Novel Pedigree Analysis Implicates DNA Repair and Chromatin
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Hypopituitarism May Be an Additional Feature of SIM1 and POU3F2 Containing 6Q16 Deletions in Children with Early Onset Obesity
Open Access Journal of Pediatric Endocrinology Case Report Hypopituitarism may be an Additional Feature of SIM1 and POU3F2 Containing 6q16 Deletions in Children with Early Onset Obesity Rutteman B1, De Rademaeker M2, Gies I1, Van den Bogaert A2, Zeevaert R1, Vanbesien J1 and De Abstract Schepper J1* Over the past decades, 6q16 deletions have become recognized as a 1Department of Pediatric Endocrinology, UZ Brussel, frequent cause of the Prader-Willi-like syndrome. Involvement of SIM1 in the Belgium deletion has been linked to the development of obesity. Although SIM1 together 2Department of Genetics, UZ Brussel, Belgium with POU3F2, which is located close to the SIM1 locus, are involved in pituitary *Corresponding author: De Schepper J, Department development and function, pituitary dysfunction has not been reported frequently of Pediatric Endocrinology, UZ Brussel, Belgium in cases of 6q16.1q16.3 deletion involving both genes. Here we report on a case of a girl with typical Prader-Willi-like symptoms including early-onset Received: December 23, 2016; Accepted: February 21, hyperphagic obesity, hypotonia, short hands and feet and neuro-psychomotor 2017; Published: February 22, 2017 development delay. Furthermore, she suffered from central diabetes insipidus, central hypothyroidism and hypocortisolism. Her genetic defect is a 6q16.1q16.3 deletion, including SIM1 and POU3F2. We recommend searching for a 6q16 deletion in children with early onset hyperphagic obesity associated with biological signs of hypopituitarism and/or polyuria-polydipsia syndrome. The finding of a combined SIM1 and POU3F2 deletion should prompt monitoring for the development of hypopituitarism, if not already present at diagnosis. Keywords: 6q16 deletion; SIM1; POU3F2; Prader-Willi-like syndrome; Hypopituitarism Introduction insipidus and of some specific pituitary hormone deficiencies in children with a combined SIM1 and POU3F2 deficiency. -

Appendix 2. Significantly Differentially Regulated Genes in Term Compared with Second Trimester Amniotic Fluid Supernatant
Appendix 2. Significantly Differentially Regulated Genes in Term Compared With Second Trimester Amniotic Fluid Supernatant Fold Change in term vs second trimester Amniotic Affymetrix Duplicate Fluid Probe ID probes Symbol Entrez Gene Name 1019.9 217059_at D MUC7 mucin 7, secreted 424.5 211735_x_at D SFTPC surfactant protein C 416.2 206835_at STATH statherin 363.4 214387_x_at D SFTPC surfactant protein C 295.5 205982_x_at D SFTPC surfactant protein C 288.7 1553454_at RPTN repetin solute carrier family 34 (sodium 251.3 204124_at SLC34A2 phosphate), member 2 238.9 206786_at HTN3 histatin 3 161.5 220191_at GKN1 gastrokine 1 152.7 223678_s_at D SFTPA2 surfactant protein A2 130.9 207430_s_at D MSMB microseminoprotein, beta- 99.0 214199_at SFTPD surfactant protein D major histocompatibility complex, class II, 96.5 210982_s_at D HLA-DRA DR alpha 96.5 221133_s_at D CLDN18 claudin 18 94.4 238222_at GKN2 gastrokine 2 93.7 1557961_s_at D LOC100127983 uncharacterized LOC100127983 93.1 229584_at LRRK2 leucine-rich repeat kinase 2 HOXD cluster antisense RNA 1 (non- 88.6 242042_s_at D HOXD-AS1 protein coding) 86.0 205569_at LAMP3 lysosomal-associated membrane protein 3 85.4 232698_at BPIFB2 BPI fold containing family B, member 2 84.4 205979_at SCGB2A1 secretoglobin, family 2A, member 1 84.3 230469_at RTKN2 rhotekin 2 82.2 204130_at HSD11B2 hydroxysteroid (11-beta) dehydrogenase 2 81.9 222242_s_at KLK5 kallikrein-related peptidase 5 77.0 237281_at AKAP14 A kinase (PRKA) anchor protein 14 76.7 1553602_at MUCL1 mucin-like 1 76.3 216359_at D MUC7 mucin 7, -

