NEST Desktop
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-
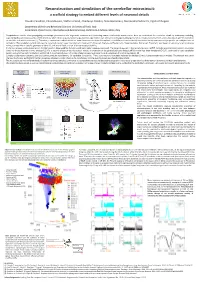
Reconstruction and Simulation of the Cerebellar Microcircuit: a Scaffold Strategy to Embed Different Levels of Neuronal Details
Reconstruction and simulation of the cerebellar microcircuit: a scaffold strategy to embed different levels of neuronal details Claudia Casellato, Elisa Marenzi, Stefano Casali, Chaitanya Medini, Alice Geminiani, Alessandra Pedrocchi, Egidio D’Angelo Department of Brain and Behavioral Sciences, University of Pavia, Italy Department of Electronics, Information and Bioengineering, Politecnico di Milano, Milan, Italy Computational models allow propagating microscopic phenomena into large‐scale networks and inferencing causal relationships across scales. Here we reconstruct the cerebellar circuit by bottom‐up modeling, reproducing the peculiar properties of this structure, which shows a quasi‐crystalline geometrical organization well defined by convergence/divergence ratios of neuronal connections and by the anisotropic 3D orientation of dendritic and axonal processes [1]. Therefore, a cerebellum scaffold model has been developed and tested. It maintains scalability and can be flexibly handled to incorporate neuronal properties on multiple scales of complexity. The cerebellar scaffold includes the canonical neuron types: Granular cell, Golgi cell, Purkinje cell, Stellate and Basket cells, Deep Cerebellar Nuclei cell. Placement was based on density and encumbrance values, connectivity on specific geometry of dendritic and axonal fields, and on distance‐based probability. In the first release, spiking point‐neuron models based on Integrate&Fire dynamics with exponential synapses were used. The network was run in the neural simulator pyNEST. Complex spatiotemporal patterns of activity, similar to those observed in vivo, emerged [2]. For a second release of the microcircuit model, an extension of the generalized Leaky Integrate&Fire model has been developed (E‐GLIF), optimized for each cerebellar neuron type and inserted into the built scaffold [3]. -

The Human Brain Project—Synergy Between Neuroscience, Computing, Informatics, and Brain-Inspired Technologies
COMMUNITY PAGE The Human Brain ProjectÐSynergy between neuroscience, computing, informatics, and brain-inspired technologies 1,2 3 4 5 Katrin AmuntsID *, Alois C. Knoll , Thomas LippertID , Cyriel M. A. PennartzID , 6 7 8 9 10 Philippe RyvlinID , Alain DestexheID , Viktor K. Jirsa , Egidio D'Angelo , Jan G. BjaalieID 1 Institute for Neuroscience and Medicine (INM-1), Forschungszentrum JuÈlich, Germany, 2 C. and O. Vogt Institute for Brain Research, University Hospital DuÈsseldorf, Heinrich Heine University DuÈsseldorf, DuÈsseldorf, Germany, 3 Institut fuÈr Informatik VI, Technische UniversitaÈt MuÈnchen, Garching bei MuÈnchen, a1111111111 Germany, 4 JuÈlich Supercomputing Centre, Institute for Advanced Simulation, Forschungszentrum JuÈlich, a1111111111 Germany, 5 Swammerdam Institute for Life Sciences, Faculty of Science, University of Amsterdam, the a1111111111 Netherlands, 6 Department of Clinical Neurosciences, Centre Hospitalo-Universitaire Vaudois (CHUV) and a1111111111 University of Lausanne, Lausanne, Switzerland, 7 Unite de Neurosciences, Information & Complexite a1111111111 (UNIC), Centre National de la Recherche Scientifique (CNRS), Gif-sur-Yvette, France, 8 Institut de Neurosciences des Systèmes, Inserm UMR1106, Aix-Marseille UniversiteÂ, Faculte de MeÂdecine, Marseille, France, 9 Department of Brain and Behavioral Science, Unit of Neurophysiology, University of Pavia, Pavia, Italy, 10 Institute of Basic Medical Sciences, University of Oslo, Oslo, Norway * [email protected] OPEN ACCESS Citation: Amunts K, Knoll AC, Lippert T, Pennartz CMA, Ryvlin P, Destexhe A, et al. (2019) The Abstract Human Brain ProjectÐSynergy between neuroscience, computing, informatics, and brain- The Human Brain Project (HBP) is a European flagship project with a 10-year horizon aim- inspired technologies. PLoS Biol 17(7): e3000344. ing to understand the human brain and to translate neuroscience knowledge into medicine https://doi.org/10.1371/journal.pbio.3000344 and technology. -

