Primepcr™Assay Validation Report
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Supplementary Table 3 Complete List of RNA-Sequencing Analysis of Gene Expression Changed by ≥ Tenfold Between Xenograft and Cells Cultured in 10%O2
Supplementary Table 3 Complete list of RNA-Sequencing analysis of gene expression changed by ≥ tenfold between xenograft and cells cultured in 10%O2 Expr Log2 Ratio Symbol Entrez Gene Name (culture/xenograft) -7.182 PGM5 phosphoglucomutase 5 -6.883 GPBAR1 G protein-coupled bile acid receptor 1 -6.683 CPVL carboxypeptidase, vitellogenic like -6.398 MTMR9LP myotubularin related protein 9-like, pseudogene -6.131 SCN7A sodium voltage-gated channel alpha subunit 7 -6.115 POPDC2 popeye domain containing 2 -6.014 LGI1 leucine rich glioma inactivated 1 -5.86 SCN1A sodium voltage-gated channel alpha subunit 1 -5.713 C6 complement C6 -5.365 ANGPTL1 angiopoietin like 1 -5.327 TNN tenascin N -5.228 DHRS2 dehydrogenase/reductase 2 leucine rich repeat and fibronectin type III domain -5.115 LRFN2 containing 2 -5.076 FOXO6 forkhead box O6 -5.035 ETNPPL ethanolamine-phosphate phospho-lyase -4.993 MYO15A myosin XVA -4.972 IGF1 insulin like growth factor 1 -4.956 DLG2 discs large MAGUK scaffold protein 2 -4.86 SCML4 sex comb on midleg like 4 (Drosophila) Src homology 2 domain containing transforming -4.816 SHD protein D -4.764 PLP1 proteolipid protein 1 -4.764 TSPAN32 tetraspanin 32 -4.713 N4BP3 NEDD4 binding protein 3 -4.705 MYOC myocilin -4.646 CLEC3B C-type lectin domain family 3 member B -4.646 C7 complement C7 -4.62 TGM2 transglutaminase 2 -4.562 COL9A1 collagen type IX alpha 1 chain -4.55 SOSTDC1 sclerostin domain containing 1 -4.55 OGN osteoglycin -4.505 DAPL1 death associated protein like 1 -4.491 C10orf105 chromosome 10 open reading frame 105 -4.491 -

Supplementary Table 1: Adhesion Genes Data Set
Supplementary Table 1: Adhesion genes data set PROBE Entrez Gene ID Celera Gene ID Gene_Symbol Gene_Name 160832 1 hCG201364.3 A1BG alpha-1-B glycoprotein 223658 1 hCG201364.3 A1BG alpha-1-B glycoprotein 212988 102 hCG40040.3 ADAM10 ADAM metallopeptidase domain 10 133411 4185 hCG28232.2 ADAM11 ADAM metallopeptidase domain 11 110695 8038 hCG40937.4 ADAM12 ADAM metallopeptidase domain 12 (meltrin alpha) 195222 8038 hCG40937.4 ADAM12 ADAM metallopeptidase domain 12 (meltrin alpha) 165344 8751 hCG20021.3 ADAM15 ADAM metallopeptidase domain 15 (metargidin) 189065 6868 null ADAM17 ADAM metallopeptidase domain 17 (tumor necrosis factor, alpha, converting enzyme) 108119 8728 hCG15398.4 ADAM19 ADAM metallopeptidase domain 19 (meltrin beta) 117763 8748 hCG20675.3 ADAM20 ADAM metallopeptidase domain 20 126448 8747 hCG1785634.2 ADAM21 ADAM metallopeptidase domain 21 208981 8747 hCG1785634.2|hCG2042897 ADAM21 ADAM metallopeptidase domain 21 180903 53616 hCG17212.4 ADAM22 ADAM metallopeptidase domain 22 177272 8745 hCG1811623.1 ADAM23 ADAM metallopeptidase domain 23 102384 10863 hCG1818505.1 ADAM28 ADAM metallopeptidase domain 28 119968 11086 hCG1786734.2 ADAM29 ADAM metallopeptidase domain 29 205542 11085 hCG1997196.1 ADAM30 ADAM metallopeptidase domain 30 148417 80332 hCG39255.4 ADAM33 ADAM metallopeptidase domain 33 140492 8756 hCG1789002.2 ADAM7 ADAM metallopeptidase domain 7 122603 101 hCG1816947.1 ADAM8 ADAM metallopeptidase domain 8 183965 8754 hCG1996391 ADAM9 ADAM metallopeptidase domain 9 (meltrin gamma) 129974 27299 hCG15447.3 ADAMDEC1 ADAM-like, -

Investigation of the Underlying Hub Genes and Molexular Pathogensis in Gastric Cancer by Integrated Bioinformatic Analyses
