The LIM Domain of Zyxin Is Sufficient for Force-Induced Accumulation Of
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Bioinformatic Analysis of Structure and Function of LIM Domains of Human Zyxin Family Proteins
International Journal of Molecular Sciences Article Bioinformatic Analysis of Structure and Function of LIM Domains of Human Zyxin Family Proteins M. Quadir Siddiqui 1,† , Maulik D. Badmalia 1,† and Trushar R. Patel 1,2,3,* 1 Alberta RNA Research and Training Institute, Department of Chemistry and Biochemistry, University of Lethbridge, 4401 University Drive, Lethbridge, AB T1K 3M4, Canada; [email protected] (M.Q.S.); [email protected] (M.D.B.) 2 Department of Microbiology, Immunology and Infectious Disease, Cumming School of Medicine, University of Calgary, 3330 Hospital Drive, Calgary, AB T2N 4N1, Canada 3 Li Ka Shing Institute of Virology, University of Alberta, Edmonton, AB T6G 2E1, Canada * Correspondence: [email protected] † These authors contributed equally to the work. Abstract: Members of the human Zyxin family are LIM domain-containing proteins that perform critical cellular functions and are indispensable for cellular integrity. Despite their importance, not much is known about their structure, functions, interactions and dynamics. To provide insights into these, we used a set of in-silico tools and databases and analyzed their amino acid sequence, phylogeny, post-translational modifications, structure-dynamics, molecular interactions, and func- tions. Our analysis revealed that zyxin members are ohnologs. Presence of a conserved nuclear export signal composed of LxxLxL/LxxxLxL consensus sequence, as well as a possible nuclear localization signal, suggesting that Zyxin family members may have nuclear and cytoplasmic roles. The molecular modeling and structural analysis indicated that Zyxin family LIM domains share Citation: Siddiqui, M.Q.; Badmalia, similarities with transcriptional regulators and have positively charged electrostatic patches, which M.D.; Patel, T.R. -

Alpha;-Actinin-4 Promotes Metastasis in Gastric Cancer
Laboratory Investigation (2017) 97, 1084–1094 © 2017 USCAP, Inc All rights reserved 0023-6837/17 α-Actinin-4 promotes metastasis in gastric cancer Xin Liu and Kent-Man Chu Metastasis increases the mortality rate of gastric cancer, which is the third leading cause of cancer-associated deaths worldwide. This study aims to identify the genes promoting metastasis of gastric cancer (GC). A human cell motility PCR array was used to analyze a pair of tumor and non-tumor tissue samples from a patient with stage IV GC (T3N3M1). Expression of the dysregulated genes was then evaluated in GC tissue samples (n = 10) and cell lines (n =6) via qPCR. Expression of α-actinin-4 (ACTN4) was validated in a larger sample size (n = 47) by qPCR, western blot and immunohistochemistry. Knockdown of ACTN4 with specific siRNAs was performed in GC cells, and adhesion assays, transwell invasion assays and migration assays were used to evaluate the function of these cells. Expression of potential targets of ACTN4 were then evaluated by qPCR. Thirty upregulated genes (greater than twofold) were revealed by the PCR array. We focused on ACTN4 because it was upregulated in 6 out of 10 pairs of tissue samples and 5 out of 6 GC cell lines. Further study indicated that ACTN4 was upregulated in 22/32 pairs of tissue samples at stage III & IV (P = 0.0069). Knockdown of ACTN4 in GC cells showed no significant effect on cell proliferation, but significantly increased cell-matrix adhesion, as well as reduced migration and invasion of AGS, MKN7 and NCI-N87 cells. -
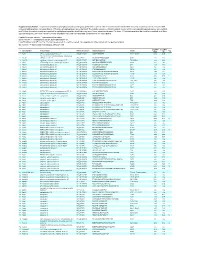
Supplementary Table 1
Supplementary Table 1. Large-scale quantitative phosphoproteomic profiling was performed on paired vehicle- and hormone-treated mTAL-enriched suspensions (n=3). A total of 654 unique phosphopeptides corresponding to 374 unique phosphoproteins were identified. The peptide sequence, phosphorylation site(s), and the corresponding protein name, gene symbol, and RefSeq Accession number are reported for each phosphopeptide identified in any one of three experimental pairs. For those 414 phosphopeptides that could be quantified in all three experimental pairs, the mean Hormone:Vehicle abundance ratio and corresponding standard error are also reported. Peptide Sequence column: * = phosphorylated residue Site(s) column: ^ = ambiguously assigned phosphorylation site Log2(H/V) Mean and SE columns: H = hormone-treated, V = vehicle-treated, n/a = peptide not observable in all 3 experimental pairs Sig. column: * = significantly changed Log 2(H/V), p<0.05 Log (H/V) Log (H/V) # Gene Symbol Protein Name Refseq Accession Peptide Sequence Site(s) 2 2 Sig. Mean SE 1 Aak1 AP2-associated protein kinase 1 NP_001166921 VGSLT*PPSS*PK T622^, S626^ 0.24 0.95 PREDICTED: ATP-binding cassette, sub-family A 2 Abca12 (ABC1), member 12 XP_237242 GLVQVLS*FFSQVQQQR S251^ 1.24 2.13 3 Abcc10 multidrug resistance-associated protein 7 NP_001101671 LMT*ELLS*GIRVLK T464, S468 -2.68 2.48 4 Abcf1 ATP-binding cassette sub-family F member 1 NP_001103353 QLSVPAS*DEEDEVPVPVPR S109 n/a n/a 5 Ablim1 actin-binding LIM protein 1 NP_001037859 PGSSIPGS*PGHTIYAK S51 -3.55 1.81 6 Ablim1 actin-binding -

