The Effects of Ecology and Evolution on Avian Flight Morphology Natalie Wright
Total Page:16
File Type:pdf, Size:1020Kb

Load more
Recommended publications
-
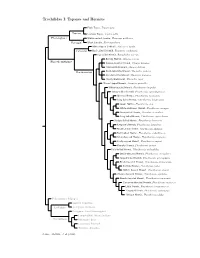
Topazes and Hermits
Trochilidae I: Topazes and Hermits Fiery Topaz, Topaza pyra Topazini Crimson Topaz, Topaza pella Florisuginae White-necked Jacobin, Florisuga mellivora Florisugini Black Jacobin, Florisuga fusca White-tipped Sicklebill, Eutoxeres aquila Eutoxerini Buff-tailed Sicklebill, Eutoxeres condamini Saw-billed Hermit, Ramphodon naevius Bronzy Hermit, Glaucis aeneus Phaethornithinae Rufous-breasted Hermit, Glaucis hirsutus ?Hook-billed Hermit, Glaucis dohrnii Threnetes ruckeri Phaethornithini Band-tailed Barbthroat, Pale-tailed Barbthroat, Threnetes leucurus ?Sooty Barbthroat, Threnetes niger ?Broad-tipped Hermit, Anopetia gounellei White-bearded Hermit, Phaethornis hispidus Tawny-bellied Hermit, Phaethornis syrmatophorus Mexican Hermit, Phaethornis mexicanus Long-billed Hermit, Phaethornis longirostris Green Hermit, Phaethornis guy White-whiskered Hermit, Phaethornis yaruqui Great-billed Hermit, Phaethornis malaris Long-tailed Hermit, Phaethornis superciliosus Straight-billed Hermit, Phaethornis bourcieri Koepcke’s Hermit, Phaethornis koepckeae Needle-billed Hermit, Phaethornis philippii Buff-bellied Hermit, Phaethornis subochraceus Scale-throated Hermit, Phaethornis eurynome Sooty-capped Hermit, Phaethornis augusti Planalto Hermit, Phaethornis pretrei Pale-bellied Hermit, Phaethornis anthophilus Stripe-throated Hermit, Phaethornis striigularis Gray-chinned Hermit, Phaethornis griseogularis Black-throated Hermit, Phaethornis atrimentalis Reddish Hermit, Phaethornis ruber ?White-browed Hermit, Phaethornis stuarti ?Dusky-throated Hermit, Phaethornis squalidus Streak-throated Hermit, Phaethornis rupurumii Cinnamon-throated Hermit, Phaethornis nattereri Little Hermit, Phaethornis longuemareus ?Tapajos Hermit, Phaethornis aethopygus ?Minute Hermit, Phaethornis idaliae Polytminae: Mangos Lesbiini: Coquettes Lesbiinae Coeligenini: Brilliants Patagonini: Giant Hummingbird Lampornithini: Mountain-Gems Tro chilinae Mellisugini: Bees Cynanthini: Emeralds Trochilini: Amazilias Source: McGuire et al. (2014).. -

A Comprehensive Multilocus Phylogeny of the Neotropical Cotingas
Molecular Phylogenetics and Evolution 81 (2014) 120–136 Contents lists available at ScienceDirect Molecular Phylogenetics and Evolution journal homepage: www.elsevier.com/locate/ympev A comprehensive multilocus phylogeny of the Neotropical cotingas (Cotingidae, Aves) with a comparative evolutionary analysis of breeding system and plumage dimorphism and a revised phylogenetic classification ⇑ Jacob S. Berv 1, Richard O. Prum Department of Ecology and Evolutionary Biology and Peabody Museum of Natural History, Yale University, P.O. Box 208105, New Haven, CT 06520, USA article info abstract Article history: The Neotropical cotingas (Cotingidae: Aves) are a group of passerine birds that are characterized by Received 18 April 2014 extreme diversity in morphology, ecology, breeding system, and behavior. Here, we present a compre- Revised 24 July 2014 hensive phylogeny of the Neotropical cotingas based on six nuclear and mitochondrial loci (7500 bp) Accepted 6 September 2014 for a sample of 61 cotinga species in all 25 genera, and 22 species of suboscine outgroups. Our taxon sam- Available online 16 September 2014 ple more than doubles the number of cotinga species studied in previous analyses, and allows us to test the monophyly of the cotingas as well as their intrageneric relationships with high resolution. We ana- Keywords: lyze our genetic data using a Bayesian species tree method, and concatenated Bayesian and maximum Phylogenetics likelihood methods, and present a highly supported phylogenetic hypothesis. We confirm the monophyly Bayesian inference Species-tree of the cotingas, and present the first phylogenetic evidence for the relationships of Phibalura flavirostris as Sexual selection the sister group to Ampelion and Doliornis, and the paraphyly of Lipaugus with respect to Tijuca. -