bioRxiv preprint doi: https://doi.org/10.1101/2020.12.20.423656; this version posted December 22, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Investigation of the underlying hub genes and molexular pathogensis in gastric cancer by integrated bioinformatic analyses Basavaraj Vastrad1, Chanabasayya Vastrad*2 1. Department of Biochemistry, Basaveshwar College of Pharmacy, Gadag, Karnataka 582103, India. 2. Biostatistics and Bioinformatics, Chanabasava Nilaya, Bharthinagar, Dharwad 580001, Karanataka, India. * Chanabasayya Vastrad [email protected] Ph: +919480073398 Chanabasava Nilaya, Bharthinagar, Dharwad 580001 , Karanataka, India bioRxiv preprint doi: https://doi.org/10.1101/2020.12.20.423656; this version posted December 22, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Abstract The high mortality rate of gastric cancer (GC) is in part due to the absence of initial disclosure of its biomarkers. The recognition of important genes associated in GC is therefore recommended to advance clinical prognosis, diagnosis and and treatment outcomes. The current investigation used the microarray dataset GSE113255 RNA seq data from the Gene Expression Omnibus database to diagnose differentially expressed genes (DEGs). Pathway and gene ontology enrichment analyses were performed, and a proteinprotein interaction network, modules, target genes - miRNA regulatory network and target genes - TF regulatory network were constructed and analyzed. Finally, validation of hub genes was performed. The 1008 DEGs identified consisted of 505 up regulated genes and 503 down regulated genes. -

Atlas Journal
Atlas of Genetics and Cytogenetics in Oncology and Haematology Home Genes Leukemias Solid Tumours Cancer-Prone Deep Insight Portal Teaching X Y 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 NA Atlas Journal Atlas Journal versus Atlas Database: the accumulation of the issues of the Journal constitutes the body of the Database/Text-Book. TABLE OF CONTENTS Volume 12, Number 6, Nov-Dec 2008 Previous Issue / Next Issue Genes BCL8 (B-cell CLL/lymphoma 8) (15q11). Silvia Rasi, Gianluca Gaidano. Atlas Genet Cytogenet Oncol Haematol 2008; 12 (6): 781-784. [Full Text] [PDF] URL : http://atlasgeneticsoncology.org/Genes/BCL8ID781ch15q11.html CDC25A (Cell division cycle 25A) (3p21). Dipankar Ray, Hiroaki Kiyokawa. Atlas Genet Cytogenet Oncol Haematol 2008; 12 (6): 785-791. [Full Text] [PDF] URL : http://atlasgeneticsoncology.org/Genes/CDC25AID40004ch3p21.html CDC73 (cell division cycle 73, Paf1/RNA polymerase II complex component, homolog (S. cerevisiae)) (1q31.2). Leslie Farber, Bin Tean Teh. Atlas Genet Cytogenet Oncol Haematol 2008; 12 (6): 792-797. [Full Text] [PDF] URL : http://atlasgeneticsoncology.org/Genes/CDC73D181ch1q31.html EIF3C (eukaryotic translation initiation factor 3, subunit C) (16p11.2). Daniel R Scoles. Atlas Genet Cytogenet Oncol Haematol 2008; 12 (6): 798-802. [Full Text] [PDF] URL : http://atlasgeneticsoncology.org/Genes/EIF3CID44187ch16p11.html ELAC2 (elaC homolog 2 (E. coli)) (17p11.2). Yang Chen, Sean Tavtigian, Donna Shattuck. Atlas Genet Cytogenet Oncol Haematol 2008; 12 (6): 803-806. [Full Text] [PDF] URL : http://atlasgeneticsoncology.org/Genes/ELAC2ID40437ch17p11.html FOXM1 (forkhead box M1) (12p13). Jamila Laoukili, Monica Alvarez Fernandez, René H Medema. -

RSPO4 Is the Major Gene in Autosomal-Recessive Anonychia
