Structural and Functional Analysis of the Cdk13/Cyclin K Complex
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Investigating the Role of Cdk11in Animal Cytokinesis
Investigating the Role of CDK11 in Animal Cytokinesis by Thomas Clifford Panagiotou A thesis submitted in conformity with the requirements for the degree of Master of Science Department of Molecular Genetics University of Toronto © Copyright by Thomas Clifford Panagiotou (2020) Investigating the Role of CDK11 in Animal Cytokinesis Thomas Clifford Panagiotou Master of Science Department of Molecular Genetics University of Toronto 2020 Abstract Finely tuned spatio-temporal regulation of cell division is required for genome stability. Cytokinesis constitutes the final stages of cell division, from chromosome segregation to the physical separation of cells, abscission. Abscission is tightly regulated to ensure it occurs after earlier cytokinetic events, like the maturation of the stem body, the regulatory platform for abscission. Active Aurora B kinase enforces the abscission checkpoint, which blocks abscission until chromosomes have been cleared from the cytokinetic machinery. Currently, it is unclear how this checkpoint is overcome. Here, I demonstrate that the cyclin-dependent kinase CDK11 is required for cytokinesis. Both inhibition and depletion of CDK11 block abscission. Furthermore, the mitosis-specific CDK11p58 kinase localizes to the stem body, where its kinase activity rescues the defects of CDK11 depletion and inhibition. These results suggest a model whereby CDK11p58 antagonizes Aurora B kinase to overcome the abscission checkpoint to allow for successful completion of cytokinesis. ii Acknowledgments I am very grateful for the support of my family and friends throughout my studies. I would also like to express my deep gratitude to Wilde Lab members, both past and present, for their advice and collaboration. In particular, I am very grateful to Matthew Renshaw, whose work comprises part of this thesis. -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

Cyclin K Interacts with Β-Catenin to Induce Cyclin D1 Expression And
Theranostics 2020, Vol. 10, Issue 24 11144 Ivyspring International Publisher Theranostics 2020; 10(24): 11144-11158. doi: 10.7150/thno.42578 Research Paper Cyclin K interacts with β-catenin to induce Cyclin D1 expression and facilitates tumorigenesis and radioresistance in lung cancer Guojun Yao*, Jing Tang*, Xijie Yang, Ye Zhao, Rui Zhou, Rui Meng, Sheng Zhang, Xiaorong Dong, Tao Zhang, Kunyu Yang, Gang Wu and Shuangbing Xu Cancer Center, Union Hospital, Tongji Medical College, Huazhong University of Science and Technology, Wuhan 430022, China. *These authors contributed equally to this work. Corresponding author: Shuangbing Xu or Gang Wu, Cancer Center, Union Hospital, Tongji Medical College, Huazhong University of Science and Technology, Wuhan 430022, China. E-mail: [email protected] or [email protected]. © The author(s). This is an open access article distributed under the terms of the Creative Commons Attribution License (https://creativecommons.org/licenses/by/4.0/). See http://ivyspring.com/terms for full terms and conditions. Received: 2019.11.29; Accepted: 2020.08.24; Published: 2020.09.11 Abstract Rationale: Radioresistance remains the major cause of local relapse and distant metastasis in lung cancer. However, the underlying molecular mechanisms remain poorly defined. This study aimed to investigate the role and regulatory mechanism of Cyclin K in lung cancer radioresistance. Methods: Expression levels of Cyclin K were measured by immunohistochemistry in human lung cancer tissues and adjacent normal lung tissues. Cell growth and proliferation, neutral comet and foci formation assays, G2/M checkpoint and a xenograft mouse model were used for functional analyses. Gene expression was examined by RNA sequencing and quantitative real-time PCR. -
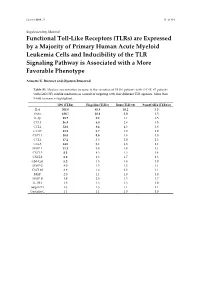
Are Expressed by a Majority of Primary Human Acute Myeloid Leukemia Cells and Inducibility of the TLR Signaling Pathway Is Associated with a More Favorable Phenotype
Cancers 2019, 11 S1 of S19 Supplementary Material Functional Toll-Like Receptors (TLRs) are Expressed by a Majority of Primary Human Acute Myeloid Leukemia Cells and Inducibility of the TLR Signaling Pathway is Associated with a More Favorable Phenotype Annette K. Brenner and Øystein Bruserud Table S1. Median concentration increase in the secretion of 19 (16 patients with G-CSF, 67 patients with GM-CSF) soluble mediators as a result of targeting with four different TLR agonists. More than 5-fold increase is highlighted. LPS (TLR4) Flagellin (TLR5) R848 (TLR7/8) Pam3CSK4 (TLR1/2) IL-6 301.0 45.9 10.2 5.3 TNFα 188.7 10.6 5.0 1.5 IL-1β 89.7 9.2 1.1 1.5 CCL3 56.4 6.0 2.4 1.5 CCL2 52.6 9.4 4.3 2.8 G-CSF 51.9 2.7 1.0 1.0 CXCL1 38.0 5.8 1.4 1.5 CCL4 17.2 3.3 2.0 1.3 CCL5 14.5 2.1 1.3 1.1 MMP-1 11.1 3.4 1.4 1.1 CXCL5 8.3 4.3 1.3 1.4 CXCL8 6.0 4.3 1.7 1.3 GM-CSF 5.2 1.5 1.4 1.0 MMP-2 4.0 1.5 1.2 1.1 CXCL10 3.9 1.6 2.2 1.1 HGF 2.0 1.1 1.0 1.0 MMP-9 1.9 2.5 1.3 1.7 IL-1RA 1.8 1.3 1.3 1.0 Serpin E1 1.3 1.3 1.1 1.1 Cystatin C 1.1 1.1 1.0 1.0 Cancers 2019, 11 S2 of S19 Table S2. -

