Bioinformatics Approach for the Identification of Fragile Regions On
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

The First Case of Congenital Myasthenic Syndrome Caused by A
G C A T T A C G G C A T genes Case Report The First Case of Congenital Myasthenic Syndrome Caused by a Large Homozygous Deletion in the C-Terminal Region of COLQ (Collagen Like Tail Subunit of Asymmetric Acetylcholinesterase) Protein Nicola Laforgia 1 , Lucrezia De Cosmo 1, Orazio Palumbo 2 , Carlotta Ranieri 3, Michela Sesta 4, Donatella Capodiferro 1, Antonino Pantaleo 3 , Pierluigi Iapicca 5 , Patrizia Lastella 6, Manuela Capozza 1 , Federico Schettini 1, Nenad Bukvic 7 , Rosanna Bagnulo 3 and Nicoletta Resta 3,7,* 1 Section of Neonatology and Neonatal Intensive Care Unit, Department of Biomedical Science and Human Oncology (DIMO), University of Bari “Aldo Moro”, 70124 Bari, Italy; [email protected] (N.L.); [email protected] (L.D.C.); [email protected] (D.C.); [email protected] (M.C.); [email protected] (F.S.) 2 Division of Medical Genetics, Fondazione IRCCS Casa Sollievo della Sofferenza, 71013 San Giovanni Rotondo, Italy; [email protected] 3 Division of Medical Genetics, Department of Biomedical Sciences and Human Oncology (DIMO), University of Bari “Aldo Moro”, 70124 Bari, Italy; [email protected] (C.R.); [email protected] (A.P.); [email protected] (R.B.) 4 Neurology Unit, University Hospital Consortium Corporation Polyclinic of Bari, 70124 Bari, Italy; [email protected] 5 SOPHiA GENETICS SA HQ, 1025 Saint-Sulpice, Switzerland; [email protected] 6 Rare Diseases Centre—Internal Medicine Unit “C. Frugoni”, Polyclinic of Bari, 70124 Bari, Italy; [email protected] 7 Medical Genetics Section, University Hospital Consortium Corporation Polyclinic of Bari, 70124 Bari, Italy; [email protected] * Correspondence: [email protected]; Tel.: +39-0805593619 Received: 17 November 2020; Accepted: 15 December 2020; Published: 18 December 2020 Abstract: Congenital myasthenic syndromes (CMSs) are caused by mutations in genes that encode proteins involved in the organization, maintenance, function, or modification of the neuromuscular junction. -

Limb Girdle Myasthenia with Digenic RAPSN and a Novel Disease Gene AK9 Mutations
European Journal of Human Genetics (2017) 25, 192–199 & 2017 Macmillan Publishers Limited, part of Springer Nature. All rights reserved 1018-4813/17 www.nature.com/ejhg ARTICLE Limb girdle myasthenia with digenic RAPSN and a novel disease gene AK9 mutations Ching-Wan Lam*,1,3, Ka-Sing Wong2,3, Ho-Wan Leung2 and Chun-Yiu Law1 Though dysfunction of neuromuscular junction (NMJ) is associated with congenital myasthenic syndrome (CMS), the proteins involved in neuromuscular transmission have not been completely identified. In this study, we aimed to identify a novel CMS gene in a consanguineous family with limb-girdle type CMS. Homozygosity mapping of the novel CMS gene was performed using high-density single-nucleotide polymorphism microarrays. The variants in CMS gene were identified by whole-exome sequencing (WES) and Sanger sequencing. A 20 MB-region of homozygosity (ROH) was mapped on chromosome 6q15–21. This was the only ROH that present in all clinically affected siblings and absent in all clinically unaffected siblings. WES showed a novel variant of AK9 gene located in this ROH. This variant was a start-gain mutation and introduced a cryptic 5′-UTR signal in intron 5 of the AK9 gene. The normal splicing signal would be interfered by the cryptic translation signal leading to defective splicing. Another 25 MB-ROH was found on chromosome 11p13–q12 in all siblings. WES showed a homozygous RAPSN pathogenic variant in this ROH. Since RAPSN-associated limb-girdle type CMS was only manifested in AK9 homozygous variant carriers, the disease phenotype was of digenic inheritance, and was determined by the novel disease modifier AK9 which provides NTPs for N-glycosylation. -