View metadata, citation and similar papers at core.ac.uk brought to you by CORE provided by Elsevier - Publisher Connector ORIGINAL ARTICLE RSPO4 Is the Major Gene in Autosomal-Recessive Anonychia and Mutations Cluster in the Furin-Like Cysteine-Rich Domains of the Wnt Signaling Ligand R-spondin 4 Nadina Ortiz Bru¨chle1, Jorge Frank2,3, Valeska Frank1, Jan Senderek1, Ahmet Akar4, Erol Koc4, Dimitris Rigopoulos5, Maurice van Steensel2,3, Klaus Zerres1 and Carsten Bergmann1 Congenital anonychia is a rare autosomal-recessive disorder characterized by the absence of finger- and toenails. Recently, we and others identified the secreted Wnt signaling ligand R-spondin 4 (RSPO4) as the first gene known to be responsible for inherited anonychia. R-spondins are secreted proteins that activate the Wnt/ b-catenin signaling pathway. This puts anonychia on the growing list of congenital malformation syndromes caused by Wnt signaling pathway defects. Here, we expand the RSPO4 mutational spectrum by identification of the previously unknown mutations c.190C4T (p.Arg64Cys) in exon 2 and c.301C4T (p.Gln101X) in exon 3, thereby corroborating R-spondin 4 as the major protein in autosomal-recessive anonychia. Almost all RSPO4 mutations detected so far affect the highly conserved exons 2 and 3. Thus, we postulate that RSPO4 mutations preferentially cluster in the furin-like cysteine-rich domains of R-spondin 4, which is in line with experimental data proposing that for b-catenin stabilization, a shortened protein comprising just these two regions is sufficient. Journal of Investigative Dermatology (2008) 128, 791–796; doi:10.1038/sj.jid.5701088; published online 4 October 2007 INTRODUCTION even within a given family. -

Altered Gene Expression in Circulating Immune Cells Following a 24-Hour Passive Dehydration
University of Connecticut OpenCommons@UConn Master's Theses University of Connecticut Graduate School 5-10-2020 Altered Gene Expression in Circulating Immune Cells Following a 24-Hour Passive Dehydration Aidan Fiol [email protected] Follow this and additional works at: https://opencommons.uconn.edu/gs_theses Recommended Citation Fiol, Aidan, "Altered Gene Expression in Circulating Immune Cells Following a 24-Hour Passive Dehydration" (2020). Master's Theses. 1480. https://opencommons.uconn.edu/gs_theses/1480 This work is brought to you for free and open access by the University of Connecticut Graduate School at OpenCommons@UConn. It has been accepted for inclusion in Master's Theses by an authorized administrator of OpenCommons@UConn. For more information, please contact [email protected]. Altered Gene Expression in Circulating Immune Cells Following a 24- Hour Passive Dehydration Aidan Fiol B.S., University of Connecticut, 2018 A Thesis Submitted in Partial Fulfillment of the Requirements for the Degree of Master of Science At the University of Connecticut 2020 i copyright by Aidan Fiol 2020 ii APPROVAL PAGE Master of Science Thesis Altered Gene Expression in Circulating Immune Cells Following a 24- Hour Passive Dehydration Presented by Aidan Fiol, B.S. Major Advisor__________________________________________________________________ Elaine Choung-Hee Lee, Ph.D. Associate Advisor_______________________________________________________________ Douglas J. Casa, Ph.D. Associate Advisor_______________________________________________________________ Robert A. Huggins, Ph.D. University of Connecticut 2020 iii ACKNOWLEDGEMENTS I’d like to thank my committee members. Dr. Lee, you have been an amazing advisor, mentor and friend to me these past few years. Your advice, whether it was how to be a better writer or scientist, or just general life advice given on our way to get coffee, has helped me grow as a researcher and as a person. -

WO 2014/028907 Al 20 February 2014 (20.02.2014) P O P C T