Colombia Mega II 1St – 30Th November 2016 (30 Days) Trip Report
Colombia Mega II 1st – 30th November 2016 (30 Days) Trip Report Black Manakin by Trevor Ellery Trip Report compiled by tour leader: Trevor Ellery Trip Report – RBL Colombia - Mega II 2016 2 ___________________________________________________________________________________ Top ten birds of the trip as voted for by the Participants: 1. Ocellated Tapaculo 6. Blue-and-yellow Macaw 2. Rainbow-bearded Thornbill 7. Red-ruffed Fruitcrow 3. Multicolored Tanager 8. Sungrebe 4. Fiery Topaz 9. Buffy Helmetcrest 5. Sword-billed Hummingbird 10. White-capped Dipper Tour Summary This was one again a fantastic trip across the length and breadth of the world’s birdiest nation. Highlights were many and included everything from the flashy Fiery Topazes and Guianan Cock-of- the-Rocks of the Mitu lowlands to the spectacular Rainbow-bearded Thornbills and Buffy Helmetcrests of the windswept highlands. In between, we visited just about every type of habitat that it is possible to bird in Colombia and shared many special moments: the diminutive Lanceolated Monklet that perched above us as we sheltered from the rain at the Piha Reserve, the showy Ochre-breasted Antpitta we stumbled across at an antswarm at Las Tangaras Reserve, the Ocellated Tapaculo (voted bird of the trip) that paraded in front of us at Rio Blanco, and the male Vermilion Cardinal, in all his crimson glory, that we enjoyed in the Guajira desert on the final morning of the trip. If you like seeing lots of birds, lots of specialities, lots of endemics and enjoy birding in some of the most stunning scenery on earth, then this trip is pretty unbeatable. -

Reference File
References added since publication of 2007 CRC Handbook of Avian Body Masses Abadie, K. B., J. Pérez Z., and M. Valverde. 2006. Primer reporte de colonias del Martín Peruano Progne murphyi. Cotinga 24:99-101. Ackerman, J. T., J. Y. Takekawa, J. D. Bluso, J. L. Yee, and C. A. Eagles-Smith. 2008. Gender identification of Caspian Terns using external morphology and discriminant function analysis. Wilson Journal of Ornithology 120:378-383. Alarcos, S., C. de la Cruz, E. Solís, J. Valencia, and M. J. García-Baquero. 2007. Sex determination of Iberian Azure-winged Magpies Cyanopica cyanus cooki by discriminant analysis of external measurements. Ringing & Migration 23:211-216. Albayrak, T., A. Besnard, and A. Erdoğan. 2011. Morphometric variation and population relationships of Krüeper’s Nuthatch (Sitta krueperi) in Turkey. Wilson Journal of Ornithology 123:734-740. Aleixo, A., C. E. B. Portes, A. Whittaker, J. D. Weckstein, L. Pedreira Gonzaga, K. J. Zimmer, C. C. Ribas, and J. M. Bates. 2013. Molecular systematics and taxonomic revision of the Curve-billed Scythebill complex (Campylorhamphus procurvoides: Dendrocolaptidae), with description of a new species from western Amazonian Brazil. Pp. 253-257, In: del Hoyo, J., A Elliott, J. Sargatal, and D.A. Christie (eds). Handbook of the birds of the world. Special volume: new species and global index. Lynx Edicions, Barcelona, Spain. Volume 1. Alfano, A. 2014. Pygmy Nightjar (Nyctopolus hirundinaeus). Neotropical Birds Online (T.S. Schulenberg, ed.). Cornell Laboratory of Ornithology, Ithaca, NY. Alvarenga, H. M. F., E. Höfling, and L. F. Silveira. 2002. Notharchus swainsoni (Gray, 1846) é uma espécie válida. -

