The Long Non-Coding RNA PCA3: an Update of Its Functions and Clinical Applications As a Biomarker in Prostate Cancer
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

S41467-020-18249-3.Pdf
ARTICLE https://doi.org/10.1038/s41467-020-18249-3 OPEN Pharmacologically reversible zonation-dependent endothelial cell transcriptomic changes with neurodegenerative disease associations in the aged brain Lei Zhao1,2,17, Zhongqi Li 1,2,17, Joaquim S. L. Vong2,3,17, Xinyi Chen1,2, Hei-Ming Lai1,2,4,5,6, Leo Y. C. Yan1,2, Junzhe Huang1,2, Samuel K. H. Sy1,2,7, Xiaoyu Tian 8, Yu Huang 8, Ho Yin Edwin Chan5,9, Hon-Cheong So6,8, ✉ ✉ Wai-Lung Ng 10, Yamei Tang11, Wei-Jye Lin12,13, Vincent C. T. Mok1,5,6,14,15 &HoKo 1,2,4,5,6,8,14,16 1234567890():,; The molecular signatures of cells in the brain have been revealed in unprecedented detail, yet the ageing-associated genome-wide expression changes that may contribute to neurovas- cular dysfunction in neurodegenerative diseases remain elusive. Here, we report zonation- dependent transcriptomic changes in aged mouse brain endothelial cells (ECs), which pro- minently implicate altered immune/cytokine signaling in ECs of all vascular segments, and functional changes impacting the blood–brain barrier (BBB) and glucose/energy metabolism especially in capillary ECs (capECs). An overrepresentation of Alzheimer disease (AD) GWAS genes is evident among the human orthologs of the differentially expressed genes of aged capECs, while comparative analysis revealed a subset of concordantly downregulated, functionally important genes in human AD brains. Treatment with exenatide, a glucagon-like peptide-1 receptor agonist, strongly reverses aged mouse brain EC transcriptomic changes and BBB leakage, with associated attenuation of microglial priming. We thus revealed tran- scriptomic alterations underlying brain EC ageing that are complex yet pharmacologically reversible. -

Downloaded from the Ensembl Database) Using the BWA MEM Algorithm (Version
bioRxiv preprint doi: https://doi.org/10.1101/2020.02.17.953281; this version posted March 18, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. 1 Identification of characteristic genomic markers in human hepatoma Huh7 and Huh7.5.1-8 2 cell lines 3 Masaki Kawamoto1, Toshiyuki Yamaji2, Kyoko Saito2, Kazuhiro Satomura3, Toshinori Endo3, 4 Masayoshi Fukasawa2, Kentaro Hanada2,*, and Naoki Osada3,4,* 5 1. Graduate School of Information Science and Technology, Hokkaido University, Sapporo 6 060-0814, Japan 7 2. Department of Biochemistry & Cell Biology, National Institute of Infectious Diseases, 8 Tokyo 162-8640, Japan 9 3. Faculty of Information Science and Technology, Hokkaido University, Sapporo 060-0814, 10 Japan 11 4. Global Station for Big Data and Cybersecurity, GI-CoRE, Hokkaido University, Sapporo 12 060-0814, Japan 13 14 *Correspondence 15 Kentaro Hanada 16 Email: [email protected] 17 Naoki Osada 18 Email: [email protected] 1 bioRxiv preprint doi: https://doi.org/10.1101/2020.02.17.953281; this version posted March 18, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. 19 Abstract 20 The human hepatoma-derived Huh7 cell line and its derivatives (Huh7.5 and Huh7.5.1) have been 21 widely used as a convenient experimental substitute for primary hepatocytes. -