Associated with Low Dehydrodolichol Diphosphate Synthase (DHDDS) Activity S
Sabry et al. Orphanet Journal of Rare Diseases (2016) 11:84 DOI 10.1186/s13023-016-0468-1 RESEARCH Open Access A case of fatal Type I congenital disorders of glycosylation (CDG I) associated with low dehydrodolichol diphosphate synthase (DHDDS) activity S. Sabry1,2,3,4, S. Vuillaumier-Barrot1,2,5, E. Mintet1,2, M. Fasseu1,2, V. Valayannopoulos6, D. Héron7,8, N. Dorison8, C. Mignot7,8,9, N. Seta5,10, I. Chantret1,2, T. Dupré1,2,5 and S. E. H. Moore1,2* Abstract Background: Type I congenital disorders of glycosylation (CDG-I) are mostly complex multisystemic diseases associated with hypoglycosylated serum glycoproteins. A subgroup harbour mutations in genes necessary for the biosynthesis of the dolichol-linked oligosaccharide (DLO) precursor that is essential for protein N-glycosylation. Here, our objective was to identify the molecular origins of disease in such a CDG-Ix patient presenting with axial hypotonia, peripheral hypertonia, enlarged liver, micropenis, cryptorchidism and sensorineural deafness associated with hypo glycosylated serum glycoproteins. Results: Targeted sequencing of DNA revealed a splice site mutation in intron 5 and a non-sense mutation in exon 4 of the dehydrodolichol diphosphate synthase gene (DHDDS). Skin biopsy fibroblasts derived from the patient revealed ~20 % residual DHDDS mRNA, ~35 % residual DHDDS activity, reduced dolichol-phosphate, truncated DLO and N-glycans, and an increased ratio of [2-3H]mannose labeled glycoprotein to [2-3H]mannose labeled DLO. Predicted truncated DHDDS transcripts did not complement rer2-deficient yeast. SiRNA-mediated down-regulation of DHDDS in human hepatocellular carcinoma HepG2 cells largely mirrored the biochemical phenotype of cells from the patient. -

Genetic Background of Ataxia in Children Younger Than 5 Years in Finland E444
Volume 6, Number 4, August 2020 Neurology.org/NG A peer-reviewed clinical and translational neurology open access journal ARTICLE Genetic background of ataxia in children younger than 5 years in Finland e444 ARTICLE Cerebral arteriopathy associated with heterozygous variants in the casitas B-lineage lymphoma gene e448 ARTICLE Somatic SLC35A2 mosaicism correlates with clinical fi ndings in epilepsy brain tissuee460 ARTICLE Synonymous variants associated with Alzheimer disease in multiplex families e450 Academy Officers Neurology® is a registered trademark of the American Academy of Neurology (registration valid in the United States). James C. Stevens, MD, FAAN, President Neurology® Genetics (eISSN 2376-7839) is an open access journal published Orly Avitzur, MD, MBA, FAAN, President Elect online for the American Academy of Neurology, 201 Chicago Avenue, Ann H. Tilton, MD, FAAN, Vice President Minneapolis, MN 55415, by Wolters Kluwer Health, Inc. at 14700 Citicorp Drive, Bldg. 3, Hagerstown, MD 21742. Business offices are located at Two Carlayne E. Jackson, MD, FAAN, Secretary Commerce Square, 2001 Market Street, Philadelphia, PA 19103. Production offices are located at 351 West Camden Street, Baltimore, MD 21201-2436. Janis M. Miyasaki, MD, MEd, FRCPC, FAAN, Treasurer © 2020 American Academy of Neurology. Ralph L. Sacco, MD, MS, FAAN, Past President Neurology® Genetics is an official journal of the American Academy of Neurology. Journal website: Neurology.org/ng, AAN website: AAN.com CEO, American Academy of Neurology Copyright and Permission Information: Please go to the journal website (www.neurology.org/ng) and click the Permissions tab for the relevant Mary E. Post, MBA, CAE article. Alternatively, send an email to [email protected]. -

