Human Cyclin-Dependent Kinase 12 (CDK12), Kinase Domain a Target Enabling Package (TEP)
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Investigating the Role of Cdk11in Animal Cytokinesis
Investigating the Role of CDK11 in Animal Cytokinesis by Thomas Clifford Panagiotou A thesis submitted in conformity with the requirements for the degree of Master of Science Department of Molecular Genetics University of Toronto © Copyright by Thomas Clifford Panagiotou (2020) Investigating the Role of CDK11 in Animal Cytokinesis Thomas Clifford Panagiotou Master of Science Department of Molecular Genetics University of Toronto 2020 Abstract Finely tuned spatio-temporal regulation of cell division is required for genome stability. Cytokinesis constitutes the final stages of cell division, from chromosome segregation to the physical separation of cells, abscission. Abscission is tightly regulated to ensure it occurs after earlier cytokinetic events, like the maturation of the stem body, the regulatory platform for abscission. Active Aurora B kinase enforces the abscission checkpoint, which blocks abscission until chromosomes have been cleared from the cytokinetic machinery. Currently, it is unclear how this checkpoint is overcome. Here, I demonstrate that the cyclin-dependent kinase CDK11 is required for cytokinesis. Both inhibition and depletion of CDK11 block abscission. Furthermore, the mitosis-specific CDK11p58 kinase localizes to the stem body, where its kinase activity rescues the defects of CDK11 depletion and inhibition. These results suggest a model whereby CDK11p58 antagonizes Aurora B kinase to overcome the abscission checkpoint to allow for successful completion of cytokinesis. ii Acknowledgments I am very grateful for the support of my family and friends throughout my studies. I would also like to express my deep gratitude to Wilde Lab members, both past and present, for their advice and collaboration. In particular, I am very grateful to Matthew Renshaw, whose work comprises part of this thesis. -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

Cyclin K Interacts with Β-Catenin to Induce Cyclin D1 Expression And
Theranostics 2020, Vol. 10, Issue 24 11144 Ivyspring International Publisher Theranostics 2020; 10(24): 11144-11158. doi: 10.7150/thno.42578 Research Paper Cyclin K interacts with β-catenin to induce Cyclin D1 expression and facilitates tumorigenesis and radioresistance in lung cancer Guojun Yao*, Jing Tang*, Xijie Yang, Ye Zhao, Rui Zhou, Rui Meng, Sheng Zhang, Xiaorong Dong, Tao Zhang, Kunyu Yang, Gang Wu and Shuangbing Xu Cancer Center, Union Hospital, Tongji Medical College, Huazhong University of Science and Technology, Wuhan 430022, China. *These authors contributed equally to this work. Corresponding author: Shuangbing Xu or Gang Wu, Cancer Center, Union Hospital, Tongji Medical College, Huazhong University of Science and Technology, Wuhan 430022, China. E-mail: [email protected] or [email protected]. © The author(s). This is an open access article distributed under the terms of the Creative Commons Attribution License (https://creativecommons.org/licenses/by/4.0/). See http://ivyspring.com/terms for full terms and conditions. Received: 2019.11.29; Accepted: 2020.08.24; Published: 2020.09.11 Abstract Rationale: Radioresistance remains the major cause of local relapse and distant metastasis in lung cancer. However, the underlying molecular mechanisms remain poorly defined. This study aimed to investigate the role and regulatory mechanism of Cyclin K in lung cancer radioresistance. Methods: Expression levels of Cyclin K were measured by immunohistochemistry in human lung cancer tissues and adjacent normal lung tissues. Cell growth and proliferation, neutral comet and foci formation assays, G2/M checkpoint and a xenograft mouse model were used for functional analyses. Gene expression was examined by RNA sequencing and quantitative real-time PCR. -
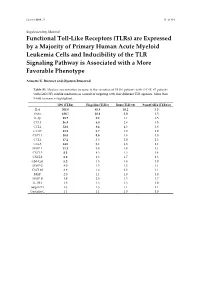
Are Expressed by a Majority of Primary Human Acute Myeloid Leukemia Cells and Inducibility of the TLR Signaling Pathway Is Associated with a More Favorable Phenotype
Cancers 2019, 11 S1 of S19 Supplementary Material Functional Toll-Like Receptors (TLRs) are Expressed by a Majority of Primary Human Acute Myeloid Leukemia Cells and Inducibility of the TLR Signaling Pathway is Associated with a More Favorable Phenotype Annette K. Brenner and Øystein Bruserud Table S1. Median concentration increase in the secretion of 19 (16 patients with G-CSF, 67 patients with GM-CSF) soluble mediators as a result of targeting with four different TLR agonists. More than 5-fold increase is highlighted. LPS (TLR4) Flagellin (TLR5) R848 (TLR7/8) Pam3CSK4 (TLR1/2) IL-6 301.0 45.9 10.2 5.3 TNFα 188.7 10.6 5.0 1.5 IL-1β 89.7 9.2 1.1 1.5 CCL3 56.4 6.0 2.4 1.5 CCL2 52.6 9.4 4.3 2.8 G-CSF 51.9 2.7 1.0 1.0 CXCL1 38.0 5.8 1.4 1.5 CCL4 17.2 3.3 2.0 1.3 CCL5 14.5 2.1 1.3 1.1 MMP-1 11.1 3.4 1.4 1.1 CXCL5 8.3 4.3 1.3 1.4 CXCL8 6.0 4.3 1.7 1.3 GM-CSF 5.2 1.5 1.4 1.0 MMP-2 4.0 1.5 1.2 1.1 CXCL10 3.9 1.6 2.2 1.1 HGF 2.0 1.1 1.0 1.0 MMP-9 1.9 2.5 1.3 1.7 IL-1RA 1.8 1.3 1.3 1.0 Serpin E1 1.3 1.3 1.1 1.1 Cystatin C 1.1 1.1 1.0 1.0 Cancers 2019, 11 S2 of S19 Table S2. -