The Best of Costa Rica March 19–31, 2019
THE BEST OF COSTA RICA MARCH 19–31, 2019 Buffy-crowned Wood-Partridge © David Ascanio LEADERS: DAVID ASCANIO & MAURICIO CHINCHILLA LIST COMPILED BY: DAVID ASCANIO VICTOR EMANUEL NATURE TOURS, INC. 2525 WALLINGWOOD DRIVE, SUITE 1003 AUSTIN, TEXAS 78746 WWW.VENTBIRD.COM THE BEST OF COSTA RICA March 19–31, 2019 By David Ascanio Photo album: https://www.flickr.com/photos/davidascanio/albums/72157706650233041 It’s about 02:00 AM in San José, and we are listening to the widespread and ubiquitous Clay-colored Robin singing outside our hotel windows. Yet, it was still too early to experience the real explosion of bird song, which usually happens after dawn. Then, after 05:30 AM, the chorus started when a vocal Great Kiskadee broke the morning silence, followed by the scratchy notes of two Hoffmann´s Woodpeckers, a nesting pair of Inca Doves, the ascending and monotonous song of the Yellow-bellied Elaenia, and the cacophony of an (apparently!) engaged pair of Rufous-naped Wrens. This was indeed a warm welcome to magical Costa Rica! To complement the first morning of birding, two boreal migrants, Baltimore Orioles and a Tennessee Warbler, joined the bird feast just outside the hotel area. Broad-billed Motmot . Photo: D. Ascanio © Victor Emanuel Nature Tours 2 The Best of Costa Rica, 2019 After breakfast, we drove towards the volcanic ring of Costa Rica. Circling the slope of Poas volcano, we eventually reached the inspiring Bosque de Paz. With its hummingbird feeders and trails transecting a beautiful moss-covered forest, this lodge offered us the opportunity to see one of Costa Rica´s most difficult-to-see Grallaridae, the Scaled Antpitta. -

Volume 2. Animals
AC20 Doc. 8.5 Annex (English only/Seulement en anglais/Únicamente en inglés) REVIEW OF SIGNIFICANT TRADE ANALYSIS OF TRADE TRENDS WITH NOTES ON THE CONSERVATION STATUS OF SELECTED SPECIES Volume 2. Animals Prepared for the CITES Animals Committee, CITES Secretariat by the United Nations Environment Programme World Conservation Monitoring Centre JANUARY 2004 AC20 Doc. 8.5 – p. 3 Prepared and produced by: UNEP World Conservation Monitoring Centre, Cambridge, UK UNEP WORLD CONSERVATION MONITORING CENTRE (UNEP-WCMC) www.unep-wcmc.org The UNEP World Conservation Monitoring Centre is the biodiversity assessment and policy implementation arm of the United Nations Environment Programme, the world’s foremost intergovernmental environmental organisation. UNEP-WCMC aims to help decision-makers recognise the value of biodiversity to people everywhere, and to apply this knowledge to all that they do. The Centre’s challenge is to transform complex data into policy-relevant information, to build tools and systems for analysis and integration, and to support the needs of nations and the international community as they engage in joint programmes of action. UNEP-WCMC provides objective, scientifically rigorous products and services that include ecosystem assessments, support for implementation of environmental agreements, regional and global biodiversity information, research on threats and impacts, and development of future scenarios for the living world. Prepared for: The CITES Secretariat, Geneva A contribution to UNEP - The United Nations Environment Programme Printed by: UNEP World Conservation Monitoring Centre 219 Huntingdon Road, Cambridge CB3 0DL, UK © Copyright: UNEP World Conservation Monitoring Centre/CITES Secretariat The contents of this report do not necessarily reflect the views or policies of UNEP or contributory organisations. -

Peru: from the Cusco Andes to the Manu
The critically endangered Royal Cinclodes - our bird-of-the-trip (all photos taken on this tour by Pete Morris) PERU: FROM THE CUSCO ANDES TO THE MANU 26 JULY – 12 AUGUST 2017 LEADERS: PETE MORRIS and GUNNAR ENGBLOM This brand new itinerary really was a tour of two halves! For the frst half of the tour we really were up on the roof of the world, exploring the Andes that surround Cusco up to altitudes in excess of 4000m. Cold clear air and fantastic snow-clad peaks were the order of the day here as we went about our task of seeking out a number of scarce, localized and seldom-seen endemics. For the second half of the tour we plunged down off of the mountains and took the long snaking Manu Road, right down to the Amazon basin. Here we traded the mountainous peaks for vistas of forest that stretched as far as the eye could see in one of the planet’s most diverse regions. Here, the temperatures rose in line with our ever growing list of sightings! In all, we amassed a grand total of 537 species of birds, including 36 which provided audio encounters only! As we all know though, it’s not necessarily the shear number of species that counts, but more the quality, and we found many high quality species. New species for the Birdquest life list included Apurimac Spinetail, Vilcabamba Thistletail, Am- pay (still to be described) and Vilcabamba Tapaculos and Apurimac Brushfnch, whilst other montane goodies included the stunning Bearded Mountaineer, White-tufted Sunbeam the critically endangered Royal Cinclodes, 1 BirdQuest Tour Report: Peru: From the Cusco Andes to The Manu 2017 www.birdquest-tours.com These wonderful Blue-headed Macaws were a brilliant highlight near to Atalaya. -