Simulation with NEST, an Example of a Full-Scale Spiking Neuronal Network Model - Seminar Paper
Simulation with NEST, an example of a full-scale spiking neuronal network model - seminar paper Till Schumann, CES, 293576 Computational and Systems Neuroscience (INM-6) 1 Introduction to Computational Neuroscience 1.1 Motivation Computational neuroscience is part of the computational biology, which, besides other meth- ods, relies on modeling to understand various aspects of biological systems. Computational neuroscience itself focuses on the nervous system. It is a growing field of research. With the fast development of computer systems and the growing availability of experimental data, computational simulations get more important. The computational power, which is available now and will be available in the next years, allows simulations of mammalian brains. Even a simulation of the human brain seems to be doable in the upcoming years. Modeling nervous systems helps us to understand the functionality of the human brain. It can help us to un- derstand different kinds of diseases like Alzheimer's and can help to develop novel therapies. Since the human brain is not accessible to direct experimental studies models of the brain are essential to understand its functionality. Computational modeling allows to construct neuron models that are based on cell-level data obtained from experiments. In contrast to the human, brain experimental data from animal experiments are widely available. Because of related structures these data is used to improve and validate models of the brain. In combination with neuronal simulations these data allows a first look into the functionality of nervous sys- tems. The paper The cell-type specific cortical microcircuit: relating structure and activity in a full-scale spiking network model shows the usability of current models and simulation tools. -

NEST by Example: an Introduction to the Neural Simulation Tool NEST Version 2.6.0
NEST by Example: An Introduction to the Neural Simulation Tool NEST Version 2.6.0 Marc-Oliver Gewaltig1, Abigail Morrison2, and Hans Ekkehard Plesser3, 2 1Blue Brain Project, Ecole Polytechnique Federale de Lausanne, QI-J, Lausanne 1015, Switzerland 2Institute of Neuroscience and Medicine (INM-6) Functional Neural Circuits Group, J¨ulich Research Center, 52425 J¨ulich, Germany 3Dept of Mathematical Sciences and Technology, Norwegian University of Life Sciences, PO Box 5003, 1432 Aas, Norway Abstract The neural simulation tool NEST can simulate small to very large networks of point-neurons or neurons with a few compartments. In this chapter, we show by example how models are programmed and simulated in NEST. This document is based on a preprint version of Gewaltig et al. (2012) and has been updated for NEST 2.6 (r11744). Updated to 2.6.0 Hans E. Plesser, December 2014 Updated to 2.4.0 Hans E. Plesser, June 2014 Updated to 2.2.2 Hans E. Plesser & Marc-Oliver Gewaltig, December 2012 1 Introduction NEST is a simulator for networks of point neurons, that is, neuron models that collapse the morphol- ogy (geometry) of dendrites, axons, and somata into either a single compartment or a small number of compartments (Gewaltig and Diesmann, 2007). This simplification is useful for questions about the dynamics of large neuronal networks with complex connectivity. In this text, we give a practical intro- duction to neural simulations with NEST. We describe how network models are defined and simulated, how simulations can be run in parallel, using multiple cores or computer clusters, and how parts of a model can be randomized. -

THE NEURAL SIMULATION TOOL NEST Nd HPAC Platform Training
THE NEURAL SIMULATION TOOL NEST 2nd HPAC Platform Training November 25, 2019 Jochen M. Eppler ([email protected]) SimLab Neuroscience Member of the Helmholtz Association OUTLINE Introduction Neuronal simulations Technological background Developing new models Performance This presentation is provided under the terms of the Creative Commons Attribution-ShareAlike License 4.0. Member of the Helmholtz Association November 25, 2019 Slide 1 NEST = NEURAL SIMULATION TOOL Point neurons and neurons with few electrical compartments Phenomenological synapse models (STDP, STP) + gap junctions, neuromodulation and structural plasticity Frameworks for rate models and binary neurons Support for neuroscience interfaces (MUSIC, libneurosim) Highly efficient C++ core with a Python frontend Hybrid parallelization (OpenMP+MPI) Same code from laptops to supercomputers Member of the Helmholtz Association November 25, 2019 Slide 2 NEST DESIGN GOALS High accuracy and flexibility Each neuron model is assigned an appropriate solver Exact integration is used for suitable neuron models Spikes are usually restricted to the computation time grid Spike interaction in continuous time for some models Constant quality assurance Automated unittest suite included in NEST build Continuous integration for all repository checkins Code review for all code contributions NEST’s development is always driven by scientific needs Member of the Helmholtz Association November 25, 2019 Slide 3 WHEN TO USE NEST? Growth Regulator/ Glu Glu factor Hormone Point neuron RTK mGluR GPCR NMDAR -