Macropinocytosis Requires Gal-3 in a Subset of Patient-Derived Glioblastoma Stem Cells
ARTICLE https://doi.org/10.1038/s42003-021-02258-z OPEN Macropinocytosis requires Gal-3 in a subset of patient-derived glioblastoma stem cells Laetitia Seguin1,8, Soline Odouard2,8, Francesca Corlazzoli 2,8, Sarah Al Haddad2, Laurine Moindrot2, Marta Calvo Tardón3, Mayra Yebra4, Alexey Koval5, Eliana Marinari2, Viviane Bes3, Alexandre Guérin 6, Mathilde Allard2, Sten Ilmjärv6, Vladimir L. Katanaev 5, Paul R. Walker3, Karl-Heinz Krause6, Valérie Dutoit2, ✉ Jann N. Sarkaria 7, Pierre-Yves Dietrich2 & Érika Cosset 2 Recently, we involved the carbohydrate-binding protein Galectin-3 (Gal-3) as a druggable target for KRAS-mutant-addicted lung and pancreatic cancers. Here, using glioblastoma patient-derived stem cells (GSCs), we identify and characterize a subset of Gal-3high glio- 1234567890():,; blastoma (GBM) tumors mainly within the mesenchymal subtype that are addicted to Gal-3- mediated macropinocytosis. Using both genetic and pharmacologic inhibition of Gal-3, we showed a significant decrease of GSC macropinocytosis activity, cell survival and invasion, in vitro and in vivo. Mechanistically, we demonstrate that Gal-3 binds to RAB10, a member of the RAS superfamily of small GTPases, and β1 integrin, which are both required for macro- pinocytosis activity and cell survival. Finally, by defining a Gal-3/macropinocytosis molecular signature, we could predict sensitivity to this dependency pathway and provide proof-of- principle for innovative therapeutic strategies to exploit this Achilles’ heel for a significant and unique subset of GBM patients. 1 University Côte d’Azur, CNRS UMR7284, INSERM U1081, Institute for Research on Cancer and Aging (IRCAN), Nice, France. 2 Laboratory of Tumor Immunology, Department of Oncology, Center for Translational Research in Onco-Hematology, Swiss Cancer Center Léman (SCCL), Geneva University Hospitals, University of Geneva, Geneva, Switzerland. -

Single-Cell RNA-Seq Analysis of the Brainstem of Mutant SOD1 Mice Reveals Perturbed Cell Types and Pathways of Amyotrophic Lateral Sclerosis
HHS Public Access Author manuscript Author ManuscriptAuthor Manuscript Author Neurobiol Manuscript Author Dis. Author manuscript; Manuscript Author available in PMC 2020 September 26. Published in final edited form as: Neurobiol Dis. 2020 July ; 141: 104877. doi:10.1016/j.nbd.2020.104877. Single-cell RNA-seq analysis of the brainstem of mutant SOD1 mice reveals perturbed cell types and pathways of amyotrophic lateral sclerosis Wenting Liua, Sharmila Venugopala, Sana Majida, In Sook Ahna, Graciel Diamantea, Jason Honga, Xia Yanga,b,c,*, Scott H. Chandlera,b,* aDepartment of Integrative Biology & Physiology, University of California, 2024 Terasaki Bld, 610 Charles E. Young Dr. East, Los Angeles, USA bBrain Research Institute, University of California, Los Angeles, USA cInstitute for Quantitative and Computational Biosciences, University of California, Los Angeles, USA Abstract Amyotrophic lateral sclerosis (ALS) is a neurodegenerative disease in which motor neurons throughout the brain and spinal cord progressively degenerate resulting in muscle atrophy, paralysis and death. Recent studies using animal models of ALS implicate multiple cell-types (e.g., astrocytes and microglia) in ALS pathogenesis in the spinal motor systems. To ascertain cellular vulnerability and cell-type specific mechanisms of ALS in the brainstem that orchestrates oral-motor functions, we conducted parallel single cell RNA sequencing (scRNA-seq) analysis using the high-throughput Drop-seq method. We isolated 1894 and 3199 cells from the brainstem of wildtype and mutant SOD1 symptomatic mice respectively, at postnatal day 100. We recovered major known cell types and neuronal subpopulations, such as interneurons and motor neurons, and trigeminal ganglion (TG) peripheral sensory neurons, as well as, previously uncharacterized interneuron subtypes. -

