The Cancer Genome Atlas Dataset-Based Analysis of Aberrantly Expressed Genes by Geneanalytics in Thymoma Associated Myasthenia Gravis: Focusing on T Cells
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Prioritization and Evaluation of Depression Candidate Genes by Combining Multidimensional Data Resources
Prioritization and Evaluation of Depression Candidate Genes by Combining Multidimensional Data Resources Chung-Feng Kao1, Yu-Sheng Fang2, Zhongming Zhao3, Po-Hsiu Kuo1,2,4* 1 Department of Public Health and Institute of Epidemiology and Preventive Medicine, College of Public Health, National Taiwan University, Taipei, Taiwan, 2 Institute of Clinical Medicine, School of Medicine, National Cheng-Kung University, Tainan, Taiwan, 3 Departments of Biomedical Informatics and Psychiatry, Vanderbilt University School of Medicine, Nashville, Tennessee, United States of America, 4 Research Center for Genes, Environment and Human Health, National Taiwan University, Taipei, Taiwan Abstract Background: Large scale and individual genetic studies have suggested numerous susceptible genes for depression in the past decade without conclusive results. There is a strong need to review and integrate multi-dimensional data for follow up validation. The present study aimed to apply prioritization procedures to build-up an evidence-based candidate genes dataset for depression. Methods: Depression candidate genes were collected in human and animal studies across various data resources. Each gene was scored according to its magnitude of evidence related to depression and was multiplied by a source-specific weight to form a combined score measure. All genes were evaluated through a prioritization system to obtain an optimal weight matrix to rank their relative importance with depression using the combined scores. The resulting candidate gene list for depression (DEPgenes) was further evaluated by a genome-wide association (GWA) dataset and microarray gene expression in human tissues. Results: A total of 5,055 candidate genes (4,850 genes from human and 387 genes from animal studies with 182 being overlapped) were included from seven data sources. -

Whole-Genome Microarray Detects Deletions and Loss of Heterozygosity of Chromosome 3 Occurring Exclusively in Metastasizing Uveal Melanoma
Anatomy and Pathology Whole-Genome Microarray Detects Deletions and Loss of Heterozygosity of Chromosome 3 Occurring Exclusively in Metastasizing Uveal Melanoma Sarah L. Lake,1 Sarah E. Coupland,1 Azzam F. G. Taktak,2 and Bertil E. Damato3 PURPOSE. To detect deletions and loss of heterozygosity of disease is fatal in 92% of patients within 2 years of diagnosis. chromosome 3 in a rare subset of fatal, disomy 3 uveal mela- Clinical and histopathologic risk factors for UM metastasis noma (UM), undetectable by fluorescence in situ hybridization include large basal tumor diameter (LBD), ciliary body involve- (FISH). ment, epithelioid cytomorphology, extracellular matrix peri- ϩ ETHODS odic acid-Schiff-positive (PAS ) loops, and high mitotic M . Multiplex ligation-dependent probe amplification 3,4 5 (MLPA) with the P027 UM assay was performed on formalin- count. Prescher et al. showed that a nonrandom genetic fixed, paraffin-embedded (FFPE) whole tumor sections from 19 change, monosomy 3, correlates strongly with metastatic death, and the correlation has since been confirmed by several disomy 3 metastasizing UMs. Whole-genome microarray analy- 3,6–10 ses using a single-nucleotide polymorphism microarray (aSNP) groups. Consequently, fluorescence in situ hybridization were performed on frozen tissue samples from four fatal dis- (FISH) detection of chromosome 3 using a centromeric probe omy 3 metastasizing UMs and three disomy 3 tumors with Ͼ5 became routine practice for UM prognostication; however, 5% years’ metastasis-free survival. to 20% of disomy 3 UM patients unexpectedly develop metas- tases.11 Attempts have therefore been made to identify the RESULTS. Two metastasizing UMs that had been classified as minimal region(s) of deletion on chromosome 3.12–15 Despite disomy 3 by FISH analysis of a small tumor sample were found these studies, little progress has been made in defining the key on MLPA analysis to show monosomy 3. -