An Overview of Mind Uploading
IOSR Journal of Engineering (IOSR JEN) www.iosrjen.org ISSN (e): 2250-3021, ISSN (p): 2278-8719 PP 18-25 An Overview of Mind Uploading Mrs. Mausami Sawarkar1, Mr. Dhiraj Rane2 Dept of CSE, Priydarshani J L CoE, Nagpur Dept. of CS, GHRIIT, Nagpur Abstract: Mind uploading is an ongoing area of active research, bringing together ideas from neuroscience, computer science, engineering, and philosophy. Realistically, mind uploading likely lies many decades in the future, but the short-term offers the possibility of advanced neural prostheses that may benefit us. Mind uploading is the conceptual futuristic technology of transferring human minds tocomputer hardware using whole-brain emulation process. In this research work, we have discuss the brief review of the technological prospectsfor mind uploading, a range of philosophical and ethical aspects of the technology. We have also summarizes various issues for mind uploading. There are many technologies are working on these issues will be summarize. These include questions about whether uploads will have consciousness and whether uploading willpreserve personal identity, as well as what impact on society a working uploading technology is likelyto have and whether these impacts are desirable. The issue of whether we ought to move forwardstowards uploading technology remains as unclear as ever. Keywords: Mind Uploading, Whole brain simulation, I. Introduction Mind uploading is an ongoing area of active research, bringing together ideas from neuroscience, computer science, engineering, and philosophy. Realistically, mind uploading likely lies many decades in the future, but the short-term offers the possibility of advanced neural prostheses that may benefit us. Mind uploading is a popular term for a process by which the mind, a collection of memories, personality, and attributes of a specific individual, is transferred from its original biological brain to an artificial computational substrate. -

Brain Modelling As a Service: the Virtual Brain on EBRAINS
Brain Modelling as a Service: The Virtual Brain on EBRAINS Michael Schirnera,b,c,d,e, Lia Domidef, Dionysios Perdikisa,b, Paul Triebkorna,b,g, Leon Stefanovskia,b, Roopa Paia,b, Paula Prodanf, Bogdan Valeanf, Jessica Palmera,b, Chloê Langforda,b, André Blickensdörfera,b, Michiel van der Vlagh, Sandra Diaz-Pierh, Alexander Peyserh, Wouter Klijnh, Dirk Pleiteri,j, Anne Nahmi, Oliver Schmidk, Marmaduke Woodmang, Lyuba Zehll, Jan Fousekg, Spase Petkoskig, Lionel Kuschg, Meysam Hashemig, Daniele Marinazzom,n, Jean-François Mangino, Agnes Flöelp,q, Simisola Akintoyer, Bernd Carsten Stahls, Michael Cepict, Emily Johnsont, Gustavo Decou,v,w,x, Anthony R. McIntoshy, Claus C. Hilgetagz, a, Marc Morgank, Bernd Schulleri, Alex Uptonb, Colin McMurtrieb, Timo Dickscheidl, Jan G. Bjaaliec, Katrin Amuntsl,d, Jochen Mersmanne, Viktor Jirsag, Petra Rittera,b,c,d,e,* aBerlin Institute of Health at Charité – Universitätsmedizin Berlin, Charitéplatz 1, 10117 Berlin, Germany bCharité – Universitätsmedizin Berlin, corporate memBer of Freie Universität Berlin and HumBoldt Universität zu Berlin, Department of Neurology with Experimental Neurology, Charitéplatz 1, 10117 Berlin, Germany cBernstein Focus State Dependencies of Learning & Bernstein Center for Computational Neuroscience, Berlin, Germany dEinstein Center for Neuroscience Berlin, Charitéplatz 1, 10117 Berlin, Germany eEinstein Center Digital Future, Wilhelmstraße 67, 10117 Berlin, Germany fCodemart, Str. Petofi Sandor, Cluj Napoca, Romania gInstitut de Neurosciences des Systèmes, Aix Marseille Université, -