Role and Regulation of the P53-Homolog P73 in the Transformation of Normal Human Fibroblasts
Role and regulation of the p53-homolog p73 in the transformation of normal human fibroblasts Dissertation zur Erlangung des naturwissenschaftlichen Doktorgrades der Bayerischen Julius-Maximilians-Universität Würzburg vorgelegt von Lars Hofmann aus Aschaffenburg Würzburg 2007 Eingereicht am Mitglieder der Promotionskommission: Vorsitzender: Prof. Dr. Dr. Martin J. Müller Gutachter: Prof. Dr. Michael P. Schön Gutachter : Prof. Dr. Georg Krohne Tag des Promotionskolloquiums: Doktorurkunde ausgehändigt am Erklärung Hiermit erkläre ich, dass ich die vorliegende Arbeit selbständig angefertigt und keine anderen als die angegebenen Hilfsmittel und Quellen verwendet habe. Diese Arbeit wurde weder in gleicher noch in ähnlicher Form in einem anderen Prüfungsverfahren vorgelegt. Ich habe früher, außer den mit dem Zulassungsgesuch urkundlichen Graden, keine weiteren akademischen Grade erworben und zu erwerben gesucht. Würzburg, Lars Hofmann Content SUMMARY ................................................................................................................ IV ZUSAMMENFASSUNG ............................................................................................. V 1. INTRODUCTION ................................................................................................. 1 1.1. Molecular basics of cancer .......................................................................................... 1 1.2. Early research on tumorigenesis ................................................................................. 3 1.3. Developing -

Human Induced Pluripotent Stem Cell–Derived Podocytes Mature Into Vascularized Glomeruli Upon Experimental Transplantation
BASIC RESEARCH www.jasn.org Human Induced Pluripotent Stem Cell–Derived Podocytes Mature into Vascularized Glomeruli upon Experimental Transplantation † Sazia Sharmin,* Atsuhiro Taguchi,* Yusuke Kaku,* Yasuhiro Yoshimura,* Tomoko Ohmori,* ‡ † ‡ Tetsushi Sakuma, Masashi Mukoyama, Takashi Yamamoto, Hidetake Kurihara,§ and | Ryuichi Nishinakamura* *Department of Kidney Development, Institute of Molecular Embryology and Genetics, and †Department of Nephrology, Faculty of Life Sciences, Kumamoto University, Kumamoto, Japan; ‡Department of Mathematical and Life Sciences, Graduate School of Science, Hiroshima University, Hiroshima, Japan; §Division of Anatomy, Juntendo University School of Medicine, Tokyo, Japan; and |Japan Science and Technology Agency, CREST, Kumamoto, Japan ABSTRACT Glomerular podocytes express proteins, such as nephrin, that constitute the slit diaphragm, thereby contributing to the filtration process in the kidney. Glomerular development has been analyzed mainly in mice, whereas analysis of human kidney development has been minimal because of limited access to embryonic kidneys. We previously reported the induction of three-dimensional primordial glomeruli from human induced pluripotent stem (iPS) cells. Here, using transcription activator–like effector nuclease-mediated homologous recombination, we generated human iPS cell lines that express green fluorescent protein (GFP) in the NPHS1 locus, which encodes nephrin, and we show that GFP expression facilitated accurate visualization of nephrin-positive podocyte formation in -