Karla Alejandra Vizcarra Zevallos Análise Da Função De Genes
Karla Alejandra Vizcarra Zevallos Análise da função de genes candidatos à manutenção da inativação do cromossomo X em humanos Dissertação apresentada ao Pro- grama de Pós‐Graduação Inter- unidades em Biotecnologia USP/ Instituto Butantan/ IPT, para obtenção do Título de Mestre em Ciências. São Paulo 2017 Karla Alejandra Vizcarra Zevallos Análise da função de genes candidatos à manutenção da inativação do cromossomo X em humanos Dissertação apresentada ao Pro- grama de Pós‐Graduação Inter- unidades em Biotecnologia do Instituto de Ciências Biomédicas USP/ Instituto Butantan/ IPT, para obtenção do Título de Mestre em Ciências. Área de concentração: Biotecnologia Orientadora: Profa. Dra. Lygia da Veiga Pereira Carramaschi Versão corrigida. A versão original eletrônica encontra-se disponível tanto na Biblioteca do ICB quanto na Biblioteca Digital de Teses e Dissertações da USP (BDTD) São Paulo 2017 UNIVERSIDADE DE SÃO PAULO Programa de Pós-Graduação Interunidades em Biotecnologia Universidade de São Paulo, Instituto Butantan, Instituto de Pesquisas Tecnológicas Candidato(a): Karla Alejandra Vizcarra Zevallos Título da Dissertação: Análise da função de genes candidatos à manutenção da inativação do cromossomo X em humanos Orientador: Profa. Dra. Lygia da Veiga Pereira Carramaschi A Comissão Julgadora dos trabalhos de Defesa da Dissertação de Mestrado, em sessão pública realizada a ........./......../.........., considerou o(a) candidato(a): ( ) Aprovado(a) ( ) Reprovado(a) Examinador(a): Assinatura: .............................................................................. -

Improving the Detection of Genomic Rearrangements in Short Read Sequencing Data
Improving the detection of genomic rearrangements in short read sequencing data Daniel Lee Cameron orcid.org/0000-0002-0951-7116 Doctor of Philosophy July 2017 Department of Medical Biology, University of Melbourne Bioinformatics Division, Walter and Eliza Hall Institute of Medical Research Submitted in total fulfilment of the requirements of the degree of Doctor of Philosophy Produced on archival quality paper Page i Abstract Genomic rearrangements, also known as structural variants, play a significant role in the development of cancer and in genetic disorders. With the bulk of genetic studies focusing on single nucleotide variants and small insertions and deletions, structural variants are often overlooked. In part this is due to the increased complexity of analysis required, but is also due to the increased difficulty of detection. The identification of genomic rearrangements using massively parallel sequencing remains a major challenge. To address this, I have developed the Genome Rearrangement Identification Software Suite (GRIDSS). In this thesis, I show that high sensitivity and specificity can be achieved by performing reference-guided assembly prior to variant calling and incorporating the results of this assembly into the variant calling process itself. Utilising a novel genome-wide break-end assembly approach, GRIDSS halves the false discovery rate compared to other recent methods on human cell line data. A comparison of assembly approaches reveals that incorporating read alignment information into a positional de Bruijn graph improves assembly quality compared to traditional de Bruijn graph assembly. To characterise the performance of structural variant calling software, I have performed the most comprehensive benchmarking of structural variant calling software to date. -

