Environmental Cues Regulate Epigenetic Reprogramming of Airway-Resident Memory CD8+ T Cells
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

An Evaluation of Host Factors As Novel Therapeutic Targets During Influenza Infection Using RNA Technologies
California State University, San Bernardino CSUSB ScholarWorks Electronic Theses, Projects, and Dissertations Office of aduateGr Studies 6-2018 An Evaluation of Host Factors as Novel Therapeutic Targets During Influenza Infection Using RNA Technologies Michael Ryan Haden Thompson California State University - San Bernardino Follow this and additional works at: https://scholarworks.lib.csusb.edu/etd Part of the Virology Commons Recommended Citation Thompson, Michael Ryan Haden, "An Evaluation of Host Factors as Novel Therapeutic Targets During Influenza Infection Using RNA Technologies" (2018). Electronic Theses, Projects, and Dissertations. 721. https://scholarworks.lib.csusb.edu/etd/721 This Thesis is brought to you for free and open access by the Office of aduateGr Studies at CSUSB ScholarWorks. It has been accepted for inclusion in Electronic Theses, Projects, and Dissertations by an authorized administrator of CSUSB ScholarWorks. For more information, please contact [email protected]. AN EVALUATION OF HOST FACTORS AS NOVEL THERAPEUTIC TARGETS DURING INFLUENZA INFECTION USING RNA TECHNOLOGIES _____________________ A Thesis Presented to the Faculty of California State University, San Bernardino _____________________ In Partial Fulfillment of the Requirements for the Degree Master of Science in Biology _____________________ by Michael Ryan Haden Thompson June 2018 AN EVALUATION OF HOST FACTORS AS NOVEL THERAPEUTIC TARGETS DURING INFLUENZA INFECTION USING RNA TECHNOLOGIES _____________________ A Thesis Presented to the Faculty of California State University, San Bernardino _____________________ By Michael Ryan Haden Thompson June 2018 Approved by: Dr. Laura L Newcomb, Committee Chair, Biology Dr. Nicole Bournias-Vardiabasis, Committee Member Dr. Daniel Nickerson, Committee Member © 2018 Michael Ryan Haden Thompson ABSTRACT Influenza A is a single-stranded, multi-segmented, negative sense RNA virus of the family Orthomyxoviridae and is the causative agent of seasonal Influenza. -

Mir-17-92 Fine-Tunes MYC Expression and Function to Ensure
ARTICLE Received 31 Mar 2015 | Accepted 22 Sep 2015 | Published 10 Nov 2015 DOI: 10.1038/ncomms9725 OPEN miR-17-92 fine-tunes MYC expression and function to ensure optimal B cell lymphoma growth Marija Mihailovich1, Michael Bremang1, Valeria Spadotto1, Daniele Musiani1, Elena Vitale1, Gabriele Varano2,w, Federico Zambelli3, Francesco M. Mancuso1,w, David A. Cairns1,w, Giulio Pavesi3, Stefano Casola2 & Tiziana Bonaldi1 The synergism between c-MYC and miR-17-19b, a truncated version of the miR-17-92 cluster, is well-documented during tumor initiation. However, little is known about miR-17-19b function in established cancers. Here we investigate the role of miR-17-19b in c-MYC-driven lymphomas by integrating SILAC-based quantitative proteomics, transcriptomics and 30 untranslated region (UTR) analysis upon miR-17-19b overexpression. We identify over one hundred miR-17-19b targets, of which 40% are co-regulated by c-MYC. Downregulation of a new miR-17/20 target, checkpoint kinase 2 (Chek2), increases the recruitment of HuR to c- MYC transcripts, resulting in the inhibition of c-MYC translation and thus interfering with in vivo tumor growth. Hence, in established lymphomas, miR-17-19b fine-tunes c-MYC activity through a tight control of its function and expression, ultimately ensuring cancer cell homeostasis. Our data highlight the plasticity of miRNA function, reflecting changes in the mRNA landscape and 30 UTR shortening at different stages of tumorigenesis. 1 Department of Experimental Oncology, European Institute of Oncology, Via Adamello 16, Milan 20139, Italy. 2 Units of Genetics of B cells and lymphomas, IFOM, FIRC Institute of Molecular Oncology Foundation, Milan 20139, Italy. -