Cytotoxic Effects and Changes in Gene Expression Profile
Toxicology in Vitro 34 (2016) 309–320 Contents lists available at ScienceDirect Toxicology in Vitro journal homepage: www.elsevier.com/locate/toxinvit Fusarium mycotoxin enniatin B: Cytotoxic effects and changes in gene expression profile Martina Jonsson a,⁎,MarikaJestoib, Minna Anthoni a, Annikki Welling a, Iida Loivamaa a, Ville Hallikainen c, Matti Kankainen d, Erik Lysøe e, Pertti Koivisto a, Kimmo Peltonen a,f a Chemistry and Toxicology Research Unit, Finnish Food Safety Authority (Evira), Mustialankatu 3, FI-00790 Helsinki, Finland b Product Safety Unit, Finnish Food Safety Authority (Evira), Mustialankatu 3, FI-00790 Helsinki, c The Finnish Forest Research Institute, Rovaniemi Unit, P.O. Box 16, FI-96301 Rovaniemi, Finland d Institute for Molecular Medicine Finland (FIMM), University of Helsinki, P.O. Box 20, FI-00014, Finland e Plant Health and Biotechnology, Norwegian Institute of Bioeconomy, Høyskoleveien 7, NO -1430 Ås, Norway f Finnish Safety and Chemicals Agency (Tukes), Opastinsilta 12 B, FI-00521 Helsinki, Finland article info abstract Article history: The mycotoxin enniatin B, a cyclic hexadepsipeptide produced by the plant pathogen Fusarium,isprevalentin Received 3 December 2015 grains and grain-based products in different geographical areas. Although enniatins have not been associated Received in revised form 5 April 2016 with toxic outbreaks, they have caused toxicity in vitro in several cell lines. In this study, the cytotoxic effects Accepted 28 April 2016 of enniatin B were assessed in relation to cellular energy metabolism, cell proliferation, and the induction of ap- Available online 6 May 2016 optosis in Balb 3T3 and HepG2 cells. The mechanism of toxicity was examined by means of whole genome ex- fi Keywords: pression pro ling of exposed rat primary hepatocytes. -

Human Cyclin-Dependent Kinase 12 (CDK12), Kinase Domain a Target Enabling Package (TEP)
Human Cyclin-Dependent Kinase 12 (CDK12), Kinase Domain A Target Enabling Package (TEP) Gene ID / UniProt ID / EC CDK12, 51755 / Q9NYV4/ 2.7.11.22, 2.7.11.23 CCNK, 8812 / O75909/ - Target Nominator Gregg Morin (UBC, Canada), Nathanael Gray (Harvard) SGC Authors Sarah E. Dixon-Clarke, Jonathan M. Elkins, Nathanael S. Gray, and Alex N. Bullock Collaborating Authors S.-W. Grace Cheng1, Tinghu Zhang2, Nicholas Kwiatkowski3, Calla M. Olson2, Brian J. Abraham3, Ann K. Greifenberg4, Scott B. Ficarro2, Yanke Liang2, Nancy M. Hannett3, Theresa Manz5, Mingfeng Hao2, Bartlomiej Bartkowiak6, Arno L. Greenleaf6, Jarrod A. Marto2, Matthias Geyer4, Richard A. Young3, and Gregg B. Morin1 Target PI Alex Bullock (SGC Oxford) Therapeutic Area(s) Oncology Disease Relevance CDK12 loss sensitises cancer cells to DNA damage Date approved by TEP 17th June 2016 Evaluation Group Document version Version 3 Document version date April 2018 DOI https://doi.org/10.5281/zenodo.1219680 Affiliations 1. Canada's Michael Smith Genome Sciences Centre, BC Cancer Agency 2. Department of Cancer Biology, Dana-Farber Cancer Institute 3. Whitehead Institute for Biomedical Research 4. Department of Structural Immunology, Institute of Innate Immunity 5. Pharmaceutical and Medicinal Chemistry, Saarland University 6. Department of Biochemistry, Duke University Medical Center USEFUL LINKS (Please note that the inclusion of links to external sites should not be taken as an endorsement of that site by the SGC in any way) SUMMARY OF PROJECT Cyclin-dependent kinase 12 (CDK12) phosphorylates RNA Pol II C-terminal domain (CTD) to promote transcriptional elongation of large DNA damage response genes. CDK12 is frequently mutated or amplified in cancer and its loss sensitises cells to DNA damage. -