Metabolites OH
H OH metabolites OH Article Metabolite Genome-Wide Association Study (mGWAS) and Gene-Metabolite Interaction Network Analysis Reveal Potential Biomarkers for Feed Efficiency in Pigs Xiao Wang and Haja N. Kadarmideen * Quantitative Genomics, Bioinformatics and Computational Biology Group, Department of Applied Mathematics and Computer Science, Technical University of Denmark, Richard Petersens Plads, Building 324, 2800 Kongens Lyngby, Denmark; [email protected] * Correspondence: [email protected] Received: 1 April 2020; Accepted: 11 May 2020; Published: 15 May 2020 Abstract: Metabolites represent the ultimate response of biological systems, so metabolomics is considered the link between genotypes and phenotypes. Feed efficiency is one of the most important phenotypes in sustainable pig production and is the main breeding goal trait. We utilized metabolic and genomic datasets from a total of 108 pigs from our own previously published studies that involved 59 Duroc and 49 Landrace pigs with data on feed efficiency (residual feed intake (RFI)), genotype (PorcineSNP80 BeadChip) data, and metabolomic data (45 final metabolite datasets derived from LC-MS system). Utilizing these datasets, our main aim was to identify genetic variants (single-nucleotide polymorphisms (SNPs)) that affect 45 different metabolite concentrations in plasma collected at the start and end of the performance testing of pigs categorized as high or low in their feed efficiency (based on RFI values). Genome-wide significant genetic variants could be then used as potential genetic or biomarkers in breeding programs for feed efficiency. The other objective was to reveal the biochemical mechanisms underlying genetic variation for pigs’ feed efficiency. In order to achieve these objectives, we firstly conducted a metabolite genome-wide association study (mGWAS) based on mixed linear models and found 152 genome-wide significant SNPs (p-value < 1.06 10 6) in association with 17 metabolites that included 90 significant SNPs annotated × − to 52 genes. -
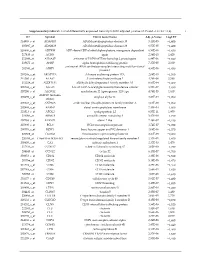
Supplementary Table S1. List of Differentially Expressed
Supplementary table S1. List of differentially expressed transcripts (FDR adjusted p‐value < 0.05 and −1.4 ≤ FC ≥1.4). 1 ID Symbol Entrez Gene Name Adj. p‐Value Log2 FC 214895_s_at ADAM10 ADAM metallopeptidase domain 10 3,11E‐05 −1,400 205997_at ADAM28 ADAM metallopeptidase domain 28 6,57E‐05 −1,400 220606_s_at ADPRM ADP‐ribose/CDP‐alcohol diphosphatase, manganese dependent 6,50E‐06 −1,430 217410_at AGRN agrin 2,34E‐10 1,420 212980_at AHSA2P activator of HSP90 ATPase homolog 2, pseudogene 6,44E‐06 −1,920 219672_at AHSP alpha hemoglobin stabilizing protein 7,27E‐05 2,330 aminoacyl tRNA synthetase complex interacting multifunctional 202541_at AIMP1 4,91E‐06 −1,830 protein 1 210269_s_at AKAP17A A‐kinase anchoring protein 17A 2,64E‐10 −1,560 211560_s_at ALAS2 5ʹ‐aminolevulinate synthase 2 4,28E‐06 3,560 212224_at ALDH1A1 aldehyde dehydrogenase 1 family member A1 8,93E‐04 −1,400 205583_s_at ALG13 ALG13 UDP‐N‐acetylglucosaminyltransferase subunit 9,50E‐07 −1,430 207206_s_at ALOX12 arachidonate 12‐lipoxygenase, 12S type 4,76E‐05 1,630 AMY1C (includes 208498_s_at amylase alpha 1C 3,83E‐05 −1,700 others) 201043_s_at ANP32A acidic nuclear phosphoprotein 32 family member A 5,61E‐09 −1,760 202888_s_at ANPEP alanyl aminopeptidase, membrane 7,40E‐04 −1,600 221013_s_at APOL2 apolipoprotein L2 6,57E‐11 1,600 219094_at ARMC8 armadillo repeat containing 8 3,47E‐08 −1,710 207798_s_at ATXN2L ataxin 2 like 2,16E‐07 −1,410 215990_s_at BCL6 BCL6 transcription repressor 1,74E‐07 −1,700 200776_s_at BZW1 basic leucine zipper and W2 domains 1 1,09E‐06 −1,570 222309_at -