Cytotoxic Effects and Changes in Gene Expression Profile
Toxicology in Vitro 34 (2016) 309–320 Contents lists available at ScienceDirect Toxicology in Vitro journal homepage: www.elsevier.com/locate/toxinvit Fusarium mycotoxin enniatin B: Cytotoxic effects and changes in gene expression profile Martina Jonsson a,⁎,MarikaJestoib, Minna Anthoni a, Annikki Welling a, Iida Loivamaa a, Ville Hallikainen c, Matti Kankainen d, Erik Lysøe e, Pertti Koivisto a, Kimmo Peltonen a,f a Chemistry and Toxicology Research Unit, Finnish Food Safety Authority (Evira), Mustialankatu 3, FI-00790 Helsinki, Finland b Product Safety Unit, Finnish Food Safety Authority (Evira), Mustialankatu 3, FI-00790 Helsinki, c The Finnish Forest Research Institute, Rovaniemi Unit, P.O. Box 16, FI-96301 Rovaniemi, Finland d Institute for Molecular Medicine Finland (FIMM), University of Helsinki, P.O. Box 20, FI-00014, Finland e Plant Health and Biotechnology, Norwegian Institute of Bioeconomy, Høyskoleveien 7, NO -1430 Ås, Norway f Finnish Safety and Chemicals Agency (Tukes), Opastinsilta 12 B, FI-00521 Helsinki, Finland article info abstract Article history: The mycotoxin enniatin B, a cyclic hexadepsipeptide produced by the plant pathogen Fusarium,isprevalentin Received 3 December 2015 grains and grain-based products in different geographical areas. Although enniatins have not been associated Received in revised form 5 April 2016 with toxic outbreaks, they have caused toxicity in vitro in several cell lines. In this study, the cytotoxic effects Accepted 28 April 2016 of enniatin B were assessed in relation to cellular energy metabolism, cell proliferation, and the induction of ap- Available online 6 May 2016 optosis in Balb 3T3 and HepG2 cells. The mechanism of toxicity was examined by means of whole genome ex- fi Keywords: pression pro ling of exposed rat primary hepatocytes. -