Comparative Transcriptome Profiling of the Human and Mouse Dorsal Root Ganglia: an RNA-Seq-Based Resource for Pain and Sensory Neuroscience Research
bioRxiv preprint doi: https://doi.org/10.1101/165431; this version posted October 13, 2017. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Title: Comparative transcriptome profiling of the human and mouse dorsal root ganglia: An RNA-seq-based resource for pain and sensory neuroscience research Short Title: Human and mouse DRG comparative transcriptomics Pradipta Ray 1, 2 #, Andrew Torck 1 , Lilyana Quigley 1, Andi Wangzhou 1, Matthew Neiman 1, Chandranshu Rao 1, Tiffany Lam 1, Ji-Young Kim 1, Tae Hoon Kim 2, Michael Q. Zhang 2, Gregory Dussor 1 and Theodore J. Price 1, # 1 The University of Texas at Dallas, School of Behavioral and Brain Sciences 2 The University of Texas at Dallas, Department of Biological Sciences # Corresponding authors Theodore J Price Pradipta Ray School of Behavioral and Brain Sciences School of Behavioral and Brain Sciences The University of Texas at Dallas The University of Texas at Dallas BSB 14.102G BSB 10.608 800 W Campbell Rd 800 W Campbell Rd Richardson TX 75080 Richardson TX 75080 972-883-4311 972-883-7262 [email protected] [email protected] Number of pages: 27 Number of figures: 9 Number of tables: 8 Supplementary Figures: 4 Supplementary Files: 6 Word count: Abstract = 219; Introduction = 457; Discussion = 1094 Conflict of interest: The authors declare no conflicts of interest Patient anonymity and informed consent: Informed consent for human tissue sources were obtained by Anabios, Inc. (San Diego, CA). Human studies: This work was approved by The University of Texas at Dallas Institutional Review Board (MR 15-237). -

ID AKI Vs Control Fold Change P Value Symbol Entrez Gene Name *In
ID AKI vs control P value Symbol Entrez Gene Name *In case of multiple probesets per gene, one with the highest fold change was selected. Fold Change 208083_s_at 7.88 0.000932 ITGB6 integrin, beta 6 202376_at 6.12 0.000518 SERPINA3 serpin peptidase inhibitor, clade A (alpha-1 antiproteinase, antitrypsin), member 3 1553575_at 5.62 0.0033 MT-ND6 NADH dehydrogenase, subunit 6 (complex I) 212768_s_at 5.50 0.000896 OLFM4 olfactomedin 4 206157_at 5.26 0.00177 PTX3 pentraxin 3, long 212531_at 4.26 0.00405 LCN2 lipocalin 2 215646_s_at 4.13 0.00408 VCAN versican 202018_s_at 4.12 0.0318 LTF lactotransferrin 203021_at 4.05 0.0129 SLPI secretory leukocyte peptidase inhibitor 222486_s_at 4.03 0.000329 ADAMTS1 ADAM metallopeptidase with thrombospondin type 1 motif, 1 1552439_s_at 3.82 0.000714 MEGF11 multiple EGF-like-domains 11 210602_s_at 3.74 0.000408 CDH6 cadherin 6, type 2, K-cadherin (fetal kidney) 229947_at 3.62 0.00843 PI15 peptidase inhibitor 15 204006_s_at 3.39 0.00241 FCGR3A Fc fragment of IgG, low affinity IIIa, receptor (CD16a) 202238_s_at 3.29 0.00492 NNMT nicotinamide N-methyltransferase 202917_s_at 3.20 0.00369 S100A8 S100 calcium binding protein A8 215223_s_at 3.17 0.000516 SOD2 superoxide dismutase 2, mitochondrial 204627_s_at 3.04 0.00619 ITGB3 integrin, beta 3 (platelet glycoprotein IIIa, antigen CD61) 223217_s_at 2.99 0.00397 NFKBIZ nuclear factor of kappa light polypeptide gene enhancer in B-cells inhibitor, zeta 231067_s_at 2.97 0.00681 AKAP12 A kinase (PRKA) anchor protein 12 224917_at 2.94 0.00256 VMP1/ mir-21likely ortholog -

A Novel Tool for Detection of Tissue and Cell Specific Marker Genes From
El Amrani et al. BMC Genomics (2015) 16:645 DOI 10.1186/s12864-015-1785-9 METHODOLOGY ARTICLE Open Access MGFM: a novel tool for detection of tissue and cell specific marker genes from microarray gene expression data Khadija El Amrani1, Harald Stachelscheid1,2, Fritz Lekschas1, Andreas Kurtz1,3* and Miguel A. Andrade-Navarro4,5 Abstract Background: Identification of marker genes associated with a specific tissue/cell type is a fundamental challenge in genetic and cell research. Marker genes are of great importance for determining cell identity, and for understanding tissue specific gene function and the molecular mechanisms underlying complex diseases. Results: We have developed a new bioinformatics tool called MGFM (Marker Gene Finder in Microarray data) to predict marker genes from microarray gene expression data. Marker genes are identified through the grouping of samples of the same type with similar marker gene expression levels. We verified our approach using two microarray data sets from the NCBI’s Gene Expression Omnibus public repository encompassing samples for similar sets of five human tissues (brain, heart, kidney, liver, and lung). Comparison with another tool for tissue-specific gene identification and validation with literature-derived established tissue markers established functionality, accuracy and simplicity of our tool. Furthermore, top ranked marker genes were experimentally validated by reverse transcriptase-polymerase chain reaction (RT-PCR). The sets of predicted marker genes associated with the five selected tissues comprised well-known genes of particular importance in these tissues. The tool is freely available from the Bioconductor web site, and it is also provided as an online application integrated into the CellFinder platform (http://cellfinder.org/analysis/marker). -

