla M. Olson2, Brian J. Abraham3, Ann K. Greifenberg4, Scott B. Ficarro2, Yanke Liang2, Nancy M. Hannett3, Theresa Manz5, Mingfeng Hao2, Bartlomiej Bartkowiak6, Arno L. Greenleaf6, Jarrod A. Marto2, Matthias Geyer4, Richard A. Young3, and Gregg B. Morin1 Target PI Alex Bullock (SGC Oxford) Therapeutic Area(s) Oncology Disease Relevance CDK12 loss sensitises cancer cells to DNA damage Date approved by TEP 17th June 2016 Evaluation Group Document version Version 3 Document version date April 2018 DOI https://doi.org/10.5281/zenodo.1219680 Affiliations 1. Canada's Michael Smith Genome Sciences Centre, BC Cancer Agency 2. Department of Cancer Biology, Dana-Farber Cancer Institute 3. Whitehead Institute for Biomedical Research 4. Department of Structural Immunology, Institute of Innate Immunity 5. Pharmaceutical and Medicinal Chemistry, Saarland University 6. Department of Biochemistry, Duke University Medical Center USEFUL LINKS (Please note that the inclusion of links to external sites should not be taken as an endorsement of that site by the SGC in any way) SUMMARY OF PROJECT Cyclin-dependent kinase 12 (CDK12) phosphorylates RNA Pol II C-terminal domain (CTD) to promote transcriptional elongation of large DNA damage response genes. CDK12 is frequently mutated or amplified in cancer and its loss sensitises cells to DNA damage. -

Protein Nuclear Import and Beyond ⇑ Laure Twyffels A,B, , Cyril Gueydan A,1, Véronique Kruys A,B,1
View metadata, citation and similar papers at core.ac.uk brought to you by CORE provided by Elsevier - Publisher Connector FEBS Letters 588 (2014) 1857–1868 journal homepage: www.FEBSLetters.org Review Transportin-1 and Transportin-2: Protein nuclear import and beyond ⇑ Laure Twyffels a,b, , Cyril Gueydan a,1, Véronique Kruys a,b,1 a Laboratoire de Biologie moléculaire du gène (CP300), Faculté des Sciences, Université Libre de Bruxelles (ULB), Belgium b Center for Microscopy and Molecular Imaging (CMMI), 6041 Gosselies, Belgium article info abstract Article history: Nearly 20 years after its identification as a new b-karyopherin mediating the nuclear import of the Received 15 February 2014 RNA-binding protein hnRNP A1, Transportin-1 is still commonly overlooked in comparison with its Revised 12 April 2014 best known cousin, Importin-b. Transportin-1 is nonetheless a considerable player in nucleo-cyto- Accepted 16 April 2014 plasmic transport. Over the past few years, significant progress has been made in the characteriza- Available online 26 April 2014 tion of the nuclear localization signals (NLSs) that Transportin-1 recognizes, thereby providing the Edited by Ulrike Kutay molecular basis of its diversified repertoire of cargoes. The recent discovery that mutations in the Transportin-dependent NLS of FUS cause mislocalization of this protein and result in amyotrophic lateral sclerosis illustrates the importance of Transportin-dependent import for human health. Keywords: Transportin Besides, new functions of Transportin-1 are emerging in processes other than nuclear import. Here, Importins we summarize what is known about Transportin-1 and the related b-karyopherin Transportin-2. Karyopherins Ó 2014 Federation of European Biochemical Societies.