Bedeutung Und Charakterisierung Der 6Q-Deletion Beim Prostatakarzinom
Bedeutung und Charakterisierung der 6q-Deletion beim Prostatakarzinom Dissertation zur Erlangung des Doktorgrades der Naturwissenschaften (Dr. rer. nat.) am Department Biologie der Fakultät Mathematik, Informatik und Naturwissenschaften der Universität Hamburg Vorgelegt von Martina Kluth Heide-Holstein Hamburg 2015 Gutachter der Dissertation Prof. Dr. Guido Sauter Prof. Dr. Thomas Dobner Datum der Disputation: 22.01.2016 Inhaltsverzeichnis 1Einleitung 1 1.1 Epidemiologie und Ätiologie des Prostatakarzinoms .............. 1 1.2 Früherkennung und Primärdiagnostik des Prostatakarzinoms .............................. 2 1.2.1 Digitale rektale Untersuchung (DRU) .................. 3 1.2.2 Prostataspezifisches Antigen (PSA) ................... 3 1.2.3 Stanzbiopsie der Prostata ........................ 5 1.3 Pathologie und Stadieneinteilung des Prostatakarzinoms ................................ 5 1.4 Therapie des Prostatakarzinoms ......................... 7 1.5 Prognosemarker beim Prostatakarzinom .................... 11 1.6 Genomische Veränderungen beim Prostatakarzinom .............. 12 1.6.1 Punktmutation .............................. 12 1.6.2 Chromosomale Zugewinne ........................ 12 1.6.3 Translokationen .............................. 13 1.6.4 Deletionen ................................. 14 1.7 Die 6q-Deletion beim Prostatakarzinom ..................... 15 1.7.1 Prävalenz der 6q-Deletion beim Prostatakarzinom ........... 15 1.7.2 Assoziation der 6q-Deletion mit dem Tumorphänotyp und der Progno- se des Prostatakarzinoms ........................ -

Agricultural University of Athens
ΓΕΩΠΟΝΙΚΟ ΠΑΝΕΠΙΣΤΗΜΙΟ ΑΘΗΝΩΝ ΣΧΟΛΗ ΕΠΙΣΤΗΜΩΝ ΤΩΝ ΖΩΩΝ ΤΜΗΜΑ ΕΠΙΣΤΗΜΗΣ ΖΩΙΚΗΣ ΠΑΡΑΓΩΓΗΣ ΕΡΓΑΣΤΗΡΙΟ ΓΕΝΙΚΗΣ ΚΑΙ ΕΙΔΙΚΗΣ ΖΩΟΤΕΧΝΙΑΣ ΔΙΔΑΚΤΟΡΙΚΗ ΔΙΑΤΡΙΒΗ Εντοπισμός γονιδιωματικών περιοχών και δικτύων γονιδίων που επηρεάζουν παραγωγικές και αναπαραγωγικές ιδιότητες σε πληθυσμούς κρεοπαραγωγικών ορνιθίων ΕΙΡΗΝΗ Κ. ΤΑΡΣΑΝΗ ΕΠΙΒΛΕΠΩΝ ΚΑΘΗΓΗΤΗΣ: ΑΝΤΩΝΙΟΣ ΚΟΜΙΝΑΚΗΣ ΑΘΗΝΑ 2020 ΔΙΔΑΚΤΟΡΙΚΗ ΔΙΑΤΡΙΒΗ Εντοπισμός γονιδιωματικών περιοχών και δικτύων γονιδίων που επηρεάζουν παραγωγικές και αναπαραγωγικές ιδιότητες σε πληθυσμούς κρεοπαραγωγικών ορνιθίων Genome-wide association analysis and gene network analysis for (re)production traits in commercial broilers ΕΙΡΗΝΗ Κ. ΤΑΡΣΑΝΗ ΕΠΙΒΛΕΠΩΝ ΚΑΘΗΓΗΤΗΣ: ΑΝΤΩΝΙΟΣ ΚΟΜΙΝΑΚΗΣ Τριμελής Επιτροπή: Aντώνιος Κομινάκης (Αν. Καθ. ΓΠΑ) Ανδρέας Κράνης (Eρευν. B, Παν. Εδιμβούργου) Αριάδνη Χάγερ (Επ. Καθ. ΓΠΑ) Επταμελής εξεταστική επιτροπή: Aντώνιος Κομινάκης (Αν. Καθ. ΓΠΑ) Ανδρέας Κράνης (Eρευν. B, Παν. Εδιμβούργου) Αριάδνη Χάγερ (Επ. Καθ. ΓΠΑ) Πηνελόπη Μπεμπέλη (Καθ. ΓΠΑ) Δημήτριος Βλαχάκης (Επ. Καθ. ΓΠΑ) Ευάγγελος Ζωίδης (Επ.Καθ. ΓΠΑ) Γεώργιος Θεοδώρου (Επ.Καθ. ΓΠΑ) 2 Εντοπισμός γονιδιωματικών περιοχών και δικτύων γονιδίων που επηρεάζουν παραγωγικές και αναπαραγωγικές ιδιότητες σε πληθυσμούς κρεοπαραγωγικών ορνιθίων Περίληψη Σκοπός της παρούσας διδακτορικής διατριβής ήταν ο εντοπισμός γενετικών δεικτών και υποψηφίων γονιδίων που εμπλέκονται στο γενετικό έλεγχο δύο τυπικών πολυγονιδιακών ιδιοτήτων σε κρεοπαραγωγικά ορνίθια. Μία ιδιότητα σχετίζεται με την ανάπτυξη (σωματικό βάρος στις 35 ημέρες, ΣΒ) και η άλλη με την αναπαραγωγική -

The Changing Chromatome As a Driver of Disease: a Panoramic View from Different Methodologies