Human Cyclin-Dependent Kinase 12 (CDK12), Kinase Domain a Target Enabling Package (TEP)
Human Cyclin-Dependent Kinase 12 (CDK12), Kinase Domain A Target Enabling Package (TEP) Gene ID / UniProt ID / EC CDK12, 51755 / Q9NYV4/ 2.7.11.22, 2.7.11.23 CCNK, 8812 / O75909/ - Target Nominator Gregg Morin (UBC, Canada), Nathanael Gray (Harvard) SGC Authors Sarah E. Dixon-Clarke, Jonathan M. Elkins, and Alex N. Bullock Collaborating Authors S.-W. Grace Cheng1, Tinghu Zhang2, Nicholas Kwiatkowski3, Calla M. Olson2, Brian J. Abraham3, Ann K. Greifenberg4, Scott B. Ficarro2, Yanke Liang2, Nancy M. Hannett3, Theresa Manz5, Mingfeng Hao2, Bartlomiej Bartkowiak6, Arno L. Greenleaf6, Jarrod A. Marto2, Matthias Geyer4, Richard A. Young3, Gregg B. Morin1 and Nathanael S. Gray2 Target PI Alex Bullock (SGC Oxford) Therapeutic Area(s) Oncology Disease Relevance CDK12 loss sensitises cancer cells to DNA damage Date approved by TEP 17th June 2016 Evaluation Group Document version Version 9 Document version date March 3rd 2021 Citation Sarah E. Dixon-Clarke, Jonathan M. Elkins, S.-W. Grace Cheng, Tinghu Zhang, Nicholas Kwiatkowski, Calla M. Olson, Brian J. Abraham, Ann K. Greifenberg, Scott B. Ficarro, Yanke Liang, Nancy M. Hannett, Theresa Manz, Mingfeng Hao, Bartlomiej Bartkowiak, Arno L. Greenleaf, Jarrod A. Marto, Matthias Geyer, Richard A. Young, Gregg B. Morin, Nathanael S. Gray and Alex N. Bullock. (2016) Human Cyclin-dependent kinase 12 (CDK12), kinase domain; A Target Enabling Package. 10.5281/zenodo.1219680. Affiliations 1. Canada's Michael Smith Genome Sciences Centre, BC Cancer Agency; 2. Department of Cancer Biology, Dana-Farber Cancer Institute; 3. Whitehead Institute for Biomedical Research; 4. Department of Structural Immunology, Institute of Innate Immunity; 5. Pharmaceutical and Medicinal Chemistry, Saarland University; 6. -