Transcriptional Regulation of RKIP in Prostate Cancer Progression
Health Science Campus FINAL APPROVAL OF DISSERTATION Doctor of Philosophy in Biomedical Sciences Transcriptional Regulation of RKIP in Prostate Cancer Progression Submitted by: Sandra Marie Beach In partial fulfillment of the requirements for the degree of Doctor of Philosophy in Biomedical Sciences Examination Committee Major Advisor: Kam Yeung, Ph.D. Academic William Maltese, Ph.D. Advisory Committee: Sonia Najjar, Ph.D. Han-Fei Ding, M.D., Ph.D. Manohar Ratnam, Ph.D. Senior Associate Dean College of Graduate Studies Michael S. Bisesi, Ph.D. Date of Defense: May 16, 2007 Transcriptional Regulation of RKIP in Prostate Cancer Progression Sandra Beach University of Toledo ACKNOWLDEGMENTS I thank my major advisor, Dr. Kam Yeung, for the opportunity to pursue my degree in his laboratory. I am also indebted to my advisory committee members past and present, Drs. Sonia Najjar, Han-Fei Ding, Manohar Ratnam, James Trempe, and Douglas Pittman for generously and judiciously guiding my studies and sharing reagents and equipment. I owe extended thanks to Dr. William Maltese as a committee member and chairman of my department for supporting my degree progress. The entire Department of Biochemistry and Cancer Biology has been most kind and helpful to me. Drs. Roy Collaco and Hong-Juan Cui have shared their excellent technical and practical advice with me throughout my studies. I thank members of the Yeung laboratory, Dr. Sungdae Park, Hui Hui Tang, Miranda Yeung for their support and collegiality. The data mining studies herein would not have been possible without the helpful advice of Dr. Robert Trumbly. I am also grateful for the exceptional assistance and shared microarray data of Dr. -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

ABCC12 Monoclonal Antibody, Clone M9II-3
ABCC12 monoclonal antibody, Gene Symbol: ABCC12 clone M9II-3 Gene Alias: MGC27071, MRP9 Catalog Number: MAB6675 Gene Summary: This gene is a member of the superfamily of ATP-binding cassette (ABC) transporters Regulatory Status: For research use only (RUO) and the encoded protein contains two ATP-binding domains and 12 transmembrane regions. ABC proteins Product Description: Rat monoclonal antibody raised transport various molecules across extra- and against partial recombinant ABCC12. intracellular membranes. ABC genes are divided into Clone Name: M9II-3 seven distinct subfamilies: ABC1, MDR/TAP, MRP, ALD, OABP, GCN20, and White. This gene is a member of Immunogen: Recombinant protein corresponding to the MRP subfamily which is involved in multi-drug amino acids 690-734 of human ABCC12. resistance. This gene and another subfamily member are arranged head-to-tail on chromosome 16q12.1. Host: Rat Increased expression of this gene is associated with breast cancer. [provided by RefSeq] Reactivity: Human References: Applications: ICC, IHC-Fr, WB 1. Multidrug resistance-associated protein 9 (ABCC12) is (See our web site product page for detailed applications present in mouse and boar sperm. Ono N, Van der information) Heijden I, Scheffer GL, Van de Wetering K, Van Deemter E, De Haas M, Boerke A, Gadella BM, De Rooij Protocols: See our web site at DG, Neefjes JJ, Groothuis TA, Oomen L, Brocks L, http://www.abnova.com/support/protocols.asp or product Ishikawa T, Borst P. Ono N, Van der Heijden I, Scheffer page for detailed protocols GL, Van de Wetering K, Van Deemter E, De Haas M, Boerke A, Gadella BM, De Rooij DG, Neefjes JJ, Specificity: M9II-3 reacts with an internal epitope of Groothuis TA, Oomen L, Brocks L, Ishikawa T, Borst P. -