Human Cyclin-Dependent Kinase 12 (CDK12), Kinase Domain a Target Enabling Package (TEP)
Human Cyclin-Dependent Kinase 12 (CDK12), Kinase Domain A Target Enabling Package (TEP) Gene ID / UniProt ID / EC CDK12, 51755 / Q9NYV4/ 2.7.11.22, 2.7.11.23 CCNK, 8812 / O75909/ - Target Nominator Gregg Morin (UBC, Canada), Nathanael Gray (Harvard) SGC Authors Sarah E. Dixon-Clarke, Jonathan M. Elkins, Nathanael S. Gray, and Alex N. Bullock Collaborating Authors S.-W. Grace Cheng1, Tinghu Zhang2, Nicholas Kwiatkowski3, Calla M. Olson2, Brian J. Abraham3, Ann K. Greifenberg4, Scott B. Ficarro2, Yanke Liang2, Nancy M. Hannett3, Theresa Manz5, Mingfeng Hao2, Bartlomiej Bartkowiak6, Arno L. Greenleaf6, Jarrod A. Marto2, Matthias Geyer4, Richard A. Young3, and Gregg B. Morin1 Target PI Alex Bullock (SGC Oxford) Therapeutic Area(s) Oncology Disease Relevance CDK12 loss sensitises cancer cells to DNA damage Date approved by TEP 17th June 2016 Evaluation Group Document version Version 3 Document version date April 2018 DOI https://doi.org/10.5281/zenodo.1219680 Affiliations 1. Canada's Michael Smith Genome Sciences Centre, BC Cancer Agency 2. Department of Cancer Biology, Dana-Farber Cancer Institute 3. Whitehead Institute for Biomedical Research 4. Department of Structural Immunology, Institute of Innate Immunity 5. Pharmaceutical and Medicinal Chemistry, Saarland University 6. Department of Biochemistry, Duke University Medical Center USEFUL LINKS (Please note that the inclusion of links to external sites should not be taken as an endorsement of that site by the SGC in any way) SUMMARY OF PROJECT Cyclin-dependent kinase 12 (CDK12) phosphorylates RNA Pol II C-terminal domain (CTD) to promote transcriptional elongation of large DNA damage response genes. CDK12 is frequently mutated or amplified in cancer and its loss sensitises cells to DNA damage. -

Cyclin-Dependent Kinases and Their Role in Inflammation, Endothelial Cell Migration
Cyclin-Dependent Kinases and their role in Inflammation, Endothelial Cell Migration and Autocrine Activity Dissertation Presented in Partial Fulfillment of the Requirements for the Degree Doctor of Philosophy in the Graduate School of The Ohio State University By Shruthi Ratnakar Shetty Graduate Program in Pharmaceutical Sciences The Ohio State University 2020 Dissertation Committee Dale Hoyt, Advisor Liva Rakotondraibe Moray Campbell Keli Hu Copyrighted by Shruthi Ratnakar Shetty 2020 Abstract Inflammation is the body’s response to infection or injury. Endothelial cells are among the different players involved in an inflammatory cascade. In response to an inflammatory stimuli such as bacterial lipopolysaccharide (LPS), endothelial cells get activated which is characterized by the production of important mediators, such as inducible nitric oxide synthase (iNOS) which, catalyzes the production of nitric oxide (NO) and reactive nitrogen species and cyclooxygenase-2 (COX-2) that catalyzes the production of prostaglandins. Though the production of these mediators is required for an inflammatory response, it is important that their levels are regulated. Continued production of iNOS results in increased accumulation of reactive nitrogen species (RNS) that might lead to cytotoxicity, whereas lack of/suppression results in endothelial and vascular dysfunction. On the other hand, severe cardiovascular, intestinal and renal side effects are observed with significant suppression of COX-2. Thus, studying factors that could regulate the levels of iNOS and COX-2 could provide useful insights for developing novel therapeutic targets. Regulation of protein levels involves control of protein induction or turnover. Since protein induction requires transcription, in this dissertation we studied the role of a promoter of transcription “Cyclin- dependent kinase 7 (CDK7)” in iNOS and COX-2 protein induction. -

Targeting Cyclin-Dependent Kinase 9 and Myeloid Cell Leukaemia 1 in MYC-Driven B-Cell Lymphoma
Targeting cyclin-dependent kinase 9 and myeloid cell leukaemia 1 in MYC-driven B-cell lymphoma Gareth Peter Gregory ORCID ID: 0000-0002-4170-0682 Thesis for Doctor of Philosophy September 2016 Sir Peter MacCallum Department of Oncology The University of Melbourne Doctor of Philosophy Submitted in total fulfilment of the degree of Abstract Aggressive B-cell lymphomas include diffuse large B-cell lymphoma, Burkitt lymphoma and intermediate forms. Despite high response rates to conventional immuno-chemotherapeutic approaches, an unmet need for novel therapeutic by resistance to chemotherapy and radiotherapy. The proto-oncogene MYC is strategies is required in the setting of relapsed and refractory disease, typified frequently dysregulated in the aggressive B-cell lymphomas, however, it has proven an elusive direct therapeutic target. MYC-dysregulated disease maintains a ‘transcriptionally-addicted’ state, whereby perturbation of A significant body of evidence is accumulating to suggest that RNA polymerase II activity may indirectly antagonise MYC activity. Furthermore, very recent studies implicate anti-apoptotic myeloid cell leukaemia 1 (MCL-1) as a critical survival determinant of MYC-driven lymphoma. This thesis utilises pharmacologic and genetic techniques in MYC-driven models of aggressive B-cell lymphoma to demonstrate that cyclin-dependent kinase 9 (CDK9) and MCL-1 are oncogenic dependencies of this subset of disease. The cyclin-dependent kinase inhibitor, dinaciclib, and more selective CDK9 inhibitors downregulation of MCL1 are used -