An Update of Wallacels Zoogeographic Regions of the World
REPORTS To examine the temporal profile of ChC produc- specification of a distinct, and probably the last, 3. G. A. Ascoli et al., Nat. Rev. Neurosci. 9, 557 (2008). tion and their correlation to laminar deployment, cohort in this lineage—the ChCs. 4. J. Szentágothai, M. A. Arbib, Neurosci. Res. Program Bull. 12, 305 (1974). we injected a single pulse of BrdU into pregnant A recent study demonstrated that progeni- CreER 5. P. Somogyi, Brain Res. 136, 345 (1977). Nkx2.1 ;Ai9 females at successive days be- tors below the ventral wall of the lateral ventricle 6. L. Sussel, O. Marin, S. Kimura, J. L. Rubenstein, tween E15 and P1 to label mitotic progenitors, (i.e., VGZ) of human infants give rise to a medial Development 126, 3359 (1999). each paired with a pulse of tamoxifen at E17 to migratory stream destined to the ventral mPFC 7. S. J. Butt et al., Neuron 59, 722 (2008). + 18 8. H. Taniguchi et al., Neuron 71, 995 (2011). label NKX2.1 cells (Fig. 3A). We first quanti- ( ). Despite species differences in the develop- 9. L. Madisen et al., Nat. Neurosci. 13, 133 (2010). fied the fraction of L2 ChCs (identified by mor- mental timing of corticogenesis, this study and 10. J. Szabadics et al., Science 311, 233 (2006). + phology) in mPFC that were also BrdU+. Although our findings raise the possibility that the NKX2.1 11. A. Woodruff, Q. Xu, S. A. Anderson, R. Yuste, Front. there was ChC production by E15, consistent progenitors in VGZ and their extended neurogenesis Neural Circuits 3, 15 (2009). -

Journal of Avian Biology JAV-00869 Wang, N
Journal of Avian Biology JAV-00869 Wang, N. and Kimball, R. T. 2016. Re-evaluating the distribution of cooperative breeding in birds: is it tightly linked with altriciality? – J. Avian Biol. doi: 10.1111/jav.00869 Supplementary material Appendix 1. Table A1. The characteristics of the 9993 species based on Jetz et al. (2012) Order Species Criteria1 Developmental K K+S K+S+I LB Mode ACCIPITRIFORMES Accipiter albogularis 0 0 0 0 1 ACCIPITRIFORMES Accipiter badius 0 0 0 0 1 ACCIPITRIFORMES Accipiter bicolor 0 0 0 0 1 ACCIPITRIFORMES Accipiter brachyurus 0 0 0 0 1 ACCIPITRIFORMES Accipiter brevipes 0 0 0 0 1 ACCIPITRIFORMES Accipiter butleri 0 0 0 0 1 ACCIPITRIFORMES Accipiter castanilius 0 0 0 0 1 ACCIPITRIFORMES Accipiter chilensis 0 0 0 0 1 ACCIPITRIFORMES Accipiter chionogaster 0 0 0 0 1 ACCIPITRIFORMES Accipiter cirrocephalus 0 0 0 0 1 ACCIPITRIFORMES Accipiter collaris 0 0 0 0 1 ACCIPITRIFORMES Accipiter cooperii 0 0 0 0 1 ACCIPITRIFORMES Accipiter erythrauchen 0 0 0 0 1 ACCIPITRIFORMES Accipiter erythronemius 0 0 0 0 1 ACCIPITRIFORMES Accipiter erythropus 0 0 0 0 1 ACCIPITRIFORMES Accipiter fasciatus 0 0 0 0 1 ACCIPITRIFORMES Accipiter francesiae 0 0 0 0 1 ACCIPITRIFORMES Accipiter gentilis 0 0 0 0 1 ACCIPITRIFORMES Accipiter griseiceps 0 0 0 0 1 ACCIPITRIFORMES Accipiter gularis 0 0 0 0 1 ACCIPITRIFORMES Accipiter gundlachi 0 0 0 0 1 ACCIPITRIFORMES Accipiter haplochrous 0 0 0 0 1 ACCIPITRIFORMES Accipiter henicogrammus 0 0 0 0 1 ACCIPITRIFORMES Accipiter henstii 0 0 0 0 1 ACCIPITRIFORMES Accipiter imitator 0 0 0 0 1 ACCIPITRIFORMES -