Comprehensive Analysis Reveals Novel Gene Signature in Head and Neck Squamous Cell Carcinoma: Predicting Is Associated with Poor Prognosis in Patients
5892 Original Article Comprehensive analysis reveals novel gene signature in head and neck squamous cell carcinoma: predicting is associated with poor prognosis in patients Yixin Sun1,2#, Quan Zhang1,2#, Lanlin Yao2#, Shuai Wang3, Zhiming Zhang1,2 1Department of Breast Surgery, The First Affiliated Hospital of Xiamen University, School of Medicine, Xiamen University, Xiamen, China; 2School of Medicine, Xiamen University, Xiamen, China; 3State Key Laboratory of Cellular Stress Biology, School of Life Sciences, Xiamen University, Xiamen, China Contributions: (I) Conception and design: Y Sun, Q Zhang; (II) Administrative support: Z Zhang; (III) Provision of study materials or patients: Y Sun, Q Zhang; (IV) Collection and assembly of data: Y Sun, L Yao; (V) Data analysis and interpretation: Y Sun, S Wang; (VI) Manuscript writing: All authors; (VII) Final approval of manuscript: All authors. #These authors contributed equally to this work. Correspondence to: Zhiming Zhang. Department of Surgery, The First Affiliated Hospital of Xiamen University, Xiamen, China. Email: [email protected]. Background: Head and neck squamous cell carcinoma (HNSC) remains an important public health problem, with classic risk factors being smoking and excessive alcohol consumption and usually has a poor prognosis. Therefore, it is important to explore the underlying mechanisms of tumorigenesis and screen the genes and pathways identified from such studies and their role in pathogenesis. The purpose of this study was to identify genes or signal pathways associated with the development of HNSC. Methods: In this study, we downloaded gene expression profiles of GSE53819 from the Gene Expression Omnibus (GEO) database, including 18 HNSC tissues and 18 normal tissues. -

Genomic Profiling Reveals Mutational Landscape in Parathyroid Carcinomas
RESEARCH ARTICLE Genomic profiling reveals mutational landscape in parathyroid carcinomas Chetanya Pandya,1 Andrew V. Uzilov,1 Justin Bellizzi,2 Chun Yee Lau,1 Aye S. Moe,1 Maya Strahl,1 Wissam Hamou,1 Leah C. Newman,1 Marc Y. Fink,1 Yevgeniy Antipin,1 Willie Yu,3 Mark Stevenson,4 Branca M. Cavaco,5 Bin T. Teh,6,7 Rajesh V. Thakker,4 Hans Morreau,8 Eric E. Schadt,1 Robert Sebra,1 Shuyu D. Li,1 Andrew Arnold,2 and Rong Chen1 1 Department of Genetics and Genomic Sciences, Icahn School of Medicine at Mount Sinai, New York, New York, USA. 2Center for Molecular Medicine, University of Connecticut School of Medicine, Farmington, Connecticut, USA. 3Center for Computational Biology, Duke–National University of Singapore Graduate Medical School, Singapore. 4Academic Endocrine Unit, Radcliffe Department of Medicine, Oxford Centre for Diabetes, Endocrinology and Metabolism (OCDEM), University of Oxford, Oxford, United Kingdom. 5Molecular Endocrinology Group, Molecular Pathobiology Research Centre Unit (UIPM) of the Portuguese Institute of Oncology from Lisbon Francisco Gentil (IPOLFG), Lisbon, Portugal. 6Laboratory of Cancer Epigenome, Division of Medical Sciences, National Cancer Centre Singapore, Singapore. 7Cancer and Stem Cell Biology Program, Duke–National University of Singapore Graduate Medical School, Singapore. 8Department of Pathology, Leiden University Medical Center, Leiden, The Netherlands. Parathyroid carcinoma (PC) is an extremely rare malignancy lacking effective therapeutic intervention. We generated and analyzed whole-exome sequencing data from 17 patients to identify somatic and germline genetic alterations. A panel of selected genes was sequenced in a 7-tumor expansion cohort. We show that 47% (8 of 17) of the tumors harbor somatic mutations in the CDC73 tumor suppressor, with germline inactivating variants in 4 of the 8 patients. -

