Characterization and Sequence Analysis of Cysteine and Glycine-Rich Protein 3 in Egyptian Native Cattle and River Native Buffalo Cdna Sequences
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Bioinformatic Analysis of Structure and Function of LIM Domains of Human Zyxin Family Proteins
International Journal of Molecular Sciences Article Bioinformatic Analysis of Structure and Function of LIM Domains of Human Zyxin Family Proteins M. Quadir Siddiqui 1,† , Maulik D. Badmalia 1,† and Trushar R. Patel 1,2,3,* 1 Alberta RNA Research and Training Institute, Department of Chemistry and Biochemistry, University of Lethbridge, 4401 University Drive, Lethbridge, AB T1K 3M4, Canada; [email protected] (M.Q.S.); [email protected] (M.D.B.) 2 Department of Microbiology, Immunology and Infectious Disease, Cumming School of Medicine, University of Calgary, 3330 Hospital Drive, Calgary, AB T2N 4N1, Canada 3 Li Ka Shing Institute of Virology, University of Alberta, Edmonton, AB T6G 2E1, Canada * Correspondence: [email protected] † These authors contributed equally to the work. Abstract: Members of the human Zyxin family are LIM domain-containing proteins that perform critical cellular functions and are indispensable for cellular integrity. Despite their importance, not much is known about their structure, functions, interactions and dynamics. To provide insights into these, we used a set of in-silico tools and databases and analyzed their amino acid sequence, phylogeny, post-translational modifications, structure-dynamics, molecular interactions, and func- tions. Our analysis revealed that zyxin members are ohnologs. Presence of a conserved nuclear export signal composed of LxxLxL/LxxxLxL consensus sequence, as well as a possible nuclear localization signal, suggesting that Zyxin family members may have nuclear and cytoplasmic roles. The molecular modeling and structural analysis indicated that Zyxin family LIM domains share Citation: Siddiqui, M.Q.; Badmalia, similarities with transcriptional regulators and have positively charged electrostatic patches, which M.D.; Patel, T.R. -

DF6216-CSRP1 Antibody
Affinity Biosciences website:www.affbiotech.com order:[email protected] CSRP1 Antibody Cat.#: DF6216 Concn.: 1mg/ml Mol.Wt.: 21kDa Size: 50ul,100ul,200ul Source: Rabbit Clonality: Polyclonal Application: WB 1:500-1:2000, IHC 1:50-1:200, ELISA(peptide) 1:20000-1:40000 *The optimal dilutions should be determined by the end user. Reactivity: Human,Mouse,Rat Purification: The antiserum was purified by peptide affinity chromatography using SulfoLink™ Coupling Resin (Thermo Fisher Scientific). Specificity: CSRP1 Antibody detects endogenous levels of total CSRP1. Immunogen: A synthesized peptide derived from human CSRP1, corresponding to a region within the internal amino acids. Uniprot: P21291 Description: This gene encodes a member of the cysteine-rich protein (CSRP) family. This gene family includes a group of LIM domain proteins, which may be involved in regulatory processes important for development and cellular differentiation. The LIM/double zinc-finger motif found in this gene product occurs in proteins with critical functions in gene regulation, cell growth, and somatic differentiation. Alternatively spliced transcript variants have been described. [provided by RefSeq, Aug 2010] Storage Condition and Rabbit IgG in phosphate buffered saline , pH 7.4, 150mM Buffer: NaCl, 0.02% sodium azide and 50% glycerol.Store at -20 °C.Stable for 12 months from date of receipt. Western blot analysis of CSRP1 expression in Mouse lung lysate 1 / 2 Affinity Biosciences website:www.affbiotech.com order:[email protected] DF6216 at 1/100 staining Mouse brain tissue by IHC-P. The sample was formaldehyde fixed and a heat mediated antigen retrieval step in citrate buffer was performed. -