(12) INTERNATIONAL APPLICATION PUBLISHED UNDER THE PATENT COOPERATION TREATY (PCT) (19) World Intellectual Property Organization International Bureau (10) International Publication Number (43) International Publication Date WO 2014/028907 Al 20 February 2014 (20.02.2014) P O P C T (51) International Patent Classification: (74) Agents: EVANS, Judith et al; P.O. Box 23 1, Manassas, C12Q 1/68 (2006.01) G01N 33/00 (2006.01) VA 20108 (US). (21) International Application Number: (81) Designated States (unless otherwise indicated, for every PCT/US20 13/055469 kind of national protection available): AE, AG, AL, AM, AO, AT, AU, AZ, BA, BB, BG, BH, BN, BR, BW, BY, (22) International Filing Date: BZ, CA, CH, CL, CN, CO, CR, CU, CZ, DE, DK, DM, 16 August 2013 (16.08.2013) DO, DZ, EC, EE, EG, ES, FI, GB, GD, GE, GH, GM, GT, (25) Filing Language: English HN, HR, HU, ID, IL, IN, IS, JP, KE, KG, KN, KP, KR, KZ, LA, LC, LK, LR, LS, LT, LU, LY, MA, MD, ME, (26) Publication Language: English MG, MK, MN, MW, MX, MY, MZ, NA, NG, NI, NO, NZ, (30) Priority Data: OM, PA, PE, PG, PH, PL, PT, QA, RO, RS, RU, RW, SA, 61/684,029 16 August 2012 (16.08.2012) US SC, SD, SE, SG, SK, SL, SM, ST, SV, SY, TH, TJ, TM, 61/718,468 25 October 2012 (25. 10.2012) US TN, TR, TT, TZ, UA, UG, US, UZ, VC, VN, ZA, ZM, 61/745,207 2 1 December 2012 (21. 12.2012) US ZW. (71) Applicant: THE TRUSTEES OF COLUMBIA UNI¬ (84) Designated States (unless otherwise indicated, for every VERSITY IN THE CITY OF NEW YORK [US/US]; kind of regional protection available): ARIPO (BW, GH, 412 Low Memorial Library, 535 West 116th Street, New GM, KE, LR, LS, MW, MZ, NA, RW, SD, SL, SZ, TZ, York, NY 10027 (US). -
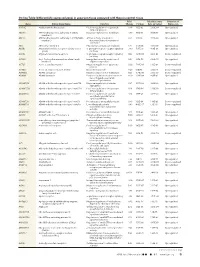
Differentially Expressed Genes in Aneurysm Tissue Compared With
On-line Table: Differentially expressed genes in aneurysm tissue compared with those in control tissue Fold False Discovery Direction of Gene Entrez Gene Name Function Change P Value Rate (q Value) Expression AADAC Arylacetamide deacetylase Positive regulation of triglyceride 4.46 1.33E-05 2.60E-04 Up-regulated catabolic process ABCA6 ATP-binding cassette, subfamily A (ABC1), Integral component of membrane 3.79 9.15E-14 8.88E-12 Up-regulated member 6 ABCC3 ATP-binding cassette, subfamily C (CFTR/MRP), ATPase activity, coupled to 6.63 1.21E-10 7.33E-09 Up-regulated member 3 transmembrane movement of substances ABI3 ABI family, member 3 Peptidyl-tyrosine phosphorylation 6.47 2.47E-05 4.56E-04 Up-regulated ACKR1 Atypical chemokine receptor 1 (Duffy blood G-protein–coupled receptor signaling 3.80 7.95E-10 4.18E-08 Up-regulated group) pathway ACKR2 Atypical chemokine receptor 2 G-protein–coupled receptor signaling 0.42 3.29E-04 4.41E-03 Down-regulated pathway ACSM1 Acyl-CoA synthetase medium-chain family Energy derivation by oxidation of 9.87 1.70E-08 6.52E-07 Up-regulated member 1 organic compounds ACTC1 Actin, ␣, cardiac muscle 1 Negative regulation of apoptotic 0.30 7.96E-06 1.65E-04 Down-regulated process ACTG2 Actin, ␥2, smooth muscle, enteric Blood microparticle 0.29 1.61E-16 2.36E-14 Down-regulated ADAM33 ADAM domain 33 Integral component of membrane 0.23 9.74E-09 3.95E-07 Down-regulated ADAM8 ADAM domain 8 Positive regulation of tumor necrosis 4.69 2.93E-04 4.01E-03 Up-regulated factor (ligand) superfamily member 11 production ADAMTS18 -

Atlas Journal