Aneuploidy: Using Genetic Instability to Preserve a Haploid Genome?
Health Science Campus FINAL APPROVAL OF DISSERTATION Doctor of Philosophy in Biomedical Science (Cancer Biology) Aneuploidy: Using genetic instability to preserve a haploid genome? Submitted by: Ramona Ramdath In partial fulfillment of the requirements for the degree of Doctor of Philosophy in Biomedical Science Examination Committee Signature/Date Major Advisor: David Allison, M.D., Ph.D. Academic James Trempe, Ph.D. Advisory Committee: David Giovanucci, Ph.D. Randall Ruch, Ph.D. Ronald Mellgren, Ph.D. Senior Associate Dean College of Graduate Studies Michael S. Bisesi, Ph.D. Date of Defense: April 10, 2009 Aneuploidy: Using genetic instability to preserve a haploid genome? Ramona Ramdath University of Toledo, Health Science Campus 2009 Dedication I dedicate this dissertation to my grandfather who died of lung cancer two years ago, but who always instilled in us the value and importance of education. And to my mom and sister, both of whom have been pillars of support and stimulating conversations. To my sister, Rehanna, especially- I hope this inspires you to achieve all that you want to in life, academically and otherwise. ii Acknowledgements As we go through these academic journeys, there are so many along the way that make an impact not only on our work, but on our lives as well, and I would like to say a heartfelt thank you to all of those people: My Committee members- Dr. James Trempe, Dr. David Giovanucchi, Dr. Ronald Mellgren and Dr. Randall Ruch for their guidance, suggestions, support and confidence in me. My major advisor- Dr. David Allison, for his constructive criticism and positive reinforcement. -

Therapeutic Targeting of Nuclear Transport
cells Review Controlling the Gatekeeper: Therapeutic Targeting of Nuclear Transport Friederike K. Kosyna * and Reinhard Depping Institute of Physiology, Center for Structural and Cell Biology in Medicine, University of Lübeck, Ratzeburger Allee 160, D-23562 Lübeck, Germany; [email protected] * Correspondence: [email protected]; Tel.: +49-451-3101-7322; Fax: +49-451-3101-7304 Received: 25 October 2018; Accepted: 17 November 2018; Published: 21 November 2018 Abstract: Nuclear transport receptors of the karyopherin superfamily of proteins transport macromolecules from one compartment to the other and are critical for both cell physiology and pathophysiology. The nuclear transport machinery is tightly regulated and essential to a number of key cellular processes since the spatiotemporally expression of many proteins and the nuclear transporters themselves is crucial for cellular activities. Dysregulation of the nuclear transport machinery results in localization shifts of specific cargo proteins and associates with the pathogenesis of disease states such as cancer, inflammation, viral illness and neurodegenerative diseases. Therefore, inhibition of the nuclear transport system has future potential for therapeutic intervention and could contribute to the elucidation of disease mechanisms. In this review, we recapitulate clue findings in the pathophysiological significance of nuclear transport processes and describe the development of nuclear transport inhibitors. Finally, clinical implications and results of the first clinical trials are discussed for the most promising nuclear transport inhibitors. Keywords: nuclear transport; exportin; importin; karyopherin; chromosome region maintenance 1 (CRM1); cancer; drug; nuclear transport inhibitor 1. Introduction The cytoplasm and the nucleoplasm are separated by the nuclear envelope in eukaryotic cells. Spatially segregation of essential cellular processes requires tight control of large molecule exchange such as RNAs, proteins, or ribonucleoprotein particles through this double membrane. -

Downloaded the “Top Edge” Version
bioRxiv preprint doi: https://doi.org/10.1101/855338; this version posted December 6, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY 4.0 International license. 1 Drosophila models of pathogenic copy-number variant genes show global and 2 non-neuronal defects during development 3 Short title: Non-neuronal defects of fly homologs of CNV genes 4 Tanzeen Yusuff1,4, Matthew Jensen1,4, Sneha Yennawar1,4, Lucilla Pizzo1, Siddharth 5 Karthikeyan1, Dagny J. Gould1, Avik Sarker1, Yurika Matsui1,2, Janani Iyer1, Zhi-Chun Lai1,2, 6 and Santhosh Girirajan1,3* 7 8 1. Department of Biochemistry and Molecular Biology, Pennsylvania State University, 9 University Park, PA 16802 10 2. Department of Biology, Pennsylvania State University, University Park, PA 16802 11 3. Department of Anthropology, Pennsylvania State University, University Park, PA 16802 12 4 contributed equally to work 13 14 *Correspondence: 15 Santhosh Girirajan, MBBS, PhD 16 205A Life Sciences Building 17 Pennsylvania State University 18 University Park, PA 16802 19 E-mail: [email protected] 20 Phone: 814-865-0674 21 1 bioRxiv preprint doi: https://doi.org/10.1101/855338; this version posted December 6, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY 4.0 International license. 22 ABSTRACT 23 While rare pathogenic copy-number variants (CNVs) are associated with both neuronal and non- 24 neuronal phenotypes, functional studies evaluating these regions have focused on the molecular 25 basis of neuronal defects. -