Computational Tools to Investigate Neurological Disorders
International Journal of Molecular Sciences Review Modeling Neurotransmission: Computational Tools to Investigate Neurological Disorders Daniela Gandolfi 1, Giulia Maria Boiani 1, Albertino Bigiani 1,2 and Jonathan Mapelli 1,2,* 1 Department of Biomedical, Metabolic and Neural Sciences, University of Modena and Reggio Emilia, Via Campi 287, 41125 Modena, Italy; daniela.gandolfi@unimore.it (D.G.); [email protected] (G.M.B.); [email protected] (A.B.) 2 Center for Neuroscience and Neurotechnology, University of Modena and Reggio Emilia, Via Campi 287, 41125 Modena, Italy * Correspondence: [email protected]; Tel.: +39-059-205-5459 Abstract: The investigation of synaptic functions remains one of the most fascinating challenges in the field of neuroscience and a large number of experimental methods have been tuned to dissect the mechanisms taking part in the neurotransmission process. Furthermore, the understanding of the insights of neurological disorders originating from alterations in neurotransmission often requires the development of (i) animal models of pathologies, (ii) invasive tools and (iii) targeted pharmacological approaches. In the last decades, additional tools to explore neurological diseases have been provided to the scientific community. A wide range of computational models in fact have been developed to explore the alterations of the mechanisms involved in neurotransmission following the emergence of neurological pathologies. Here, we review some of the advancements in the development of computational methods employed to investigate neuronal circuits with a particular focus on the application to the most diffuse neurological disorders. Citation: Gandolfi, D.; Boiani, G.M.; Bigiani, A.; Mapelli, J. Modeling Keywords: synaptic transmission; computational modeling; synaptic plasticity; neurological Neurotransmission: Computational disorders Tools to Investigate Neurological Disorders. -

NESTML: a Modeling Language for Spiking Neurons
<Vorname Nachname [et. al.]>(Hrsg.): <Buchtitel>, Lecture Notes in Informatics (LNI), Gesellschaft fur¨ Informatik, Bonn <Jahr> 1 NESTML: a modeling language for spiking neurons Dimitri Plotnikov1, Bernhard Rumpe1, Inga Blundell2, Tammo Ippen2,3, Jochen Martin Eppler4 and Abigail Morrison2,4,5 Abstract: Biological nervous systems exhibit astonishing complexity. Neuroscientists aim to capture this com- plexity by modeling and simulation of biological processes. Often very complex models are nec- essary to depict the processes, which makes it difficult to create these models. Powerful tools are thus necessary, which enable neuroscientists to express models in a comprehensive and concise way and generate efficient code for digital simulations. Several modeling languages for computational neuroscience have been proposed [Gl10, Ra11]. However, as these languages seek simulator inde- pendence they typically only support a subset of the features desired by the modeler. In this article, we present the modular and extensible domain specific language NESTML, which provides neuro- science domain concepts as first-class language constructs and supports domain experts in creating neuron models for the neural simulation tool NEST. NESTML and a set of example models are publically available on GitHub. Keywords: Simulation, modeling, biological neural networks, neuronal modeling, neuroscience, NEST, NESTML, MontiCore, domain specific language, code generation, C++. 1 Introduction Classical neuroscience investigates the biophysical processes behind single neuron behav- ior and higher brain function. The first experimental studies of the nervous system were conducted already hundreds of years ago [Sh91], but observing single-cell activity in a cell culture or slice (in vitro) or in an intact brain (in vivo) is a technically challenging task. -