Actin Cytoskeleton and Regulation of Tgfβ Signaling: Exploring Their Links
biomolecules Review Actin Cytoskeleton and Regulation of TGFβ Signaling: Exploring Their Links Roberta Melchionna 1,†, Paola Trono 1,2,†, Annalisa Tocci 1 and Paola Nisticò 1,* 1 Tumor Immunology and Immunotherapy Unit, IRCCS Regina Elena National Cancer Institute, via Chianesi 53, 00144 Rome, Italy; [email protected] (R.M.); [email protected] (P.T.); [email protected] (A.T.) 2 Institute of Biochemistry and Cell Biology, National Research Council, via Ramarini 32, 00015 Monterotondo Scalo, Rome, Italy * Correspondence: [email protected]; Tel.: +39-0652662539 † These authors contributed equally to this paper. Abstract: Human tissues, to maintain their architecture and function, respond to injuries by activating intricate biochemical and physical mechanisms that regulates intercellular communication crucial in maintaining tissue homeostasis. Coordination of the communication occurs through the activity of different actin cytoskeletal regulators, physically connected to extracellular matrix through integrins, generating a platform of biochemical and biomechanical signaling that is deregulated in cancer. Among the major pathways, a controller of cellular functions is the cytokine transforming growth factor β (TGFβ), which remains a complex and central signaling network still to be interpreted and explained in cancer progression. Here, we discuss the link between actin dynamics and TGFβ signaling with the aim of exploring their aberrant interaction in cancer. Keywords: actin cytoskeleton; actin-binding proteins; TGFβ; extracellular matrix; tumor microenvi- Citation: Melchionna, R.; Trono, P.; ronment Tocci, A.; Nisticò, P. Actin Cytoskeleton and Regulation of TGFβ Signaling: Exploring Their Links. Biomolecules 2021, 11, 336. 1. Introduction https://doi.org/10.3390/biom11020336 Actin dynamics critically affect different aspects of human health and disease, ranging Academic Editor: Vladimir from embryonic development to wound repair, inflammation, and cancer [1]. -

Spatial and Temporal Aspects of Signalling 6 1
r r r Cell Signalling Biology Michael J. Berridge Module 6 Spatial and Temporal Aspects of Signalling 6 1 Module 6 Spatial and Temporal Aspects of Signalling Synopsis The function and efficiency of cell signalling pathways are very dependent on their organization both in space and time. With regard to spatial organization, signalling components are highly organized with respect to their cellular location and how they transmit information from one region of the cell to another. This spatial organization of signalling pathways depends on the molecular interactions that occur between signalling components that use signal transduction domains to construct signalling pathways. Very often, the components responsible for information transfer mechanisms are held in place by being attached to scaffolding proteins to form macromolecular signalling complexes. Sometimes these macromolecular complexes can be organized further by being localized to specific regions of the cell, as found in lipid rafts and caveolae or in the T-tubule regions of skeletal and cardiac cells. Another feature of the spatial aspects concerns the local Another important temporal aspect is timing and signal and global aspects of signalling. The spatial organization of integration, which relates to the way in which functional signalling molecules mentioned above can lead to highly interactions between signalling pathways are determined localized signalling events, but when the signalling mo- by both the order and the timing of their presentations. lecules are more evenly distributed, signals can spread The organization of signalling systems in both time and more globally throughout the cell. In addition, signals space greatly enhances both their efficiency and versatility. -