Open Data for Differential Network Analysis in Glioma
International Journal of Molecular Sciences Article Open Data for Differential Network Analysis in Glioma , Claire Jean-Quartier * y , Fleur Jeanquartier y and Andreas Holzinger Holzinger Group HCI-KDD, Institute for Medical Informatics, Statistics and Documentation, Medical University Graz, Auenbruggerplatz 2/V, 8036 Graz, Austria; [email protected] (F.J.); [email protected] (A.H.) * Correspondence: [email protected] These authors contributed equally to this work. y Received: 27 October 2019; Accepted: 3 January 2020; Published: 15 January 2020 Abstract: The complexity of cancer diseases demands bioinformatic techniques and translational research based on big data and personalized medicine. Open data enables researchers to accelerate cancer studies, save resources and foster collaboration. Several tools and programming approaches are available for analyzing data, including annotation, clustering, comparison and extrapolation, merging, enrichment, functional association and statistics. We exploit openly available data via cancer gene expression analysis, we apply refinement as well as enrichment analysis via gene ontology and conclude with graph-based visualization of involved protein interaction networks as a basis for signaling. The different databases allowed for the construction of huge networks or specified ones consisting of high-confidence interactions only. Several genes associated to glioma were isolated via a network analysis from top hub nodes as well as from an outlier analysis. The latter approach highlights a mitogen-activated protein kinase next to a member of histondeacetylases and a protein phosphatase as genes uncommonly associated with glioma. Cluster analysis from top hub nodes lists several identified glioma-associated gene products to function within protein complexes, including epidermal growth factors as well as cell cycle proteins or RAS proto-oncogenes. -

Case Report 12Q14 Microdeletions: Additional Case Series with Confirmation of a Macrocephaly Region
Hindawi Publishing Corporation Case Reports in Genetics Volume 2015, Article ID 192071, 7 pages http://dx.doi.org/10.1155/2015/192071 Case Report 12q14 Microdeletions: Additional Case Series with Confirmation of a Macrocephaly Region Adrian Mc Cormack,1 Cynthia Sharpe,2 Nerine Gregersen,3 Warwick Smith,4 Ian Hayes,3 Alice M. George,1 and Donald R. Love1 1 Diagnostic Genetics, LabPLUS, Auckland City Hospital, P.O. Box 110031, Auckland 1148, New Zealand 2Department of Neuroservices, Starship Children’s Health, Private Bag 92024, Auckland 1142, New Zealand 3Genetic Health Service New Zealand-Northern Hub, Auckland City Hospital, Private Bag 92024, Auckland 1142, New Zealand 4Middlemore Hospital, Private Bag 93311, Otahuhu, Auckland 1640, New Zealand Correspondence should be addressed to Donald R. Love; [email protected] Received 16 April 2015; Revised 7 July 2015; Accepted 7 July 2015 Academic Editor: Patrick Morrison Copyright © 2015 Adrian Mc Cormack et al. This is an open access article distributed under the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited. To date, there have been only a few reports of patients carrying a microdeletion in chromosome 12q14. These patients usually present with pre- and postnatal growth retardation, and developmental delay. Here we report on two additional patients with both genotype and phenotype differences. Similar to previously published cases, one patient has haploinsufficiency ofthe HMGA2 gene and shows severe short stature and developmental delay. The second patient is only one of a handful without the loss of the HMGA2 gene and shows a much better growth profile, but with absolute macrocephaly. -
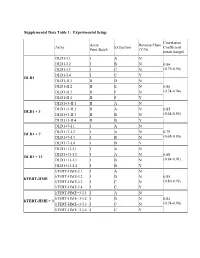
Table 3: Average Gene Expression Profiles by Chromosome
Supplemental Data Table 1: Experimental Setup Correlation Array Reverse Fluor Array Extraction Coefficient Print Batch (Y/N) mean (range) DLD1-I.1 I A N DLD1-I.2 I B N 0.86 DLD1-I.3 I C N (0.79-0.90) DLD1-I.4 I C Y DLD1 DLD1-II.1 II D N DLD1-II.2 II E N 0.86 DLD1-II.3 II F N (0.74-0.94) DLD1-II.4 II F Y DLD1+3-II.1 II A N DLD1+3-II.2 II A N 0.85 DLD1 + 3 DLD1+3-II.3 II B N (0.64-0.95) DLD1+3-II.4 II B Y DLD1+7-I.1 I A N DLD1+7-I.2 I A N 0.79 DLD1 + 7 DLD1+7-I.3 I B N (0.68-0.90) DLD1+7-I.4 I B Y DLD1+13-I.1 I A N DLD1+13-I.2 I A N 0.88 DLD1 + 13 DLD1+13-I.3 I B N (0.84-0.91) DLD1+13-I.4 I B Y hTERT-HME-I.1 I A N hTERT-HME-I.2 I B N 0.85 hTERT-HME hTERT-HME-I.3 I C N (0.80-0.92) hTERT-HME-I.4 I C Y hTERT-HME+3-I.1 I A N hTERT-HME+3-I.2 I B N 0.84 hTERT-HME + 3 hTERT-HME+3-I.3 I C N (0.74-0.90) hTERT-HME+3-I.4 I C Y Supplemental Data Table 2: Average gene expression profiles by chromosome arm DLD1 hTERT-HME Ratio.7 Ratio.1 Ratio.3 Ratio.3 Chrom. -