Screening and Identification of Hub Genes in Bladder Cancer by Bioinformatics Analysis and KIF11 Is a Potential Prognostic Biomarker
ONCOLOGY LETTERS 21: 205, 2021 Screening and identification of hub genes in bladder cancer by bioinformatics analysis and KIF11 is a potential prognostic biomarker XIAO‑CONG MO1,2*, ZI‑TONG ZHANG1,3*, MENG‑JIA SONG1,2, ZI‑QI ZHOU1,2, JIAN‑XIONG ZENG1,2, YU‑FEI DU1,2, FENG‑ZE SUN1,2, JIE‑YING YANG1,2, JUN‑YI HE1,2, YUE HUANG1,2, JIAN‑CHUAN XIA1,2 and DE‑SHENG WENG1,2 1State Key Laboratory of Oncology in South China, Collaborative Innovation Centre for Cancer Medicine; 2Department of Biotherapy, Sun Yat‑Sen University Cancer Center; 3Department of Radiation Oncology, Sun Yat‑Sen University Cancer Center, Guangzhou, Guangdong 510060, P.R. China Received July 31, 2020; Accepted December 18, 2020 DOI: 10.3892/ol.2021.12466 Abstract. Bladder cancer (BC) is the ninth most common immunohistochemistry and western blotting. In summary, lethal malignancy worldwide. Great efforts have been devoted KIF11 was significantly upregulated in BC and might act as to clarify the pathogenesis of BC, but the underlying molecular a potential prognostic biomarker. The present identification mechanisms remain unclear. To screen for the genes associated of DEGs and hub genes in BC may provide novel insight for with the progression and carcinogenesis of BC, three datasets investigating the molecular mechanisms of BC. were obtained from the Gene Expression Omnibus. A total of 37 tumor and 16 non‑cancerous samples were analyzed to Introduction identify differentially expressed genes (DEGs). Subsequently, 141 genes were identified, including 55 upregulated and Bladder cancer (BC) is the ninth most common malignancy 86 downregulated genes. The protein‑protein interaction worldwide with substantial morbidity and mortality. -

Alpha;-Actinin-4 Promotes Metastasis in Gastric Cancer
Laboratory Investigation (2017) 97, 1084–1094 © 2017 USCAP, Inc All rights reserved 0023-6837/17 α-Actinin-4 promotes metastasis in gastric cancer Xin Liu and Kent-Man Chu Metastasis increases the mortality rate of gastric cancer, which is the third leading cause of cancer-associated deaths worldwide. This study aims to identify the genes promoting metastasis of gastric cancer (GC). A human cell motility PCR array was used to analyze a pair of tumor and non-tumor tissue samples from a patient with stage IV GC (T3N3M1). Expression of the dysregulated genes was then evaluated in GC tissue samples (n = 10) and cell lines (n =6) via qPCR. Expression of α-actinin-4 (ACTN4) was validated in a larger sample size (n = 47) by qPCR, western blot and immunohistochemistry. Knockdown of ACTN4 with specific siRNAs was performed in GC cells, and adhesion assays, transwell invasion assays and migration assays were used to evaluate the function of these cells. Expression of potential targets of ACTN4 were then evaluated by qPCR. Thirty upregulated genes (greater than twofold) were revealed by the PCR array. We focused on ACTN4 because it was upregulated in 6 out of 10 pairs of tissue samples and 5 out of 6 GC cell lines. Further study indicated that ACTN4 was upregulated in 22/32 pairs of tissue samples at stage III & IV (P = 0.0069). Knockdown of ACTN4 in GC cells showed no significant effect on cell proliferation, but significantly increased cell-matrix adhesion, as well as reduced migration and invasion of AGS, MKN7 and NCI-N87 cells. -

Key Genes Regulating Skeletal Muscle Development and Growth in Farm Animals
animals Review Key Genes Regulating Skeletal Muscle Development and Growth in Farm Animals Mohammadreza Mohammadabadi 1 , Farhad Bordbar 1,* , Just Jensen 2 , Min Du 3 and Wei Guo 4 1 Department of Animal Science, Faculty of Agriculture, Shahid Bahonar University of Kerman, Kerman 77951, Iran; [email protected] 2 Center for Quantitative Genetics and Genomics, Aarhus University, 8210 Aarhus, Denmark; [email protected] 3 Washington Center for Muscle Biology, Department of Animal Sciences, Washington State University, Pullman, WA 99163, USA; [email protected] 4 Muscle Biology and Animal Biologics, Animal and Dairy Science, University of Wisconsin-Madison, Madison, WI 53558, USA; [email protected] * Correspondence: [email protected] Simple Summary: Skeletal muscle mass is an important economic trait, and muscle development and growth is a crucial factor to supply enough meat for human consumption. Thus, understanding (candidate) genes regulating skeletal muscle development is crucial for understanding molecular genetic regulation of muscle growth and can be benefit the meat industry toward the goal of in- creasing meat yields. During the past years, significant progress has been made for understanding these mechanisms, and thus, we decided to write a comprehensive review covering regulators and (candidate) genes crucial for muscle development and growth in farm animals. Detection of these genes and factors increases our understanding of muscle growth and development and is a great help for breeders to satisfy demands for meat production on a global scale. Citation: Mohammadabadi, M.; Abstract: Farm-animal species play crucial roles in satisfying demands for meat on a global scale, Bordbar, F.; Jensen, J.; Du, M.; Guo, W. -