Single-Cell Transcriptomes Reveal a Complex Cellular Landscape in the Middle Ear and Differential Capacities for Acute Response to Infection
fgene-11-00358 April 9, 2020 Time: 15:55 # 1 ORIGINAL RESEARCH published: 15 April 2020 doi: 10.3389/fgene.2020.00358 Single-Cell Transcriptomes Reveal a Complex Cellular Landscape in the Middle Ear and Differential Capacities for Acute Response to Infection Allen F. Ryan1*, Chanond A. Nasamran2, Kwang Pak1, Clara Draf1, Kathleen M. Fisch2, Nicholas Webster3 and Arwa Kurabi1 1 Departments of Surgery/Otolaryngology, UC San Diego School of Medicine, VA Medical Center, La Jolla, CA, United States, 2 Medicine/Center for Computational Biology & Bioinformatics, UC San Diego School of Medicine, VA Medical Center, La Jolla, CA, United States, 3 Medicine/Endocrinology, UC San Diego School of Medicine, VA Medical Center, La Jolla, CA, United States Single-cell transcriptomics was used to profile cells of the normal murine middle ear. Clustering analysis of 6770 transcriptomes identified 17 cell clusters corresponding to distinct cell types: five epithelial, three stromal, three lymphocyte, two monocyte, Edited by: two endothelial, one pericyte and one melanocyte cluster. Within some clusters, Amélie Bonnefond, Institut National de la Santé et de la cell subtypes were identified. While many corresponded to those cell types known Recherche Médicale (INSERM), from prior studies, several novel types or subtypes were noted. The results indicate France unexpected cellular diversity within the resting middle ear mucosa. The resolution of Reviewed by: Fabien Delahaye, uncomplicated, acute, otitis media is too rapid for cognate immunity to play a major Institut Pasteur de Lille, France role. Thus innate immunity is likely responsible for normal recovery from middle ear Nelson L. S. Tang, infection. The need for rapid response to pathogens suggests that innate immune The Chinese University of Hong Kong, China genes may be constitutively expressed by middle ear cells. -

Actin Bundling Via LIM Domains
[Plant Signaling & Behavior 3:5, 320-321; May 2008]; ©2008 Landes Bioscience Article Addendum Actin bundling via LIM domains Clément Thomas,* Monika Dieterle, Sabrina Gatti, Céline Hoffmann, Flora Moreau, Jessica Papuga and André Steinmetz Centre de Recherche Public-Santé; Val Fleuri 84; L-1526; Luxembourg Key words: Actin-binding proteins, actin-bundling, cysteine-rich proteins, cytoskeleton, LIM domain. The LIM domain is defined as a protein-protein interaction module involved in the regulation of diverse cellular processes including gene expression and cytoskeleton organization. We distribute. have recently shown that the tobacco WLIM1, a two LIM domain-containing protein, is able to bind to, stabilize and bundle actin filaments, suggesting that it participates to the regulation of actin cytoskeleton structure and dynamics. In the December issue of the Journal of Biological Chemistry we report a domain analysis not that specifically ascribes the actin-related activities of WLIM1 to its two LIM domains. Results suggest that LIM domains func- Figure 1. Domain maps for wild-type WLIM1 (A) and GFP-fused chimeric tion synergistically in the full-length protein to achieve optimal 3xWLIM1 (B). A. WLIM1 basically comprises a short N-terminal domain (Nt), two LIM domains (LIM1Do and LIM2), an interLIM spacer (IL) and a C-terminal activities. Here we briefly summarize relevant data regarding the domain (Ct). B. 3xWLIM1 consists of three tandem WLIM1 copies. This chi- actin-related properties/functions of two LIM domain-containing meric protein has been fused in C-terminus to GFP and transiently expressed proteins in plants and animals. In addition, we provide further in tobacco BY2 cells. -

Early Evolution of the LIM Homeobox Gene Family
Srivastava et al. BMC Biology 2010, 8:4 http://www.biomedcentral.com/1741-7007/8/4 RESEARCH ARTICLE Open Access Early evolution of the LIM homeobox gene family Mansi Srivastava1*, Claire Larroux2, Daniel R Lu1, Kareshma Mohanty1, Jarrod Chapman3, Bernard M Degnan2, Daniel S Rokhsar1,3 Abstract Background: LIM homeobox (Lhx) transcription factors are unique to the animal lineage and have patterning roles during embryonic development in flies, nematodes and vertebrates, with a conserved role in specifying neuronal identity. Though genes of this family have been reported in a sponge and a cnidarian, the expression patterns and functions of the Lhx family during development in non-bilaterian phyla are not known. Results: We identified Lhx genes in two cnidarians and a placozoan and report the expression of Lhx genes during embryonic development in Nematostella and the demosponge Amphimedon. Members of the six major LIM homeobox subfamilies are represented in the genomes of the starlet sea anemone, Nematostella vectensis, and the placozoan Trichoplax adhaerens. The hydrozoan cnidarian, Hydra magnipapillata, has retained four of the six Lhx subfamilies, but apparently lost two others. Only three subfamilies are represented in the haplosclerid demosponge Amphimedon queenslandica. A tandem cluster of three Lhx genes of different subfamilies and a gene containing two LIM domains in the genome of T. adhaerens (an animal without any neurons) indicates that Lhx subfamilies were generated by tandem duplication. This tandem cluster in Trichoplax is likely a remnant of the original chromosomal context in which Lhx subfamilies first appeared. Three of the six Trichoplax Lhx genes are expressed in animals in laboratory culture, as are all Lhx genes in Hydra. -