Nº Ref Uniprot Proteína Péptidos Identificados Por MS/MS 1 P01024
Document downloaded from http://www.elsevier.es, day 26/09/2021. This copy is for personal use. Any transmission of this document by any media or format is strictly prohibited. Nº Ref Uniprot Proteína Péptidos identificados 1 P01024 CO3_HUMAN Complement C3 OS=Homo sapiens GN=C3 PE=1 SV=2 por 162MS/MS 2 P02751 FINC_HUMAN Fibronectin OS=Homo sapiens GN=FN1 PE=1 SV=4 131 3 P01023 A2MG_HUMAN Alpha-2-macroglobulin OS=Homo sapiens GN=A2M PE=1 SV=3 128 4 P0C0L4 CO4A_HUMAN Complement C4-A OS=Homo sapiens GN=C4A PE=1 SV=1 95 5 P04275 VWF_HUMAN von Willebrand factor OS=Homo sapiens GN=VWF PE=1 SV=4 81 6 P02675 FIBB_HUMAN Fibrinogen beta chain OS=Homo sapiens GN=FGB PE=1 SV=2 78 7 P01031 CO5_HUMAN Complement C5 OS=Homo sapiens GN=C5 PE=1 SV=4 66 8 P02768 ALBU_HUMAN Serum albumin OS=Homo sapiens GN=ALB PE=1 SV=2 66 9 P00450 CERU_HUMAN Ceruloplasmin OS=Homo sapiens GN=CP PE=1 SV=1 64 10 P02671 FIBA_HUMAN Fibrinogen alpha chain OS=Homo sapiens GN=FGA PE=1 SV=2 58 11 P08603 CFAH_HUMAN Complement factor H OS=Homo sapiens GN=CFH PE=1 SV=4 56 12 P02787 TRFE_HUMAN Serotransferrin OS=Homo sapiens GN=TF PE=1 SV=3 54 13 P00747 PLMN_HUMAN Plasminogen OS=Homo sapiens GN=PLG PE=1 SV=2 48 14 P02679 FIBG_HUMAN Fibrinogen gamma chain OS=Homo sapiens GN=FGG PE=1 SV=3 47 15 P01871 IGHM_HUMAN Ig mu chain C region OS=Homo sapiens GN=IGHM PE=1 SV=3 41 16 P04003 C4BPA_HUMAN C4b-binding protein alpha chain OS=Homo sapiens GN=C4BPA PE=1 SV=2 37 17 Q9Y6R7 FCGBP_HUMAN IgGFc-binding protein OS=Homo sapiens GN=FCGBP PE=1 SV=3 30 18 O43866 CD5L_HUMAN CD5 antigen-like OS=Homo -

A Spatially Aware Likelihood Test to Detect Sweeps from Haplotype Distributions
bioRxiv preprint doi: https://doi.org/10.1101/2021.05.12.443825; this version posted May 13, 2021. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. A spatially aware likelihood test to detect sweeps from haplotype distributions Michael DeGiorgio1;∗, Zachary A. Szpiech2;3;∗ 1Department of Computer and Electrical Engineering and Computer Science, Florida Atlantic University, Boca Raton, FL 33431, USA 2Department of Biology, Pennsylvania State University, University Park, PA 16801, USA 3Institute for Computational and Data Sciences, Pennsylvania State University, University Park, PA 16801, USA ∗Corresponding authors: [email protected] (M.D.), [email protected] (Z.A.S.) Keywords: haplotypes, hard sweeps, soft sweeps, maximum likelihood 1 bioRxiv preprint doi: https://doi.org/10.1101/2021.05.12.443825; this version posted May 13, 2021. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. Abstract The inference of positive selection in genomes is a problem of great interest in evolutionary genomics. By identifying putative regions of the genome that contain adaptive mutations, we are able to learn about the biology of organisms and their evolutionary history. Here we introduce a composite likelihood method that identifies recently completed or ongoing positive selection by searching for extreme distortions in the spatial distribution of the haplotype fre- quency spectrum relative to the genome-wide expectation taken as neutrality. -