The Genetic Complexity of Prostate Cancer
G C A T T A C G G C A T genes Review The Genetic Complexity of Prostate Cancer Eva Compérat 1,2,3,*, Gabriel Wasinger 3 , André Oszwald 3 , Renate Kain 3 , Geraldine Cancel-Tassin 1 and Olivier Cussenot 1,4 1 CeRePP/GRC5 Predictive Onco-Urology, Sorbonne University, 75020 Paris, France; [email protected] (G.C.-T.); [email protected] (O.C.) 2 Department of Pathology, Hôpital Tenon, Sorbonne University, 75020 Paris, France 3 Department of Pathology, Medical University of Vienna, 1090 Vienna, Austria; [email protected] (G.W.); [email protected] (A.O.); [email protected] (R.K.) 4 Department of Urology, Hôpital Tenon, Sorbonne University, 75020 Paris, France * Correspondence: [email protected]; Tel.: +33-658246024 Received: 28 September 2020; Accepted: 23 November 2020; Published: 25 November 2020 Abstract: Prostate cancer (PCa) is a major concern in public health, with many genetically distinct subsets. Genomic alterations in PCa are extraordinarily complex, and both germline and somatic mutations are of great importance in the development of this tumor. The aim of this review is to provide an overview of genetic changes that can occur in the development of PCa and their role in potential therapeutic approaches. Various pathways and mechanisms proposed to play major roles in PCa are described in detail to provide an overview of current knowledge. Keywords: prostate cancer; germline mutations; somatic mutations; PTEN; TMPRSS2; ERG; androgen receptors 1. Introduction Prostate cancer (PCa) is a major concern in public health, with more than 1.1 million cases worldwide detected every year [1]. -

Structural and Functional Analysis of the Cdk13/Cyclin K Complex
MASARYK UNIVERSITY FACULTY OF SCIENCE The role of the C-terminal domain of RNA polymerase II in transcriptionally regulated process of genomic instability Ph.D. Dissertation Květa Pilařová Supervisor: Mgr. Dalibor Blažek, Ph.D. Department of biochemistry Brno 2020 Bibliografický záznam Autor: Mgr. Květa Pilařová Přírodovědecká fakulta, Masarykova univerzita Ústav biochemie Název práce: Úloha C-terminální domény RNA polymerasy II v transkripčně regulovaném procesu genomové nestability Studijní program: Biochemie Vedoucí práce: Mgr. Dalibor Blažek, Ph.D. Akademický rok: 2019/2020 Počet stran: 132 Klíčová slova: CDK12, CDK13, kinázová aktivita, analog-senzitivní kináza, C- terminální doména RNAPII, transkripce, genová exprese, přechod mezi G1/S, genomová nestabilita, nádorový biomarker Bibliographic entry Author: Mgr. Květa Pilařová Faculty of Science, Masaryk University Department of biochemistry Title of thesis: The role of the C-terminal domain of RNA polymerase II in transcriptionally regulated process of genomic instability Degree programme: Biochemistry Supervisor: Mgr. Dalibor Blažek, Ph.D. Academic year: 2019/2020 Number of pages: 132 Keywords: CDK12, CDK13, kinase activity, analogue-sensitive kinase, C- terminal domain of RNAPII, transcription, gene expression, G1/S progression, genome instability, tumour biomarker Abstrakt Transkripce protein-kódujících genů je řízena v eukaryotických buňkách RNA polymerázou II (RNAPII). Cyklin-dependentní kinázy 12 a 13 (CDK12 a CDK13) se řadí do skupiny transkripčních CDKs, které asociují s RNAPII při elongaci a fosforylují její C-terminální doménu (CTD). Obě kinázy působí také na expresi genů. Abnormální funkce těchto proteinů jsou u lidských buněk spojovány s různými typy onemocnění a v posledním desetiletí se proto staly předmětem studia výzkumů v oblasti medicíny. Jakým způsobem se CDK12 a CDK13 konkrétně podílí na regulaci transkripce a fosforylačním statusu CTD však není příliš známo. -