Primate Specific Retrotransposons, Svas, in the Evolution of Networks That Alter Brain Function
Title: Primate specific retrotransposons, SVAs, in the evolution of networks that alter brain function. Olga Vasieva1*, Sultan Cetiner1, Abigail Savage2, Gerald G. Schumann3, Vivien J Bubb2, John P Quinn2*, 1 Institute of Integrative Biology, University of Liverpool, Liverpool, L69 7ZB, U.K 2 Department of Molecular and Clinical Pharmacology, Institute of Translational Medicine, The University of Liverpool, Liverpool L69 3BX, UK 3 Division of Medical Biotechnology, Paul-Ehrlich-Institut, Langen, D-63225 Germany *. Corresponding author Olga Vasieva: Institute of Integrative Biology, Department of Comparative genomics, University of Liverpool, Liverpool, L69 7ZB, [email protected] ; Tel: (+44) 151 795 4456; FAX:(+44) 151 795 4406 John Quinn: Department of Molecular and Clinical Pharmacology, Institute of Translational Medicine, The University of Liverpool, Liverpool L69 3BX, UK, [email protected]; Tel: (+44) 151 794 5498. Key words: SVA, trans-mobilisation, behaviour, brain, evolution, psychiatric disorders 1 Abstract The hominid-specific non-LTR retrotransposon termed SINE–VNTR–Alu (SVA) is the youngest of the transposable elements in the human genome. The propagation of the most ancient SVA type A took place about 13.5 Myrs ago, and the youngest SVA types appeared in the human genome after the chimpanzee divergence. Functional enrichment analysis of genes associated with SVA insertions demonstrated their strong link to multiple ontological categories attributed to brain function and the disorders. SVA types that expanded their presence in the human genome at different stages of hominoid life history were also associated with progressively evolving behavioural features that indicated a potential impact of SVA propagation on a cognitive ability of a modern human. -
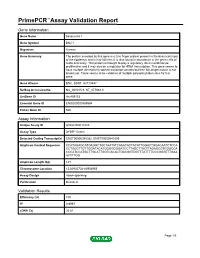
Primepcr™Assay Validation Report
PrimePCR™Assay Validation Report Gene Information Gene Name basonuclin 1 Gene Symbol BNC1 Organism Human Gene Summary The protein encoded by this gene is a zinc finger protein present in the basal cell layer of the epidermis and in hair follicles. It is also found in abundance in the germ cells of testis and ovary. This protein is thought to play a regulatory role in keratinocyte proliferation and it may also be a regulator for rRNA transcription. This gene seems to have multiple alternatively spliced transcript variants but their full-length nature is not known yet. There seems to be evidence of multiple polyadenylation sites for this gene. Gene Aliases BNC, BSN1, HsT19447 RefSeq Accession No. NC_000015.9, NT_077661.3 UniGene ID Hs.459153 Ensembl Gene ID ENSG00000169594 Entrez Gene ID 646 Assay Information Unique Assay ID qHsaCID0017223 Assay Type SYBR® Green Detected Coding Transcript(s) ENST00000345382, ENST00000541809 Amplicon Context Sequence CCATAGAGCATGAGGCTGCTAATATCAAACACTACATTGGACTGGACAATCTCCA CCTGGCTTGTTGGATACATGGGGGGGATCCTTAGCTTACTTAGAGCGTGGGCCA CCCATCCATGCTTGCATTGGTCACACTGACGGTGGTTTATTTTCCCGGGTTTGAA ACTTTGG Amplicon Length (bp) 141 Chromosome Location 15:83935724-83936955 Assay Design Intron-spanning Purification Desalted Validation Results Efficiency (%) 100 R2 0.9997 cDNA Cq 26.81 Page 1/5 PrimePCR™Assay Validation Report cDNA Tm (Celsius) 84.5 gDNA Cq 39.42 Specificity (%) 100 Information to assist with data interpretation is provided at the end of this report. Page 2/5 PrimePCR™Assay Validation Report BNC1, Human Amplification -

DF6216-CSRP1 Antibody
Affinity Biosciences website:www.affbiotech.com order:[email protected] CSRP1 Antibody Cat.#: DF6216 Concn.: 1mg/ml Mol.Wt.: 21kDa Size: 50ul,100ul,200ul Source: Rabbit Clonality: Polyclonal Application: WB 1:500-1:2000, IHC 1:50-1:200, ELISA(peptide) 1:20000-1:40000 *The optimal dilutions should be determined by the end user. Reactivity: Human,Mouse,Rat Purification: The antiserum was purified by peptide affinity chromatography using SulfoLink™ Coupling Resin (Thermo Fisher Scientific). Specificity: CSRP1 Antibody detects endogenous levels of total CSRP1. Immunogen: A synthesized peptide derived from human CSRP1, corresponding to a region within the internal amino acids. Uniprot: P21291 Description: This gene encodes a member of the cysteine-rich protein (CSRP) family. This gene family includes a group of LIM domain proteins, which may be involved in regulatory processes important for development and cellular differentiation. The LIM/double zinc-finger motif found in this gene product occurs in proteins with critical functions in gene regulation, cell growth, and somatic differentiation. Alternatively spliced transcript variants have been described. [provided by RefSeq, Aug 2010] Storage Condition and Rabbit IgG in phosphate buffered saline , pH 7.4, 150mM Buffer: NaCl, 0.02% sodium azide and 50% glycerol.Store at -20 °C.Stable for 12 months from date of receipt. Western blot analysis of CSRP1 expression in Mouse lung lysate 1 / 2 Affinity Biosciences website:www.affbiotech.com order:[email protected] DF6216 at 1/100 staining Mouse brain tissue by IHC-P. The sample was formaldehyde fixed and a heat mediated antigen retrieval step in citrate buffer was performed. -