Pycarl: a Pynn Interface for Hardware-Software Co-Simulation of Spiking Neural Network
PyCARL: A PyNN Interface for Hardware-Software Co-Simulation of Spiking Neural Network Adarsha Balaji1, Prathyusha Adiraju2, Hirak J. Kashyap3, Anup Das1;2, Jeffrey L. Krichmar3, Nikil D. Dutt3, and Francky Catthoor2;4 1Electrical and Computer Engineering, Drexel University, Philadelphia, USA 2Neuromorphic Computing, Stichting Imec Nederlands, Eindhoven, Netherlands 3Cognitive Science and Computer Science, University of California, Irvine, USA 4ESAT Department, KU Leuven and IMEC, Leuven, Belgium Correspondence Email: [email protected], [email protected], [email protected] , Abstract—We present PyCARL, a PyNN-based common To facilitate faster application development and portability Python programming interface for hardware-software co- across research institutes, a common Python programming simulation of spiking neural network (SNN). Through PyCARL, interface called PyNN is proposed [8]. PyNN provides a we make the following two key contributions. First, we pro- vide an interface of PyNN to CARLsim, a computationally- high-level abstraction of SNN models, promotes code sharing efficient, GPU-accelerated and biophysically-detailed SNN simu- and reuse, and provides a foundation for simulator-agnostic lator. PyCARL facilitates joint development of machine learning analysis, visualization and data-management tools. Many SNN models and code sharing between CARLsim and PyNN users, simulators now support interfacing with PyNN — PyNEST [9] promoting an integrated and larger neuromorphic community. for the NEST simulator, PyPCSIM [4] for the PCSIM sim- Second, we integrate cycle-accurate models of state-of-the-art neuromorphic hardware such as TrueNorth, Loihi, and DynapSE ulator, and Brian 2 [10] for the Brian simulator. Through in PyCARL, to accurately model hardware latencies, which delay PyNN, applications developed using one simulator can be an- spikes between communicating neurons, degrading performance alyzed/simulated using another simulator with minimal effort. -

The Grateful Un-Dead? Philosophical and Social Implications of Mind-Uploading
1 July 23, 2020 The grateful Un-dead? Philosophical and Social Implications of Mind-Uploading Ivan William Kelly Professor Emeritus, University of Saskatchewan, Saskatoon, SK, Canada Email address: [email protected] Abstract: The popular belief that our mind either depends on or (in stronger terms) is identical with brain functions and processes, along with the belief that advances in technological in virtual reality and computability will continue, has contributed to the contention that one-day (perhaps this century) it may be possible to transfer one’s mind (or a simulated copy) into another body (physical or virtual). This is called mind-uploading or whole brain emulation. This paper serves as an introduction to the area and some of the major issues that arise from the notion of mind- uploading. The topic of mind/brain uploading is, at present, both a philosophical and scientific minefield. This paper ties together several thoughts on the topic in the popular, philosophical, and scientific literature, and brings up some related issues that might be worth exploring. A challenge is made that uploading a mind from a brain may have intrinsic limitations on what can be uploaded: whatever is uploaded may not be recognizably human, or will be a very limited version. Keywords: mind uploading, brain emulation, mind transfer, virtual reality, robots, android, the mind-body problem, mind transfer, trans-human, 1.Introduction What if (a big if) you actually could upload or transfer your mind into a computer network or a robot? That is, you would now have your mind in a different body or ‘live in’ a virtual reality (or virtual realities). -

Brian 2: an Intuitive and Efficient Neural Simulator
bioRxiv preprint doi: https://doi.org/10.1101/595710; this version posted April 1, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY 4.0 International license. Brian 2: an intuitive and efficient neural simulator Marcel Stimberg1*, Romain Brette1, and Dan F. M. Goodman2 1Sorbonne Université, INSERM, CNRS, Institut de la Vision, Paris, France 2Department of Electrical and Electronic Engineering, Imperial College London, UK Abstract To be maximally useful for neuroscience research, neural simulators must make it possible to define original models. This is especially important because a computational experiment might not only need descriptions of neurons and synapses, but also models of interactions with the environment (e.g. muscles), or the environment itself. To preserve high performance when defining new models, current simulators offer two options: low-level programming, or mark-up languages (and other domain specific languages). The first option requires time and expertise, is prone to errors, and contributes to problems with reproducibility and replicability. The second option has limited scope, since it can only describe the range of neural models covered by the ontology. Other aspects of a computational experiment, such as the stimulation protocol, cannot be expressed within this framework. “Brian” 2 is a complete rewrite of Brian that addresses this issue by using runtime code generation with a procedural equation-oriented approach. Brian 2 enables scientists to write code that is particularly simple and concise, closely matching the way they conceptualise their models, while the technique of runtime code generation automatically transforms high level descriptions of models into efficient low level code tailored to different hardware (e.g.