The changing chromatome as a driver of disease: A panoramic view from different methodologies Isabel Espejo1, Luciano Di Croce,1,2,3 and Sergi Aranda1 1. Centre for Genomic Regulation (CRG), Barcelona Institute of Science and Technology, Dr. Aiguader 88, Barcelona 08003, Spain 2. Universitat Pompeu Fabra (UPF), Barcelona, Spain 3. ICREA, Pg. Lluis Companys 23, Barcelona 08010, Spain *Corresponding authors: Luciano Di Croce ([email protected]) Sergi Aranda ([email protected]) 1 GRAPHICAL ABSTRACT Chromatin-bound proteins regulate gene expression, replicate and repair DNA, and transmit epigenetic information. Several human diseases are highly influenced by alterations in the chromatin- bound proteome. Thus, biochemical approaches for the systematic characterization of the chromatome could contribute to identifying new regulators of cellular functionality, including those that are relevant to human disorders. 2 SUMMARY Chromatin-bound proteins underlie several fundamental cellular functions, such as control of gene expression and the faithful transmission of genetic and epigenetic information. Components of the chromatin proteome (the “chromatome”) are essential in human life, and mutations in chromatin-bound proteins are frequently drivers of human diseases, such as cancer. Proteomic characterization of chromatin and de novo identification of chromatin interactors could thus reveal important and perhaps unexpected players implicated in human physiology and disease. Recently, intensive research efforts have focused on developing strategies to characterize the chromatome composition. In this review, we provide an overview of the dynamic composition of the chromatome, highlight the importance of its alterations as a driving force in human disease (and particularly in cancer), and discuss the different approaches to systematically characterize the chromatin-bound proteome in a global manner. -

Content Based Search in Gene Expression Databases and a Meta-Analysis of Host Responses to Infection
Content Based Search in Gene Expression Databases and a Meta-analysis of Host Responses to Infection A Thesis Submitted to the Faculty of Drexel University by Francis X. Bell in partial fulfillment of the requirements for the degree of Doctor of Philosophy November 2015 c Copyright 2015 Francis X. Bell. All Rights Reserved. ii Acknowledgments I would like to acknowledge and thank my advisor, Dr. Ahmet Sacan. Without his advice, support, and patience I would not have been able to accomplish all that I have. I would also like to thank my committee members and the Biomed Faculty that have guided me. I would like to give a special thanks for the members of the bioinformatics lab, in particular the members of the Sacan lab: Rehman Qureshi, Daisy Heng Yang, April Chunyu Zhao, and Yiqian Zhou. Thank you for creating a pleasant and friendly environment in the lab. I give the members of my family my sincerest gratitude for all that they have done for me. I cannot begin to repay my parents for their sacrifices. I am eternally grateful for everything they have done. The support of my sisters and their encouragement gave me the strength to persevere to the end. iii Table of Contents LIST OF TABLES.......................................................................... vii LIST OF FIGURES ........................................................................ xiv ABSTRACT ................................................................................ xvii 1. A BRIEF INTRODUCTION TO GENE EXPRESSION............................. 1 1.1 Central Dogma of Molecular Biology........................................... 1 1.1.1 Basic Transfers .......................................................... 1 1.1.2 Uncommon Transfers ................................................... 3 1.2 Gene Expression ................................................................. 4 1.2.1 Estimating Gene Expression ............................................ 4 1.2.2 DNA Microarrays ...................................................... -