CRIP1 Antibody (C-Term) Affinity Purified Rabbit Polyclonal Antibody (Pab) Catalog # Ap4707b
10320 Camino Santa Fe, Suite G San Diego, CA 92121 Tel: 858.875.1900 Fax: 858.622.0609 CRIP1 Antibody (C-term) Affinity Purified Rabbit Polyclonal Antibody (Pab) Catalog # AP4707b Specification CRIP1 Antibody (C-term) - Product Information Application WB, IHC-P, FC,E Primary Accession P50238 Other Accession P63255, P63254, Q56K04 Reactivity Human Predicted Bovine, Mouse, Rat Host Rabbit Clonality Polyclonal Isotype Rabbit Ig Calculated MW 8533 Antigen Region 48-77 CRIP1 Antibody (C-term) - Additional Information Gene ID 1396 All lanes : Anti-CRIP1 Antibody (C-term) at 1:8000 dilution Lane 1: T47D whole cell Other Names lysate Lane 2: THP-1 whole cell lysate Cysteine-rich protein 1, CRP-1, Cysteine-rich Lysates/proteins at 20 µg per lane. heart protein, CRHP, hCRHP, Cysteine-rich Secondary Goat Anti-Rabbit IgG, (H+L), intestinal protein, CRIP, CRIP1, CRIP, CRP1 Peroxidase conjugated at 1/10000 dilution. Predicted band size : 9 kDa Blocking/Dilution Target/Specificity buffer: 5% NFDM/TBST. This CRIP1 antibody is generated from rabbits immunized with a KLH conjugated synthetic peptide between 48-77 amino acids from the C-terminal region of human CRIP1. Dilution WB~~1:8000 IHC-P~~1:50~100 FC~~1:10~50 Format Purified polyclonal antibody supplied in PBS with 0.09% (W/V) sodium azide. This antibody is purified through a protein A column, followed by peptide affinity CRIP1 Antibody (C-term) (Cat. #AP4707b) purification. IHC analysis in formalin fixed and paraffin embedded bladder carcinoma followed by Storage peroxidase conjugation of the secondary Maintain refrigerated at 2-8°C for up to 2 antibody and DAB staining. -
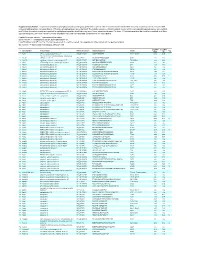
Supplementary Table 1
Supplementary Table 1. Large-scale quantitative phosphoproteomic profiling was performed on paired vehicle- and hormone-treated mTAL-enriched suspensions (n=3). A total of 654 unique phosphopeptides corresponding to 374 unique phosphoproteins were identified. The peptide sequence, phosphorylation site(s), and the corresponding protein name, gene symbol, and RefSeq Accession number are reported for each phosphopeptide identified in any one of three experimental pairs. For those 414 phosphopeptides that could be quantified in all three experimental pairs, the mean Hormone:Vehicle abundance ratio and corresponding standard error are also reported. Peptide Sequence column: * = phosphorylated residue Site(s) column: ^ = ambiguously assigned phosphorylation site Log2(H/V) Mean and SE columns: H = hormone-treated, V = vehicle-treated, n/a = peptide not observable in all 3 experimental pairs Sig. column: * = significantly changed Log 2(H/V), p<0.05 Log (H/V) Log (H/V) # Gene Symbol Protein Name Refseq Accession Peptide Sequence Site(s) 2 2 Sig. Mean SE 1 Aak1 AP2-associated protein kinase 1 NP_001166921 VGSLT*PPSS*PK T622^, S626^ 0.24 0.95 PREDICTED: ATP-binding cassette, sub-family A 2 Abca12 (ABC1), member 12 XP_237242 GLVQVLS*FFSQVQQQR S251^ 1.24 2.13 3 Abcc10 multidrug resistance-associated protein 7 NP_001101671 LMT*ELLS*GIRVLK T464, S468 -2.68 2.48 4 Abcf1 ATP-binding cassette sub-family F member 1 NP_001103353 QLSVPAS*DEEDEVPVPVPR S109 n/a n/a 5 Ablim1 actin-binding LIM protein 1 NP_001037859 PGSSIPGS*PGHTIYAK S51 -3.55 1.81 6 Ablim1 actin-binding -