A Single-Cell Transcriptomic Landscape of Primate Arterial Aging
ARTICLE https://doi.org/10.1038/s41467-020-15997-0 OPEN A single-cell transcriptomic landscape of primate arterial aging Weiqi Zhang 1,2,3,4,5,13, Shu Zhang6,7,13, Pengze Yan3,8,13, Jie Ren7,9,13, Moshi Song3,5,8, Jingyi Li2,3,8, Jinghui Lei4, Huize Pan2,3, Si Wang3,5,8, Xibo Ma3,10, Shuai Ma2,3,8, Hongyu Li2,3, Fei Sun2,3, Haifeng Wan3,5,11, ✉ ✉ ✉ Wei Li 3,5,11, Piu Chan4, Qi Zhou3,5,11, Guang-Hui Liu 2,3,4,5,8 , Fuchou Tang 6,7,9,12 & Jing Qu 3,5,11 Our understanding of how aging affects the cellular and molecular components of the vas- 1234567890():,; culature and contributes to cardiovascular diseases is still limited. Here we report a single-cell transcriptomic survey of aortas and coronary arteries in young and old cynomolgus monkeys. Our data define the molecular signatures of specialized arteries and identify eight markers discriminating aortic and coronary vasculatures. Gene network analyses characterize tran- scriptional landmarks that regulate vascular senility and position FOXO3A, a longevity- associated transcription factor, as a master regulator gene that is downregulated in six subtypes of monkey vascular cells during aging. Targeted inactivation of FOXO3A in human vascular endothelial cells recapitulates the major phenotypic defects observed in aged monkey arteries, verifying FOXO3A loss as a key driver for arterial endothelial aging. Our study provides a critical resource for understanding the principles underlying primate arterial aging and contributes important clues to future treatment of age-associated vascular disorders. 1 CAS Key Laboratory of Genomic and Precision Medicine, Beijing Institute of Genomics, Chinese Academy of Sciences, Beijing 100101, China. -

Multidrug Transporter MRP4/ABCC4 As a Key Determinant of Pancreatic
www.nature.com/scientificreports OPEN Multidrug transporter MRP4/ ABCC4 as a key determinant of pancreatic cancer aggressiveness A. Sahores1, A. Carozzo1, M. May1, N. Gómez1, N. Di Siervi1, M. De Sousa Serro1, A. Yanef1, A. Rodríguez‑González2, M. Abba3, C. Shayo2 & C. Davio1* Recent fndings show that MRP4 is critical for pancreatic ductal adenocarcinoma (PDAC) cell proliferation. Nevertheless, the signifcance of MRP4 protein levels and function in PDAC progression is still unclear. The aim of this study was to determine the role of MRP4 in PDAC tumor aggressiveness. Bioinformatic studies revealed that PDAC samples show higher MRP4 transcript levels compared to normal adjacent pancreatic tissue and circulating tumor cells express higher levels of MRP4 than primary tumors. Also, high levels of MRP4 are typical of high-grade PDAC cell lines and associate with an epithelial-mesenchymal phenotype. Moreover, PDAC patients with high levels of MRP4 depict dysregulation of pathways associated with migration, chemotaxis and cell adhesion. Silencing MRP4 in PANC1 cells reduced tumorigenicity and tumor growth and impaired cell migration. Transcriptomic analysis revealed that MRP4 silencing alters PANC1 gene expression, mainly dysregulating pathways related to cell-to-cell interactions and focal adhesion. Contrarily, MRP4 overexpression signifcantly increased BxPC-3 growth rate, produced a switch in the expression of EMT markers, and enhanced experimental metastatic incidence. Altogether, our results indicate that MRP4 is associated with a more aggressive phenotype in PDAC, boosting pancreatic tumorigenesis and metastatic capacity, which could fnally determine a fast tumor progression in PDAC patients. Pancreatic ductal adenocarcinoma (PDAC) is one of the most lethal human malignancies, due to its late diag- nosis, inherent resistance to treatment and early dissemination 1. -