Xo PANEL DNA GENE LIST
xO PANEL DNA GENE LIST ~1700 gene comprehensive cancer panel enriched for clinically actionable genes with additional biologically relevant genes (at 400 -500x average coverage on tumor) Genes A-C Genes D-F Genes G-I Genes J-L AATK ATAD2B BTG1 CDH7 CREM DACH1 EPHA1 FES G6PC3 HGF IL18RAP JADE1 LMO1 ABCA1 ATF1 BTG2 CDK1 CRHR1 DACH2 EPHA2 FEV G6PD HIF1A IL1R1 JAK1 LMO2 ABCB1 ATM BTG3 CDK10 CRK DAXX EPHA3 FGF1 GAB1 HIF1AN IL1R2 JAK2 LMO7 ABCB11 ATR BTK CDK11A CRKL DBH EPHA4 FGF10 GAB2 HIST1H1E IL1RAP JAK3 LMTK2 ABCB4 ATRX BTRC CDK11B CRLF2 DCC EPHA5 FGF11 GABPA HIST1H3B IL20RA JARID2 LMTK3 ABCC1 AURKA BUB1 CDK12 CRTC1 DCUN1D1 EPHA6 FGF12 GALNT12 HIST1H4E IL20RB JAZF1 LPHN2 ABCC2 AURKB BUB1B CDK13 CRTC2 DCUN1D2 EPHA7 FGF13 GATA1 HLA-A IL21R JMJD1C LPHN3 ABCG1 AURKC BUB3 CDK14 CRTC3 DDB2 EPHA8 FGF14 GATA2 HLA-B IL22RA1 JMJD4 LPP ABCG2 AXIN1 C11orf30 CDK15 CSF1 DDIT3 EPHB1 FGF16 GATA3 HLF IL22RA2 JMJD6 LRP1B ABI1 AXIN2 CACNA1C CDK16 CSF1R DDR1 EPHB2 FGF17 GATA5 HLTF IL23R JMJD7 LRP5 ABL1 AXL CACNA1S CDK17 CSF2RA DDR2 EPHB3 FGF18 GATA6 HMGA1 IL2RA JMJD8 LRP6 ABL2 B2M CACNB2 CDK18 CSF2RB DDX3X EPHB4 FGF19 GDNF HMGA2 IL2RB JUN LRRK2 ACE BABAM1 CADM2 CDK19 CSF3R DDX5 EPHB6 FGF2 GFI1 HMGCR IL2RG JUNB LSM1 ACSL6 BACH1 CALR CDK2 CSK DDX6 EPOR FGF20 GFI1B HNF1A IL3 JUND LTK ACTA2 BACH2 CAMTA1 CDK20 CSNK1D DEK ERBB2 FGF21 GFRA4 HNF1B IL3RA JUP LYL1 ACTC1 BAG4 CAPRIN2 CDK3 CSNK1E DHFR ERBB3 FGF22 GGCX HNRNPA3 IL4R KAT2A LYN ACVR1 BAI3 CARD10 CDK4 CTCF DHH ERBB4 FGF23 GHR HOXA10 IL5RA KAT2B LZTR1 ACVR1B BAP1 CARD11 CDK5 CTCFL DIAPH1 ERCC1 FGF3 GID4 HOXA11 -

The Role of Cdks and Cdkis in Murine Development
International Journal of Molecular Sciences Review The Role of CDKs and CDKIs in Murine Development Grace Jean Campbell , Emma Langdale Hands and Mathew Van de Pette * Epigenetic Mechanisms of Toxicology Lab, MRC Toxicology Unit, Cambridge University, Cambridge CB2 1QR, UK; [email protected] (G.J.C.); [email protected] (E.L.H.) * Correspondence: [email protected] Received: 8 July 2020; Accepted: 26 July 2020; Published: 28 July 2020 Abstract: Cyclin-dependent kinases (CDKs) and their inhibitors (CDKIs) play pivotal roles in the regulation of the cell cycle. As a result of these functions, it may be extrapolated that they are essential for appropriate embryonic development. The twenty known mouse CDKs and eight CDKIs have been studied to varying degrees in the developing mouse, but only a handful of CDKs and a single CDKI have been shown to be absolutely required for murine embryonic development. What has become apparent, as more studies have shone light on these family members, is that in addition to their primary functional role in regulating the cell cycle, many of these genes are also controlling specific cell fates by directing differentiation in various tissues. Here we review the extensive mouse models that have been generated to study the functions of CDKs and CDKIs, and discuss their varying roles in murine embryonic development, with a particular focus on the brain, pancreas and fertility. Keywords: cyclin-dependent kinase; CDK inhibitors; mouse; development; knock-out models 1. Introduction Cyclin-dependent kinases (CDKs) are proteins that, by definition, require the binding of partner cyclin proteins in order to phosphorylate a series of target proteins. -