Identification of Differentially Expressed Genes in Human Bladder Cancer Through Genome-Wide Gene Expression Profiling
521-531 24/7/06 18:28 Page 521 ONCOLOGY REPORTS 16: 521-531, 2006 521 Identification of differentially expressed genes in human bladder cancer through genome-wide gene expression profiling KAZUMORI KAWAKAMI1,3, HIDEKI ENOKIDA1, TOKUSHI TACHIWADA1, TAKENARI GOTANDA1, KENGO TSUNEYOSHI1, HIROYUKI KUBO1, KENRYU NISHIYAMA1, MASAKI TAKIGUCHI2, MASAYUKI NAKAGAWA1 and NAOHIKO SEKI3 1Department of Urology, Graduate School of Medical and Dental Sciences, Kagoshima University, 8-35-1 Sakuragaoka, Kagoshima 890-8520; Departments of 2Biochemistry and Genetics, and 3Functional Genomics, Graduate School of Medicine, Chiba University, 1-8-1 Inohana, Chuo-ku, Chiba 260-8670, Japan Received February 15, 2006; Accepted April 27, 2006 Abstract. Large-scale gene expression profiling is an effective CKS2 gene not only as a potential biomarker for diagnosing, strategy for understanding the progression of bladder cancer but also for staging human BC. This is the first report (BC). The aim of this study was to identify genes that are demonstrating that CKS2 expression is strongly correlated expressed differently in the course of BC progression and to with the progression of human BC. establish new biomarkers for BC. Specimens from 21 patients with pathologically confirmed superficial (n=10) or Introduction invasive (n=11) BC and 4 normal bladder samples were studied; samples from 14 of the 21 BC samples were subjected Bladder cancer (BC) is among the 5 most common to microarray analysis. The validity of the microarray results malignancies worldwide, and the 2nd most common tumor of was verified by real-time RT-PCR. Of the 136 up-regulated the genitourinary tract and the 2nd most common cause of genes we detected, 21 were present in all 14 BCs examined death in patients with cancer of the urinary tract (1-7). -

Role and Regulation of the P53-Homolog P73 in the Transformation of Normal Human Fibroblasts
Role and regulation of the p53-homolog p73 in the transformation of normal human fibroblasts Dissertation zur Erlangung des naturwissenschaftlichen Doktorgrades der Bayerischen Julius-Maximilians-Universität Würzburg vorgelegt von Lars Hofmann aus Aschaffenburg Würzburg 2007 Eingereicht am Mitglieder der Promotionskommission: Vorsitzender: Prof. Dr. Dr. Martin J. Müller Gutachter: Prof. Dr. Michael P. Schön Gutachter : Prof. Dr. Georg Krohne Tag des Promotionskolloquiums: Doktorurkunde ausgehändigt am Erklärung Hiermit erkläre ich, dass ich die vorliegende Arbeit selbständig angefertigt und keine anderen als die angegebenen Hilfsmittel und Quellen verwendet habe. Diese Arbeit wurde weder in gleicher noch in ähnlicher Form in einem anderen Prüfungsverfahren vorgelegt. Ich habe früher, außer den mit dem Zulassungsgesuch urkundlichen Graden, keine weiteren akademischen Grade erworben und zu erwerben gesucht. Würzburg, Lars Hofmann Content SUMMARY ................................................................................................................ IV ZUSAMMENFASSUNG ............................................................................................. V 1. INTRODUCTION ................................................................................................. 1 1.1. Molecular basics of cancer .......................................................................................... 1 1.2. Early research on tumorigenesis ................................................................................. 3 1.3. Developing -

A Sonic Hedgehog Signaling Domain in the Arterial Adventitia Supports Resident Sca1؉ Smooth Muscle Progenitor Cells
A sonic hedgehog signaling domain in the arterial adventitia supports resident Sca1؉ smooth muscle progenitor cells Jenna N. Passman*†, Xiu Rong Dong*‡, San-Pin Wu*§, Colin T. Maguire*‡, Kelly A. Hogan¶, Victoria L. Bautch*†¶, and Mark W. Majesky*†‡ʈ** *Carolina Cardiovascular Biology Center, Departments of ‡Medicine, ʈGenetics, and ¶Biology, and †Curriculum in Genetics and Molecular Biology, University of North Carolina, Chapel Hill, NC 27599 Edited by Eric N. Olson, University of Texas Southwestern Medical Center, Dallas, TX, and approved April 22, 2008 (received for review December 6, 2007) We characterize a sonic hedgehog (Shh) signaling domain re- interstitial fibroblasts (13, 14), and promotes endothelial cell stricted to the adventitial layer of artery wall that supports resi- chemotaxis and tube formation (17, 18). Therefore, Shh plays dent Sca1-positive vascular progenitor cells (AdvSca1). Using important roles in cell–cell communication during vascular patched-1 (Ptc1lacZ) and patched-2 (Ptc2lacZ) reporter mice, adven- development and adult wound repair. titial Shh signaling activity was first detected at embryonic day (E) Hedgehog (Hh) proteins signal by binding to patched-1 (Ptc1) 15.5, reached the highest levels between postnatal day 1 (P1) and or patched-2 (Ptc2) receptors and cdo/boc accessory proteins at P10, was diminished in adult vessels, and colocalized with a the cell surface, resulting in derepression of smoothened (smo) circumferential ring of Shh protein deposited between the media (19). Ptc proteins are 12-transmembrane domain receptors that ؊/؊ and adventitia. In Shh mice, AdvSca1 cells normally found at the are structurally related to the resistance-nodulation-division aortic root were either absent or greatly diminished in number. -