Novel Exopolysaccharides Produced by Lactococcus Lactis Subsp
Biosci. Biotechnol. Biochem., 77 (10), 2013–2018, 2013 Novel Exopolysaccharides Produced by Lactococcus lactis subsp. lactis, and the Diversity of epsE Genes in the Exopolysaccharide Biosynthesis Gene Clusters y Chise SUZUKI, Miho KOBAYASHI, and Hiromi KIMOTO-NIRA NARO Institute of Livestock and Grassland Science, 2 Ikenodai, Tsukuba, Ibaraki 305-0901, Japan Received April 19, 2013; Accepted June 26, 2013; Online Publication, October 7, 2013 [doi:10.1271/bbb.130322] To characterize novel variations of exopolysacchar- ovalbumin-sensitized mice.12) Strain C59 also showed ides (EPSs) produced by dairy strains of Lactococcus significant activity as a plasminogen activator.13) This lactis subsp. lactis and subsp. cremoris, the EPSs of five activity was extracted from bacterial cells by 0.1 M dairy strains of L. lactis were purified. Sugar composi- sodium carbonate buffer (pH 10.0), suggesting that tion analysis showed two novel EPSs produced by cell-surface materials of strain C59 are involved in it. strains of L. lactis subsp. lactis. One strain produced EPS and CPS synthesis is usually controlled by eps EPS lacking galactose, and the other produced EPS operons located on a chromosome or plasmid, and by containing fucose. Among the eps gene clusters of these genes outside the operon.6,14–16) Clusters can differ strains, the highly conserved epsD and its neighboring greatly among strains belonging to the same species, but epsE were sequenced. Sequence and PCR analysis the first six genes, epsRXABCD, in lactococcal eps revealed that epsE genes were strain-specific. By South- operons are highly conserved among strains.6,15,17,18) A ern blot analysis using epsD, the eps gene cluster in each tyrosine (Tyr) phosphorylation regulatory system is strain was found to locate to the chromosome or a very involved in the modulation of polysaccharide synthesis. -

Table S1 Genes with Coefficients of Variation (CV) Ratios (CVGHS/CVSD) in the Upper and Lower 10% of for Expression in Genetic Hypercalciuric Stone-Forming (GHS) And
Table S1 Genes with coefficients of variation (CV) ratios (CVGHS/CVSD) in the upper and lower 10% of for expression in Genetic Hypercalciuric Stone-forminG (GHS) and Sprague-Dawley (SD) rats, assayed usinG Affymetrix Rat Genome 230 GeneChips. The location of the microsatellite or quantitative trait locus (QTL) position (Loc) in base pairs (bp), gene initiation base pair (BP), chromosomal banding position and RGD or GenBank name are given.Rat Genome Database (RGD) (www.rgd.mcw.edu), GenBank (http://www.ncbi.nlm.nih.gov/Genbank/) and Entrez (http://www.ncbi.nlm.nih.gov/sites/entrez). Variance in gene expression was ranked according to CVGHS/CVSD ratio for all renal (8846 genes, Pctl, Renal) and duodenal genes (9038 genes, Pctl, Duod). Gene Pctl, Pctl, QTL Gene (GenBank) Renal Duod BP GenBank/RGD Description D1Mit95 Riok2 (BG371773) 91.3 52.4 55,897,246 RIO kinase 2 66,832,211 Tmc4 (BF545988) - 93.6 63,582,715 transmembrane channel-like gene family 4 Isoc2b (BM385414) 2.3 - 67,689,183 Isochorismatase domain containing 2b Suv420h2 (AW525235) 69.9 97.3 67,815,980 suppressor of variegation 4-20 homolog 2 (Drosophila) Lilrb4 (BF282961) 95.5 - 69,165,497 leukocyte immunoglobulin-like receptor, subfamily B, member 4 Zbtb45 (BI289556) - 10.0 72,912,203 zinc finger and BTB domain containing 45 Sepw1 (NM_013027) 7.5 - 76,249,869 selenoprotein W, muscle 1 Chmp2a (AW434104) 0.1 86.9 72,889,348 chromatin modifying protein 2a Tmem160 (AWS25031) 97.6 2.2 76,716,254 transmembrane protein Pnmal2 (BI282311) 99.6 - 77,277,341 PNMA-like 2 Dmpk (AI044427) 98.9 7.3 78,449,323 dystrophia myotonica-protein kinase; serine-threonine kinase Vasp (AW520792) 99.3 - 78,621,478 vasodilator-stimulated phosphoprotein Ercc1 (AA892791) 90.6 - 78,711,248 excision repair cross-complementing rodent repair deficiency, complementation G. -