Modulating the Unfolded Protein Response with ONC201 to Impact
www.nature.com/scientificreports OPEN Modulating the unfolded protein response with ONC201 to impact on radiation response in prostate cancer cells Francesca Amoroso1,7*, Kimberley Glass1, Reema Singh7, Francisco Liberal1,2, Rebecca E. Steele1,3, Sarah Maguire1, Rohinton Tarapore4, Joshua E. Allen4, Sandra Van Schaeybroeck1, Karl T. Butterworth1, Kevin Prise1, Joe M. O’Sullivan1,5, Suneil Jain1,5, David J. Waugh6 & Ian G. Mills1,7,8,9* Prostate cancer (PCa) is the most common non-cutaneous cancer in men and a notable cause of cancer mortality when it metastasises. The unfolded protein response (UPR) can be cytoprotective but when acutely activated can lead to cell death. In this study, we sought to enhance the acute activation of the UPR using radiation and ONC201, an UPR activator. Treating PCa cells with ONC201 quickly increased the expression of all the key regulators of the UPR and reduced the oxidative phosphorylation, with cell death occurring 72 h later. We exploited this time lag to sensitize prostate cancer cells to radiation through short-term treatment with ONC201. To understand how priming occurred, we performed RNA-Seq analysis and found that ONC201 suppressed the expression of cell cycle and DNA repair factors. In conclusion, we have shown that ONC201 can prime enhanced radiation response. Prostate cancer (PCa) is the most common cancer diagnosed in men and the second most common cause of cancer death afer lung cancer. According to recent projections, prostate cancer incidence rates are predict to rise by 12% in the UK between 2014 and 2035, to 233 cases per 100,000 males by 20351. -

Targeting Cyclin-Dependent Kinases in Human Cancers: from Small Molecules to Peptide Inhibitors
Cancers 2015, 7, 179-237; doi:10.3390/cancers7010179 OPEN ACCESS cancers ISSN 2072-6694 www.mdpi.com/journal/cancers Review Targeting Cyclin-Dependent Kinases in Human Cancers: From Small Molecules to Peptide Inhibitors Marion Peyressatre †, Camille Prével †, Morgan Pellerano and May C. Morris * Institut des Biomolécules Max Mousseron, IBMM-CNRS-UMR5247, 15 Av. Charles Flahault, 34093 Montpellier, France; E-Mails: [email protected] (M.P.); [email protected] (C.P.); [email protected] (M.P.) † These authors contributed equally to this work. * Author to whom correspondence should be addressed; E-Mail: [email protected]; Tel.: +33-04-1175-9624; Fax: +33-04-1175-9641. Academic Editor: Jonas Cicenas Received: 17 December 2014 / Accepted: 12 January 2015 / Published: 23 January 2015 Abstract: Cyclin-dependent kinases (CDK/Cyclins) form a family of heterodimeric kinases that play central roles in regulation of cell cycle progression, transcription and other major biological processes including neuronal differentiation and metabolism. Constitutive or deregulated hyperactivity of these kinases due to amplification, overexpression or mutation of cyclins or CDK, contributes to proliferation of cancer cells, and aberrant activity of these kinases has been reported in a wide variety of human cancers. These kinases therefore constitute biomarkers of proliferation and attractive pharmacological targets for development of anticancer therapeutics. The structural features of several of these kinases have been elucidated and their molecular mechanisms of regulation characterized in depth, providing clues for development of drugs and inhibitors to disrupt their function. However, like most other kinases, they constitute a challenging class of therapeutic targets due to their highly conserved structural features and ATP-binding pocket. -

Clinical, Molecular, and Immune Analysis of Dabrafenib-Trametinib
Supplementary Online Content Chen G, McQuade JL, Panka DJ, et al. Clinical, molecular and immune analysis of dabrafenib-trametinib combination treatment for metastatic melanoma that progressed during BRAF inhibitor monotherapy: a phase 2 clinical trial. JAMA Oncology. Published online April 28, 2016. doi:10.1001/jamaoncol.2016.0509. eMethods. eReferences. eTable 1. Clinical efficacy eTable 2. Adverse events eTable 3. Correlation of baseline patient characteristics with treatment outcomes eTable 4. Patient responses and baseline IHC results eFigure 1. Kaplan-Meier analysis of overall survival eFigure 2. Correlation between IHC and RNAseq results eFigure 3. pPRAS40 expression and PFS eFigure 4. Baseline and treatment-induced changes in immune infiltrates eFigure 5. PD-L1 expression eTable 5. Nonsynonymous mutations detected by WES in baseline tumors This supplementary material has been provided by the authors to give readers additional information about their work. © 2016 American Medical Association. All rights reserved. Downloaded From: https://jamanetwork.com/ on 09/30/2021 eMethods Whole exome sequencing Whole exome capture libraries for both tumor and normal samples were constructed using 100ng genomic DNA input and following the protocol as described by Fisher et al.,3 with the following adapter modification: Illumina paired end adapters were replaced with palindromic forked adapters with unique 8 base index sequences embedded within the adapter. In-solution hybrid selection was performed using the Illumina Rapid Capture Exome enrichment kit with 38Mb target territory (29Mb baited). The targeted region includes 98.3% of the intervals in the Refseq exome database. Dual-indexed libraries were pooled into groups of up to 96 samples prior to hybridization.