The RNA Editing Proteins
Biomolecules 2015, 5, 2338-2362; doi:10.3390/biom5042338 OPEN ACCESS biomolecules ISSN 2218-273X www.mdpi.com/journal/biomolecules/ Review New Insights into the Biological Role of Mammalian ADARs; the RNA Editing Proteins Niamh Mannion 1, Fabiana Arieti 2, Angela Gallo 3, Liam P. Keegan 2 and Mary A. O’Connell 2,* 1 Paul O’Gorman Leukaemia Research Centre, Institute of Cancer Sciences, College of Medical, Veterinary and Life Sciences, University of Glasgow, 21 Shelley Road, Glasgow G12 0ZD, UK; E-Mail: [email protected] 2 CEITEC—Central European Institute of Technology, Masaryk University, Kamenice 5, Brno 625 00, Czech Republic; E-Mails: [email protected] (F.A.); [email protected] (L.P.K.) 3 Oncohaematoogy Department, Ospedale Pediatrico Bambino Gesù (IRCCS) Viale di San Paolo, Roma 15-00146, Italy; E-Mail: [email protected] * Author to whom correspondence should be addressed; E-Mail: [email protected]; Tel.: + 420-549-495-460. Academic Editor: André P. Gerber Received: 24 July 2015 / Accepted: 11 September 2015 / Published: 30 September 2015 Abstract: The ADAR proteins deaminate adenosine to inosine in double-stranded RNA which is one of the most abundant modifications present in mammalian RNA. Inosine can have a profound effect on the RNAs that are edited, not only changing the base-pairing properties, but can also result in recoding, as inosine behaves as if it were guanosine. In mammals there are three ADAR proteins and two ADAR-related proteins (ADAD) expressed. All have a very similar modular structure; however, both their expression and biological function differ significantly. -
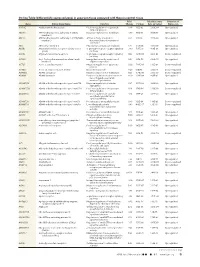
Differentially Expressed Genes in Aneurysm Tissue Compared With
On-line Table: Differentially expressed genes in aneurysm tissue compared with those in control tissue Fold False Discovery Direction of Gene Entrez Gene Name Function Change P Value Rate (q Value) Expression AADAC Arylacetamide deacetylase Positive regulation of triglyceride 4.46 1.33E-05 2.60E-04 Up-regulated catabolic process ABCA6 ATP-binding cassette, subfamily A (ABC1), Integral component of membrane 3.79 9.15E-14 8.88E-12 Up-regulated member 6 ABCC3 ATP-binding cassette, subfamily C (CFTR/MRP), ATPase activity, coupled to 6.63 1.21E-10 7.33E-09 Up-regulated member 3 transmembrane movement of substances ABI3 ABI family, member 3 Peptidyl-tyrosine phosphorylation 6.47 2.47E-05 4.56E-04 Up-regulated ACKR1 Atypical chemokine receptor 1 (Duffy blood G-protein–coupled receptor signaling 3.80 7.95E-10 4.18E-08 Up-regulated group) pathway ACKR2 Atypical chemokine receptor 2 G-protein–coupled receptor signaling 0.42 3.29E-04 4.41E-03 Down-regulated pathway ACSM1 Acyl-CoA synthetase medium-chain family Energy derivation by oxidation of 9.87 1.70E-08 6.52E-07 Up-regulated member 1 organic compounds ACTC1 Actin, ␣, cardiac muscle 1 Negative regulation of apoptotic 0.30 7.96E-06 1.65E-04 Down-regulated process ACTG2 Actin, ␥2, smooth muscle, enteric Blood microparticle 0.29 1.61E-16 2.36E-14 Down-regulated ADAM33 ADAM domain 33 Integral component of membrane 0.23 9.74E-09 3.95E-07 Down-regulated ADAM8 ADAM domain 8 Positive regulation of tumor necrosis 4.69 2.93E-04 4.01E-03 Up-regulated factor (ligand) superfamily member 11 production ADAMTS18