A Sensitized Mutagenesis Screen in Factor V Leiden Mice Identifies Novel Thrombosis
bioRxiv preprint doi: https://doi.org/10.1101/080432; this version posted February 12, 2017. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. A sensitized mutagenesis screen in Factor V Leiden mice identifies novel thrombosis suppressor loci Randal J. Westricka,b,c,h, Kärt Tombergc,e,h, Amy E. Sieberta, Guojing Zhuc, Mary E. Winnd, Sarah L. Dobiesc, Sara L. Manningc, Marisa A. Brakea, Audrey C. Cleurenc, Linzi M. Hobbsa, Lena M. Mishacka, Alexander Johnstona, Emilee N. Kotnikc, David R. Siemieniakf, Jishu Xue, Jun Z. Lie, Thomas L. Saunderse and David Ginsburgc,e,f,h aOakland University Department of Biological Sciences bOakland University Center for Data Science and Big Data Analysis cLife Sciences Institute, University of Michigan dBioinformatics and Biostatistics Core, Van Andel Research Institute eDepartment of Human Genetics, University of Michigan fHoward Hughes Medical Institute, University of Michigan gTransgenic Animal Model Core, University of Michigan hDepartments of Internal Medicine and Pediatrics, University of Michigan iThese authors contributed equally to this work Co-corresponding authors: David Ginsburg, MD. 5214 LSI Building, 210 Washtenaw Avenue, Ann Arbor MI 48109. E- mail: [email protected], Telephone: 734-647-4808, Fax: 734-936-2888. Randal Westrick, PhD. 118 Library Drive, 305 Dodge Hall, Rochester MI, 48309-4479 E-mail: [email protected], Telephone: 248-370-3577. 1 bioRxiv preprint doi: https://doi.org/10.1101/080432; this version posted February 12, 2017. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. -