Xo PANEL DNA GENE LIST
xO PANEL DNA GENE LIST ~1700 gene comprehensive cancer panel enriched for clinically actionable genes with additional biologically relevant genes (at 400 -500x average coverage on tumor) Genes A-C Genes D-F Genes G-I Genes J-L AATK ATAD2B BTG1 CDH7 CREM DACH1 EPHA1 FES G6PC3 HGF IL18RAP JADE1 LMO1 ABCA1 ATF1 BTG2 CDK1 CRHR1 DACH2 EPHA2 FEV G6PD HIF1A IL1R1 JAK1 LMO2 ABCB1 ATM BTG3 CDK10 CRK DAXX EPHA3 FGF1 GAB1 HIF1AN IL1R2 JAK2 LMO7 ABCB11 ATR BTK CDK11A CRKL DBH EPHA4 FGF10 GAB2 HIST1H1E IL1RAP JAK3 LMTK2 ABCB4 ATRX BTRC CDK11B CRLF2 DCC EPHA5 FGF11 GABPA HIST1H3B IL20RA JARID2 LMTK3 ABCC1 AURKA BUB1 CDK12 CRTC1 DCUN1D1 EPHA6 FGF12 GALNT12 HIST1H4E IL20RB JAZF1 LPHN2 ABCC2 AURKB BUB1B CDK13 CRTC2 DCUN1D2 EPHA7 FGF13 GATA1 HLA-A IL21R JMJD1C LPHN3 ABCG1 AURKC BUB3 CDK14 CRTC3 DDB2 EPHA8 FGF14 GATA2 HLA-B IL22RA1 JMJD4 LPP ABCG2 AXIN1 C11orf30 CDK15 CSF1 DDIT3 EPHB1 FGF16 GATA3 HLF IL22RA2 JMJD6 LRP1B ABI1 AXIN2 CACNA1C CDK16 CSF1R DDR1 EPHB2 FGF17 GATA5 HLTF IL23R JMJD7 LRP5 ABL1 AXL CACNA1S CDK17 CSF2RA DDR2 EPHB3 FGF18 GATA6 HMGA1 IL2RA JMJD8 LRP6 ABL2 B2M CACNB2 CDK18 CSF2RB DDX3X EPHB4 FGF19 GDNF HMGA2 IL2RB JUN LRRK2 ACE BABAM1 CADM2 CDK19 CSF3R DDX5 EPHB6 FGF2 GFI1 HMGCR IL2RG JUNB LSM1 ACSL6 BACH1 CALR CDK2 CSK DDX6 EPOR FGF20 GFI1B HNF1A IL3 JUND LTK ACTA2 BACH2 CAMTA1 CDK20 CSNK1D DEK ERBB2 FGF21 GFRA4 HNF1B IL3RA JUP LYL1 ACTC1 BAG4 CAPRIN2 CDK3 CSNK1E DHFR ERBB3 FGF22 GGCX HNRNPA3 IL4R KAT2A LYN ACVR1 BAI3 CARD10 CDK4 CTCF DHH ERBB4 FGF23 GHR HOXA10 IL5RA KAT2B LZTR1 ACVR1B BAP1 CARD11 CDK5 CTCFL DIAPH1 ERCC1 FGF3 GID4 HOXA11 -

The Role of Cdks and Cdkis in Murine Development
International Journal of Molecular Sciences Review The Role of CDKs and CDKIs in Murine Development Grace Jean Campbell , Emma Langdale Hands and Mathew Van de Pette * Epigenetic Mechanisms of Toxicology Lab, MRC Toxicology Unit, Cambridge University, Cambridge CB2 1QR, UK; [email protected] (G.J.C.); [email protected] (E.L.H.) * Correspondence: [email protected] Received: 8 July 2020; Accepted: 26 July 2020; Published: 28 July 2020 Abstract: Cyclin-dependent kinases (CDKs) and their inhibitors (CDKIs) play pivotal roles in the regulation of the cell cycle. As a result of these functions, it may be extrapolated that they are essential for appropriate embryonic development. The twenty known mouse CDKs and eight CDKIs have been studied to varying degrees in the developing mouse, but only a handful of CDKs and a single CDKI have been shown to be absolutely required for murine embryonic development. What has become apparent, as more studies have shone light on these family members, is that in addition to their primary functional role in regulating the cell cycle, many of these genes are also controlling specific cell fates by directing differentiation in various tissues. Here we review the extensive mouse models that have been generated to study the functions of CDKs and CDKIs, and discuss their varying roles in murine embryonic development, with a particular focus on the brain, pancreas and fertility. Keywords: cyclin-dependent kinase; CDK inhibitors; mouse; development; knock-out models 1. Introduction Cyclin-dependent kinases (CDKs) are proteins that, by definition, require the binding of partner cyclin proteins in order to phosphorylate a series of target proteins.