Atlas of Genetics and Cytogenetics in Oncology and Haematology Home Genes Leukemias Solid Tumours Cancer-Prone Deep Insight Portal Teaching X Y 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 NA Atlas Journal Atlas Journal versus Atlas Database: the accumulation of the issues of the Journal constitutes the body of the Database/Text-Book. TABLE OF CONTENTS Volume 12, Number 4, Jul-Aug 2008 Previous Issue / Next Issue Genes AKR1C3 (aldo-keto reductase family 1, member C3 (3-alpha hydroxysteroid dehydrogenase, type II)) (10p15.1). Hsueh Kung Lin. Atlas Genet Cytogenet Oncol Haematol 2008; Vol (12): 498-502. [Full Text] [PDF] URL : http://atlasgeneticsoncology.org/Genes/AKR1C3ID612ch10p15.html CASP1 (caspase 1, apoptosis-related cysteine peptidase (interleukin 1, beta, convertase)) (11q22.3). Yatender Kumar, Vegesna Radha, Ghanshyam Swarup. Atlas Genet Cytogenet Oncol Haematol 2008; Vol (12): 503-518. [Full Text] [PDF] URL : http://atlasgeneticsoncology.org/Genes/CASP1ID145ch11q22.html GCNT3 (glucosaminyl (N-acetyl) transferase 3, mucin type) (15q21.3). Prakash Radhakrishnan, Pi-Wan Cheng. Atlas Genet Cytogenet Oncol Haematol 2008; Vol (12): 519-524. [Full Text] [PDF] URL : http://atlasgeneticsoncology.org/Genes/GCNT3ID44105ch15q21.html HYAL2 (Hyaluronoglucosaminidase 2) (3p21.3). Lillian SN Chow, Kwok-Wai Lo. Atlas Genet Cytogenet Oncol Haematol 2008; Vol (12): 525-529. [Full Text] [PDF] URL : http://atlasgeneticsoncology.org/Genes/HYAL2ID40904ch3p21.html LMO2 (LIM domain only 2 (rhombotin-like 1)) (11p13) - updated. Pieter Van Vlierberghe, Jean Loup Huret. Atlas Genet Cytogenet Oncol Haematol 2008; Vol (12): 530-535. [Full Text] [PDF] URL : http://atlasgeneticsoncology.org/Genes/RBTN2ID34.html PEBP1 (phosphatidylethanolamine binding protein 1) (12q24.23). -

Transcriptome-Wide Analysis Reveals Hallmarks of Human Intestine Development and Maturation in Vitro and in Vivo Stacy R
Stem Cell Reports, Volume 4 Supplemental Information Transcriptome-wide Analysis Reveals Hallmarks of Human Intestine Development and Maturation In Vitro and In Vivo Stacy R. Finkbeiner, David R. Hill, Christopher H. Altheim, Priya H. Dedhia, Matthew J. Taylor, Yu-Hwai Tsai, Alana M. Chin, Maxime M. Mahe, Carey L. Watson, Jennifer J. Freeman, Roy Nattiv, Matthew Thomson, Ophir D. Klein, Noah F. Shroyer, Michael A. Helmrath, Daniel H. Teitelbaum, Peter J. Dempsey, and Jason R. Spence Figure S1, related to Figure 1. (A) Principal component analysis was conducted HIOs (n=3 independently derived H9- HIOs), adult distal small intestine (n=4 independent biological specimens), adult duodenum (n=2 independent biological specimens), and fetal small intestine (n=6 independent biological specimens) (see also Table S1). The top two principal components account for 76.8% of the variation in the data and show that HIOs group with fetal intestinal tissues. (B) A dendrogram was generated based on hierarchical clustering of the gene sets using Canberra distance (B). Fetal tissues are more similar to each other than to their respective adult tissues and HIOs cluster within the same clade as the fetal tissues. Branch lengths indicate the level of dissimilarity between samples, with longer branch lengths indicating a greater level of dissimilarity. Red labels at branch point correspond to the Approximately Unbiased (AU) p- value. AU > 95 indicates that a given branch assignment is strongly supported by the data. Green labels at each branch point correspond to the bootstrap probability (BP) of a cluster, defined as the frequency of a given relationship among the bootstrap replicates. -

Homozygosity Mapping and Sequencing Identify Two Genes That