Enrichment of in Vivo Transcription Data from Dietary Intervention
Hulst et al. Genes & Nutrition (2017) 12:11 DOI 10.1186/s12263-017-0559-1 RESEARCH Open Access Enrichment of in vivo transcription data from dietary intervention studies with in vitro data provides improved insight into gene regulation mechanisms in the intestinal mucosa Marcel Hulst1,3* , Alfons Jansman2, Ilonka Wijers1, Arjan Hoekman1, Stéphanie Vastenhouw3, Marinus van Krimpen2, Mari Smits1,3 and Dirkjan Schokker1 Abstract Background: Gene expression profiles of intestinal mucosa of chickens and pigs fed over long-term periods (days/ weeks) with a diet rich in rye and a diet supplemented with zinc, respectively, or of chickens after a one-day amoxicillin treatment of chickens, were recorded recently. Such dietary interventions are frequently used to modulate animal performance or therapeutically for monogastric livestock. In this study, changes in gene expression induced by these three interventions in cultured “Intestinal Porcine Epithelial Cells” (IPEC-J2) recorded after a short-term period of 2 and 6 hours, were compared to the in vivo gene expression profiles in order to evaluate the capability of this in vitro bioassay in predicting in vivo responses. Methods: Lists of response genes were analysed with bioinformatics programs to identify common biological pathways induced in vivo as well as in vitro. Furthermore, overlapping genes and pathways were evaluated for possible involvement in the biological processes induced in vivo by datamining and consulting literature. Results: For all three interventions, only a limited number of identical genes and a few common biological processes/ pathways were found to be affected by the respective interventions. However, several enterocyte-specific regulatory and secreted effector proteins that responded in vitro could be related to processes regulated in vivo, i.e. -
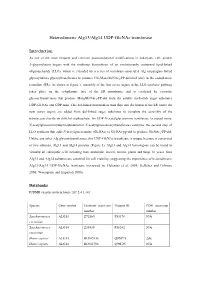
Alg13/Alg14 UDP-Glcnac Transferase
Heterodimeric Alg13/Alg14 UDP-GlcNAc transferase Introduction As one of the most frequent and common posttranslational modifications in eukaryotic cell, protein N-glycosylation begins with the multistep biosynthesis of an evolutionarily conserved lipid-linked oligosaccharide (LLO), which is extended by a series of membrane-associated Alg (asparagine-linked glycosylation) glycosyltransferases to produce Glc3Man9GlcNAc2-PP-dolichol (dol) in the endoplasmic reticulum (ER). As shown in figure 1, assembly of the first seven sugars in the LLO synthetic pathway takes place on the cytoplasmic face of the ER membrane, and is catalyzed by cytosolic glycosyltransferases that produce Man5GlcNAc2-PP-dol from the soluble nucleotide sugar substrates UDP-GlcNAc and GDP-man. This dol-linked intermediate then flips into the lumen of the ER where the next seven sugars are added from dol-linked sugar substrates to complete the assembly of the tetradecasaccharide on dolichol diphosphate. An UDP-N-acetylglucosamine transferase (accepted name: N-acetylglucosaminyldiphosphodolichol N-acetylglucosaminyltransferase) catalyzes the second step of LLO synthesis that adds N-acetylglucosamine (GlcNAc) to GlcNAc-pp-dol to produce GlcNAc2-PP-dol. Unlike any other Alg glycosyltransferases, this UDP-GlcNAc transferase is unique because it comprised of two subunits, Alg13 and Alg14 proteins (Figure 1). Alg13 and Alg14 homologous can be found in virtually all eukaryotic cells including man, mammals, insects, worms, plants and fungi. In yeast, both Alg13 and Alg14 subunits -

Small Chromosomal Regions Position Themselves Autonomously According to Their Chromatin Class
Downloaded from genome.cshlp.org on October 6, 2021 - Published by Cold Spring Harbor Laboratory Press Research Small chromosomal regions position themselves autonomously according to their chromatin class Harmen J.G. van de Werken,1,2 Josien C. Haan,3 Yana Feodorova,4,7 Dominika Bijos,5 An Weuts,3 Koen Theunis,3 Sjoerd J.B. Holwerda,2,8 Wouter Meuleman,5,9 Ludo Pagie,5 Katharina Thanisch,4,10 Parveen Kumar,3 Heinrich Leonhardt,4 Peter Marynen,6 † Bas van Steensel,5 Thierry Voet,3 Wouter de Laat,2 Irina Solovei,4 and Boris Joffe4, 1Cancer Computational Biology Center, Erasmus MC Cancer Institute & Department of Urology, Erasmus MC Cancer Institute, Erasmus University Medical Center, 3015 CN, Rotterdam, The Netherlands; 2Hubrecht Institute-KNAW & University Medical Center Utrecht, 3584 CT Utrecht, The Netherlands; 3Laboratory of Reproductive Genomics, Department of Human Genetics, KU Leuven, Leuven, 3000, Belgium; 4Department of Biology II, Ludwig Maximilians University Munich, 82152 Planegg-Martinsried, Germany; 5Division of Gene Regulation, Netherlands Cancer Institute, 1066 CX Amsterdam, The Netherlands; 6Human Genome Laboratory, Department of Human Genetics, KU Leuven, Leuven, 3000, Belgium The spatial arrangement of chromatin is linked to the regulation of nuclear processes. One striking aspect of nuclear orga- nization is the spatial segregation of heterochromatic and euchromatic domains. The mechanisms of this chromatin segre- gation are still poorly understood. In this work, we investigated the link between the primary genomic sequence and chromatin domains. We analyzed the spatial intranuclear arrangement of a human artificial chromosome (HAC) in a xen- ospecific mouse background in comparison to an orthologous region of native mouse chromosome. -