Human Induced Pluripotent Stem Cell–Derived Podocytes Mature Into Vascularized Glomeruli Upon Experimental Transplantation
BASIC RESEARCH www.jasn.org Human Induced Pluripotent Stem Cell–Derived Podocytes Mature into Vascularized Glomeruli upon Experimental Transplantation † Sazia Sharmin,* Atsuhiro Taguchi,* Yusuke Kaku,* Yasuhiro Yoshimura,* Tomoko Ohmori,* ‡ † ‡ Tetsushi Sakuma, Masashi Mukoyama, Takashi Yamamoto, Hidetake Kurihara,§ and | Ryuichi Nishinakamura* *Department of Kidney Development, Institute of Molecular Embryology and Genetics, and †Department of Nephrology, Faculty of Life Sciences, Kumamoto University, Kumamoto, Japan; ‡Department of Mathematical and Life Sciences, Graduate School of Science, Hiroshima University, Hiroshima, Japan; §Division of Anatomy, Juntendo University School of Medicine, Tokyo, Japan; and |Japan Science and Technology Agency, CREST, Kumamoto, Japan ABSTRACT Glomerular podocytes express proteins, such as nephrin, that constitute the slit diaphragm, thereby contributing to the filtration process in the kidney. Glomerular development has been analyzed mainly in mice, whereas analysis of human kidney development has been minimal because of limited access to embryonic kidneys. We previously reported the induction of three-dimensional primordial glomeruli from human induced pluripotent stem (iPS) cells. Here, using transcription activator–like effector nuclease-mediated homologous recombination, we generated human iPS cell lines that express green fluorescent protein (GFP) in the NPHS1 locus, which encodes nephrin, and we show that GFP expression facilitated accurate visualization of nephrin-positive podocyte formation in -

Actin Cytoskeleton and Regulation of Tgfβ Signaling: Exploring Their Links
biomolecules Review Actin Cytoskeleton and Regulation of TGFβ Signaling: Exploring Their Links Roberta Melchionna 1,†, Paola Trono 1,2,†, Annalisa Tocci 1 and Paola Nisticò 1,* 1 Tumor Immunology and Immunotherapy Unit, IRCCS Regina Elena National Cancer Institute, via Chianesi 53, 00144 Rome, Italy; [email protected] (R.M.); [email protected] (P.T.); [email protected] (A.T.) 2 Institute of Biochemistry and Cell Biology, National Research Council, via Ramarini 32, 00015 Monterotondo Scalo, Rome, Italy * Correspondence: [email protected]; Tel.: +39-0652662539 † These authors contributed equally to this paper. Abstract: Human tissues, to maintain their architecture and function, respond to injuries by activating intricate biochemical and physical mechanisms that regulates intercellular communication crucial in maintaining tissue homeostasis. Coordination of the communication occurs through the activity of different actin cytoskeletal regulators, physically connected to extracellular matrix through integrins, generating a platform of biochemical and biomechanical signaling that is deregulated in cancer. Among the major pathways, a controller of cellular functions is the cytokine transforming growth factor β (TGFβ), which remains a complex and central signaling network still to be interpreted and explained in cancer progression. Here, we discuss the link between actin dynamics and TGFβ signaling with the aim of exploring their aberrant interaction in cancer. Keywords: actin cytoskeleton; actin-binding proteins; TGFβ; extracellular matrix; tumor microenvi- Citation: Melchionna, R.; Trono, P.; ronment Tocci, A.; Nisticò, P. Actin Cytoskeleton and Regulation of TGFβ Signaling: Exploring Their Links. Biomolecules 2021, 11, 336. 1. Introduction https://doi.org/10.3390/biom11020336 Actin dynamics critically affect different aspects of human health and disease, ranging Academic Editor: Vladimir from embryonic development to wound repair, inflammation, and cancer [1].