A High-Throughput Approach to Uncover Novel Roles of APOBEC2, a Functional Orphan of the AID/APOBEC Family
Rockefeller University Digital Commons @ RU Student Theses and Dissertations 2018 A High-Throughput Approach to Uncover Novel Roles of APOBEC2, a Functional Orphan of the AID/APOBEC Family Linda Molla Follow this and additional works at: https://digitalcommons.rockefeller.edu/ student_theses_and_dissertations Part of the Life Sciences Commons A HIGH-THROUGHPUT APPROACH TO UNCOVER NOVEL ROLES OF APOBEC2, A FUNCTIONAL ORPHAN OF THE AID/APOBEC FAMILY A Thesis Presented to the Faculty of The Rockefeller University in Partial Fulfillment of the Requirements for the degree of Doctor of Philosophy by Linda Molla June 2018 © Copyright by Linda Molla 2018 A HIGH-THROUGHPUT APPROACH TO UNCOVER NOVEL ROLES OF APOBEC2, A FUNCTIONAL ORPHAN OF THE AID/APOBEC FAMILY Linda Molla, Ph.D. The Rockefeller University 2018 APOBEC2 is a member of the AID/APOBEC cytidine deaminase family of proteins. Unlike most of AID/APOBEC, however, APOBEC2’s function remains elusive. Previous research has implicated APOBEC2 in diverse organisms and cellular processes such as muscle biology (in Mus musculus), regeneration (in Danio rerio), and development (in Xenopus laevis). APOBEC2 has also been implicated in cancer. However the enzymatic activity, substrate or physiological target(s) of APOBEC2 are unknown. For this thesis, I have combined Next Generation Sequencing (NGS) techniques with state-of-the-art molecular biology to determine the physiological targets of APOBEC2. Using a cell culture muscle differentiation system, and RNA sequencing (RNA-Seq) by polyA capture, I demonstrated that unlike the AID/APOBEC family member APOBEC1, APOBEC2 is not an RNA editor. Using the same system combined with enhanced Reduced Representation Bisulfite Sequencing (eRRBS) analyses I showed that, unlike the AID/APOBEC family member AID, APOBEC2 does not act as a 5-methyl-C deaminase. -

Single Cell Transcriptional and Chromatin Accessibility Profiling Redefine Cellular Heterogeneity in the Adult Human Kidney
ARTICLE https://doi.org/10.1038/s41467-021-22368-w OPEN Single cell transcriptional and chromatin accessibility profiling redefine cellular heterogeneity in the adult human kidney Yoshiharu Muto 1,7, Parker C. Wilson 2,7, Nicolas Ledru 1, Haojia Wu1, Henrik Dimke 3,4, ✉ Sushrut S. Waikar 5 & Benjamin D. Humphreys 1,6 1234567890():,; The integration of single cell transcriptome and chromatin accessibility datasets enables a deeper understanding of cell heterogeneity. We performed single nucleus ATAC (snATAC- seq) and RNA (snRNA-seq) sequencing to generate paired, cell-type-specific chromatin accessibility and transcriptional profiles of the adult human kidney. We demonstrate that snATAC-seq is comparable to snRNA-seq in the assignment of cell identity and can further refine our understanding of functional heterogeneity in the nephron. The majority of differ- entially accessible chromatin regions are localized to promoters and a significant proportion are closely associated with differentially expressed genes. Cell-type-specific enrichment of transcription factor binding motifs implicates the activation of NF-κB that promotes VCAM1 expression and drives transition between a subpopulation of proximal tubule epithelial cells. Our multi-omics approach improves the ability to detect unique cell states within the kidney and redefines cellular heterogeneity in the proximal tubule and thick ascending limb. 1 Division of Nephrology, Department of Medicine, Washington University in St. Louis, St. Louis, MO, USA. 2 Department of Pathology and Immunology, Washington University in St. Louis, St. Louis, MO, USA. 3 Department of Cardiovascular and Renal Research, Institute of Molecular Medicine, University of Southern Denmark, Odense, Denmark. 4 Department of Nephrology, Odense University Hospital, Odense, Denmark.