Investigating Developmental and Epileptic Encephalopathy Using Drosophila Melanogaster
International Journal of Molecular Sciences Review Investigating Developmental and Epileptic Encephalopathy Using Drosophila melanogaster Akari Takai 1 , Masamitsu Yamaguchi 2,3, Hideki Yoshida 2 and Tomohiro Chiyonobu 1,* 1 Department of Pediatrics, Graduate School of Medical Science, Kyoto Prefectural University of Medicine, Kyoto 602-8566, Japan; [email protected] 2 Department of Applied Biology, Kyoto Institute of Technology, Matsugasaki, Sakyo-ku, Kyoto 603-8585, Japan; [email protected] (M.Y.); [email protected] (H.Y.) 3 Kansai Gakken Laboratory, Kankyo Eisei Yakuhin Co. Ltd., Kyoto 619-0237, Japan * Correspondence: [email protected] Received: 15 August 2020; Accepted: 1 September 2020; Published: 3 September 2020 Abstract: Developmental and epileptic encephalopathies (DEEs) are the spectrum of severe epilepsies characterized by early-onset, refractory seizures occurring in the context of developmental regression or plateauing. Early infantile epileptic encephalopathy (EIEE) is one of the earliest forms of DEE, manifesting as frequent epileptic spasms and characteristic electroencephalogram findings in early infancy. In recent years, next-generation sequencing approaches have identified a number of monogenic determinants underlying DEE. In the case of EIEE, 85 genes have been registered in Online Mendelian Inheritance in Man as causative genes. Model organisms are indispensable tools for understanding the in vivo roles of the newly identified causative genes. In this review, we first present an overview of epilepsy and its genetic etiology, especially focusing on EIEE and then briefly summarize epilepsy research using animal and patient-derived induced pluripotent stem cell (iPSC) models. The Drosophila model, which is characterized by easy gene manipulation, a short generation time, low cost and fewer ethical restrictions when designing experiments, is optimal for understanding the genetics of DEE. -

Table S1. 103 Ferroptosis-Related Genes Retrieved from the Genecards
Table S1. 103 ferroptosis-related genes retrieved from the GeneCards. Gene Symbol Description Category GPX4 Glutathione Peroxidase 4 Protein Coding AIFM2 Apoptosis Inducing Factor Mitochondria Associated 2 Protein Coding TP53 Tumor Protein P53 Protein Coding ACSL4 Acyl-CoA Synthetase Long Chain Family Member 4 Protein Coding SLC7A11 Solute Carrier Family 7 Member 11 Protein Coding VDAC2 Voltage Dependent Anion Channel 2 Protein Coding VDAC3 Voltage Dependent Anion Channel 3 Protein Coding ATG5 Autophagy Related 5 Protein Coding ATG7 Autophagy Related 7 Protein Coding NCOA4 Nuclear Receptor Coactivator 4 Protein Coding HMOX1 Heme Oxygenase 1 Protein Coding SLC3A2 Solute Carrier Family 3 Member 2 Protein Coding ALOX15 Arachidonate 15-Lipoxygenase Protein Coding BECN1 Beclin 1 Protein Coding PRKAA1 Protein Kinase AMP-Activated Catalytic Subunit Alpha 1 Protein Coding SAT1 Spermidine/Spermine N1-Acetyltransferase 1 Protein Coding NF2 Neurofibromin 2 Protein Coding YAP1 Yes1 Associated Transcriptional Regulator Protein Coding FTH1 Ferritin Heavy Chain 1 Protein Coding TF Transferrin Protein Coding TFRC Transferrin Receptor Protein Coding FTL Ferritin Light Chain Protein Coding CYBB Cytochrome B-245 Beta Chain Protein Coding GSS Glutathione Synthetase Protein Coding CP Ceruloplasmin Protein Coding PRNP Prion Protein Protein Coding SLC11A2 Solute Carrier Family 11 Member 2 Protein Coding SLC40A1 Solute Carrier Family 40 Member 1 Protein Coding STEAP3 STEAP3 Metalloreductase Protein Coding ACSL1 Acyl-CoA Synthetase Long Chain Family Member 1 Protein