A Meta-Analysis of the Effects of High-LET Ionizing Radiations in Human Gene Expression
Supplementary Materials A Meta-Analysis of the Effects of High-LET Ionizing Radiations in Human Gene Expression Table S1. Statistically significant DEGs (Adj. p-value < 0.01) derived from meta-analysis for samples irradiated with high doses of HZE particles, collected 6-24 h post-IR not common with any other meta- analysis group. This meta-analysis group consists of 3 DEG lists obtained from DGEA, using a total of 11 control and 11 irradiated samples [Data Series: E-MTAB-5761 and E-MTAB-5754]. Ensembl ID Gene Symbol Gene Description Up-Regulated Genes ↑ (2425) ENSG00000000938 FGR FGR proto-oncogene, Src family tyrosine kinase ENSG00000001036 FUCA2 alpha-L-fucosidase 2 ENSG00000001084 GCLC glutamate-cysteine ligase catalytic subunit ENSG00000001631 KRIT1 KRIT1 ankyrin repeat containing ENSG00000002079 MYH16 myosin heavy chain 16 pseudogene ENSG00000002587 HS3ST1 heparan sulfate-glucosamine 3-sulfotransferase 1 ENSG00000003056 M6PR mannose-6-phosphate receptor, cation dependent ENSG00000004059 ARF5 ADP ribosylation factor 5 ENSG00000004777 ARHGAP33 Rho GTPase activating protein 33 ENSG00000004799 PDK4 pyruvate dehydrogenase kinase 4 ENSG00000004848 ARX aristaless related homeobox ENSG00000005022 SLC25A5 solute carrier family 25 member 5 ENSG00000005108 THSD7A thrombospondin type 1 domain containing 7A ENSG00000005194 CIAPIN1 cytokine induced apoptosis inhibitor 1 ENSG00000005381 MPO myeloperoxidase ENSG00000005486 RHBDD2 rhomboid domain containing 2 ENSG00000005884 ITGA3 integrin subunit alpha 3 ENSG00000006016 CRLF1 cytokine receptor like -

Co-Expression Networks Reveal the Tissue-Specific Regulation of Transcription and Splicing
Downloaded from genome.cshlp.org on September 26, 2021 - Published by Cold Spring Harbor Laboratory Press Method Co-expression networks reveal the tissue-specific regulation of transcription and splicing Ashis Saha,1 Yungil Kim,1,6 Ariel D.H. Gewirtz,2,6 Brian Jo,2 Chuan Gao,3 Ian C. McDowell,4 The GTEx Consortium,7 Barbara E. Engelhardt,5 and Alexis Battle1 1Department of Computer Science, Johns Hopkins University, Baltimore, Maryland 21218, USA; 2Program in Quantitative and Computational Biology, Princeton University, Princeton, New Jersey 08540, USA; 3Department of Statistical Science, Duke University, Durham, North Carolina 27708, USA; 4Program in Computational Biology and Bioinformatics, Duke University, Durham, North Carolina 27708, USA; 5Department of Computer Science and Center for Statistics and Machine Learning, Princeton University, Princeton, New Jersey 08540, USA Gene co-expression networks capture biologically important patterns in gene expression data, enabling functional analyses of genes, discovery of biomarkers, and interpretation of genetic variants. Most network analyses to date have been limited to assessing correlation between total gene expression levels in a single tissue or small sets of tissues. Here, we built networks that additionally capture the regulation of relative isoform abundance and splicing, along with tissue-specific connections unique to each of a diverse set of tissues. We used the Genotype-Tissue Expression (GTEx) project v6 RNA sequencing data across 50 tissues and 449 individuals. First, we developed a framework called Transcriptome-Wide Networks (TWNs) for combining total expression and relative isoform levels into a single sparse network, capturing the interplay between the reg- ulation of splicing and transcription.