Manipulation of Focal Adhesion Signaling by Pathogenic Microbes
International Journal of Molecular Sciences Review Manipulation of Focal Adhesion Signaling by Pathogenic Microbes Korinn N. Murphy 1,2 and Amanda J. Brinkworth 2,* 1 School of Molecular Biosciences, Washington State University, Pullman, WA 99164, USA; [email protected] 2 Department of Pathology and Microbiology, University of Nebraska Medical Center, Omaha, NE 68198, USA * Correspondence: [email protected]; Tel.: +1-402-836-9777 Abstract: Focal adhesions (FAs) serve as dynamic signaling hubs within the cell. They connect intracellular actin to the extracellular matrix (ECM) and respond to environmental cues. In doing so, these structures facilitate important processes such as cell–ECM adhesion and migration. Pathogenic microbes often modify the host cell actin cytoskeleton in their pursuit of an ideal replicative niche or during invasion to facilitate uptake. As actin-interfacing structures, FA dynamics are also intimately tied to actin cytoskeletal organization. Indeed, exploitation of FAs is another avenue by which pathogenic microbes ensure their uptake, survival and dissemination. This is often achieved through the secretion of effector proteins which target specific protein components within the FA. Molecular mimicry of the leucine–aspartic acid (LD) motif or vinculin-binding domains (VBDs) commonly found within FA proteins is a common microbial strategy. Other effectors may induce post-translational modifications to FA proteins through the regulation of phosphorylation sites or proteolytic cleavage. In this review, we present an overview of the regulatory mechanisms governing host cell FAs, and provide examples of how pathogenic microbes have evolved to co-opt them to their own advantage. Recent technological advances pose exciting opportunities for delving deeper into the mechanistic details by which pathogenic microbes modify FAs. -

A Chromosome-Centric Human Proteome Project (C-HPP) To
computational proteomics Laboratory for Computational Proteomics www.FenyoLab.org E-mail: [email protected] Facebook: NYUMC Computational Proteomics Laboratory Twitter: @CompProteomics Perspective pubs.acs.org/jpr A Chromosome-centric Human Proteome Project (C-HPP) to Characterize the Sets of Proteins Encoded in Chromosome 17 † ‡ § ∥ ‡ ⊥ Suli Liu, Hogune Im, Amos Bairoch, Massimo Cristofanilli, Rui Chen, Eric W. Deutsch, # ¶ △ ● § † Stephen Dalton, David Fenyo, Susan Fanayan,$ Chris Gates, , Pascale Gaudet, Marina Hincapie, ○ ■ △ ⬡ ‡ ⊥ ⬢ Samir Hanash, Hoguen Kim, Seul-Ki Jeong, Emma Lundberg, George Mias, Rajasree Menon, , ∥ □ △ # ⬡ ▲ † Zhaomei Mu, Edouard Nice, Young-Ki Paik, , Mathias Uhlen, Lance Wells, Shiaw-Lin Wu, † † † ‡ ⊥ ⬢ ⬡ Fangfei Yan, Fan Zhang, Yue Zhang, Michael Snyder, Gilbert S. Omenn, , Ronald C. Beavis, † # and William S. Hancock*, ,$, † Barnett Institute and Department of Chemistry and Chemical Biology, Northeastern University, Boston, Massachusetts 02115, United States ‡ Stanford University, Palo Alto, California, United States § Swiss Institute of Bioinformatics (SIB) and University of Geneva, Geneva, Switzerland ∥ Fox Chase Cancer Center, Philadelphia, Pennsylvania, United States ⊥ Institute for System Biology, Seattle, Washington, United States ¶ School of Medicine, New York University, New York, United States $Department of Chemistry and Biomolecular Sciences, Macquarie University, Sydney, NSW, Australia ○ MD Anderson Cancer Center, Houston, Texas, United States ■ Yonsei University College of Medicine, Yonsei University,