Akkad et al. Canine Genetics and Epidemiology (2015) 2:5 DOI 10.1186/s40575-015-0018-5 RESEARCH Open Access Homozygosity mapping and sequencing identify two genes that might contribute to pointing behavior in hunting dogs Denis A Akkad1*†, Wanda M Gerding1†, Robin B Gasser2 and Jörg T Epplen1,3 Abstract Background: The domestic dog represents an important model for studying the genetics of behavior. In spite of technological advances in genomics and phenomics, the genetic basis of most specific canine behaviors is largely unknown. Some breeds of hunting dogs exhibit a behavioral trait called “pointing” (a prolonged halt of movement to indicate the position of a game animal). Here, the genomes of pointing dogs (Large Munsterlander and Weimaraner) were compared with those of behaviorally distinct herding dogs (Berger des Pyrenées and Schapendoes). We assumed (i) that these four dog breeds initially represented inbred populations and (ii) that selective breeding for pointing behavior promotes an enrichment of the genetic trait in a homozygous state. Results: The homozygosity mapping of 52 dogs (13 of each of the four breeds) followed by subsequent interval resequencing identified fixed genetic differences on chromosome 22 between pointers and herding dogs. In addition, we identified one non-synonomous variation in each of the coding genes SETDB2 and CYSLTR2 that might have a functional consequence. Genetic analysis of additional hunting and non-hunting dogs revealed consistent homozygosity forthesetwovariationsinsixofsevenpointingbreeds. Conclusions: Based on the present findings, we propose that, together with other genetic, training and/or environmental factors, the nucleotide and associated amino acid variations identified in genes SETDB2 and CYSLTR2 contribute to pointing behavior. -

A Personalized Genomics Approach of the Prostate Cancer
cells Article A Personalized Genomics Approach of the Prostate Cancer Sanda Iacobas 1 and Dumitru A. Iacobas 2,* 1 Department of Pathology, New York Medical College, Valhalla, NY 10595, USA; [email protected] 2 Personalized Genomics Laboratory, Center for Computational Systems Biology, Roy G Perry College of Engineering, Prairie View A&M University, Prairie View, TX 77446, USA * Correspondence: [email protected]; Tel.: +1-936-261-9926 Abstract: Decades of research identified genomic similarities among prostate cancer patients and proposed general solutions for diagnostic and treatments. However, each human is a dynamic unique with never repeatable transcriptomic topology and no gene therapy is good for everybody. Therefore, we propose the Genomic Fabric Paradigm (GFP) as a personalized alternative to the biomarkers approach. Here, GFP is applied to three (one primary—“A”, and two secondary—“B” & “C”) cancer nodules and the surrounding normal tissue (“N”) from a surgically removed prostate tumor. GFP proved for the first time that, in addition to the expression levels, cancer alters also the cellular control of the gene expression fluctuations and remodels their networking. Substantial differences among the profiled regions were found in the pathways of P53-signaling, apoptosis, prostate cancer, block of differentiation, evading apoptosis, immortality, insensitivity to anti-growth signals, proliferation, resistance to chemotherapy, and sustained angiogenesis. ENTPD2, AP5M1 BAIAP2L1, and TOR1A were identified as the master regulators of the “A”, “B”, “C”, and “N” regions, and potential consequences of ENTPD2 manipulation were analyzed. The study shows that GFP can fully characterize the transcriptomic complexity of a heterogeneous prostate tumor and identify the most influential genes in each cancer nodule.