Screening and Identification of Hub Genes in Bladder Cancer by Bioinformatics Analysis and KIF11 Is a Potential Prognostic Biomarker
ONCOLOGY LETTERS 21: 205, 2021 Screening and identification of hub genes in bladder cancer by bioinformatics analysis and KIF11 is a potential prognostic biomarker XIAO‑CONG MO1,2*, ZI‑TONG ZHANG1,3*, MENG‑JIA SONG1,2, ZI‑QI ZHOU1,2, JIAN‑XIONG ZENG1,2, YU‑FEI DU1,2, FENG‑ZE SUN1,2, JIE‑YING YANG1,2, JUN‑YI HE1,2, YUE HUANG1,2, JIAN‑CHUAN XIA1,2 and DE‑SHENG WENG1,2 1State Key Laboratory of Oncology in South China, Collaborative Innovation Centre for Cancer Medicine; 2Department of Biotherapy, Sun Yat‑Sen University Cancer Center; 3Department of Radiation Oncology, Sun Yat‑Sen University Cancer Center, Guangzhou, Guangdong 510060, P.R. China Received July 31, 2020; Accepted December 18, 2020 DOI: 10.3892/ol.2021.12466 Abstract. Bladder cancer (BC) is the ninth most common immunohistochemistry and western blotting. In summary, lethal malignancy worldwide. Great efforts have been devoted KIF11 was significantly upregulated in BC and might act as to clarify the pathogenesis of BC, but the underlying molecular a potential prognostic biomarker. The present identification mechanisms remain unclear. To screen for the genes associated of DEGs and hub genes in BC may provide novel insight for with the progression and carcinogenesis of BC, three datasets investigating the molecular mechanisms of BC. were obtained from the Gene Expression Omnibus. A total of 37 tumor and 16 non‑cancerous samples were analyzed to Introduction identify differentially expressed genes (DEGs). Subsequently, 141 genes were identified, including 55 upregulated and Bladder cancer (BC) is the ninth most common malignancy 86 downregulated genes. The protein‑protein interaction worldwide with substantial morbidity and mortality. -

Accurate Breakpoint Mapping in Apparently Balanced Translocation Families with Discordant Phenotypes Using Whole Genome Mate-Pair Sequencing
Accurate Breakpoint Mapping in Apparently Balanced Translocation Families with Discordant Phenotypes Using Whole Genome Mate-Pair Sequencing Aristidou, Constantia; Koufaris, Costas; Theodosiou, Athina; Bak, Mads; Mehrjouy, Mana M; Behjati, Farkhondeh; Tanteles, George; Christophidou-Anastasiadou, Violetta; Tommerup, Niels; Sismani, Carolina Published in: PLOS ONE DOI: 10.1371/journal.pone.0169935 Publication date: 2017 Document version Publisher's PDF, also known as Version of record Document license: CC BY Citation for published version (APA): Aristidou, C., Koufaris, C., Theodosiou, A., Bak, M., Mehrjouy, M. M., Behjati, F., Tanteles, G., Christophidou- Anastasiadou, V., Tommerup, N., & Sismani, C. (2017). Accurate Breakpoint Mapping in Apparently Balanced Translocation Families with Discordant Phenotypes Using Whole Genome Mate-Pair Sequencing. PLOS ONE, 12(1), [e0169935]. https://doi.org/10.1371/journal.pone.0169935 Download date: 29. sep.. 2021 RESEARCH ARTICLE Accurate Breakpoint Mapping in Apparently Balanced Translocation Families with Discordant Phenotypes Using Whole Genome Mate-Pair Sequencing Constantia Aristidou1,2, Costas Koufaris1, Athina Theodosiou1, Mads Bak3, Mana M. Mehrjouy3, Farkhondeh Behjati4, George Tanteles5, Violetta Christophidou- Anastasiadou6, Niels Tommerup3, Carolina Sismani1,2* a1111111111 a1111111111 1 Department of Cytogenetics and Genomics, The Cyprus Institute of Neurology and Genetics, Nicosia, Cyprus, 2 The Cyprus School of Molecular Medicine, The Cyprus Institute of Neurology and Genetics, a1111111111