The Effects of Ecology and Evolution on Avian Flight Morphology Natalie Wright
University of New Mexico UNM Digital Repository Biology ETDs Electronic Theses and Dissertations 7-1-2015 The effects of ecology and evolution on avian flight morphology Natalie Wright Follow this and additional works at: https://digitalrepository.unm.edu/biol_etds Recommended Citation Wright, Natalie. "The effects of ecology and evolution on avian flight morphology." (2015). https://digitalrepository.unm.edu/ biol_etds/116 This Dissertation is brought to you for free and open access by the Electronic Theses and Dissertations at UNM Digital Repository. It has been accepted for inclusion in Biology ETDs by an authorized administrator of UNM Digital Repository. For more information, please contact [email protected]. Natalie A. Wright Candidate Biology Department This dissertation is approved, and it is acceptable in quality and form for publication: Approved by the Dissertation Committee: Christopher C. Witt, Chairperson Jeffery Long Blair Wolf James Brown i THE EFFECTS OF ECOLOGY AND EVOLUTION ON AVIAN FLIGHT MORPHOLOGY by NATALIE A. WRIGHT B.S., Zoology, University of Florida, 2005 M.S., Zoology, University of Florida, 2009 DISSERTATION Submitted in Partial Fulfillment of the Requirements for the Degree of Doctor of Philosophy Biology The University of New Mexico Albuquerque, New Mexico August, 2015 ii Dedication This dissertation is dedicated to Bob Dickerman, whose enthusiastic contributions to ornithology will continue to influence and aid many new generations of biologists. He collected a great many of the specimens used in this manuscript. Without his generous support of the Museum of Southwestern Biology ornithology collection, this dissertation would not have been possible. He is greatly missed and remembered fondly. iii Acknowledgements I thank my advisor, Chris Witt, for his support, encouragement, and guidance over the past six years. -

Alpha Codes for 2168 Bird Species (And 113 Non-Species Taxa) in Accordance with the 62Nd AOU Supplement (2021), Sorted Taxonomically
Four-letter (English Name) and Six-letter (Scientific Name) Alpha Codes for 2168 Bird Species (and 113 Non-Species Taxa) in accordance with the 62nd AOU Supplement (2021), sorted taxonomically Prepared by Peter Pyle and David F. DeSante The Institute for Bird Populations www.birdpop.org ENGLISH NAME 4-LETTER CODE SCIENTIFIC NAME 6-LETTER CODE Highland Tinamou HITI Nothocercus bonapartei NOTBON Great Tinamou GRTI Tinamus major TINMAJ Little Tinamou LITI Crypturellus soui CRYSOU Thicket Tinamou THTI Crypturellus cinnamomeus CRYCIN Slaty-breasted Tinamou SBTI Crypturellus boucardi CRYBOU Choco Tinamou CHTI Crypturellus kerriae CRYKER White-faced Whistling-Duck WFWD Dendrocygna viduata DENVID Black-bellied Whistling-Duck BBWD Dendrocygna autumnalis DENAUT West Indian Whistling-Duck WIWD Dendrocygna arborea DENARB Fulvous Whistling-Duck FUWD Dendrocygna bicolor DENBIC Emperor Goose EMGO Anser canagicus ANSCAN Snow Goose SNGO Anser caerulescens ANSCAE + Lesser Snow Goose White-morph LSGW Anser caerulescens caerulescens ANSCCA + Lesser Snow Goose Intermediate-morph LSGI Anser caerulescens caerulescens ANSCCA + Lesser Snow Goose Blue-morph LSGB Anser caerulescens caerulescens ANSCCA + Greater Snow Goose White-morph GSGW Anser caerulescens atlantica ANSCAT + Greater Snow Goose Intermediate-morph GSGI Anser caerulescens atlantica ANSCAT + Greater Snow Goose Blue-morph GSGB Anser caerulescens atlantica ANSCAT + Snow X Ross's Goose Hybrid SRGH Anser caerulescens x rossii ANSCAR + Snow/Ross's Goose SRGO Anser caerulescens/rossii ANSCRO Ross's Goose -

Use of Floral Nectar in Heliconia Stilesii Daniels by Three Species of Hermit Hummingbirds
The Condor89~779-787 0 The Cooper OrnithologicalSociety 1987 ECOLOGICAL FITTING: USE OF FLORAL NECTAR IN HELICONIA STILESII DANIELS BY THREE SPECIES OF HERMIT HUMMINGBIRDS FRANK B. GILL The Academyof Natural Sciences,Philadelphia, PA 19103 Abstract. Three speciesof hermit hummingbirds-a specialist(Eutoxeres aquikz), a gen- eralist (Phaethornissuperciliosus), and a thief (Threnetesruckerz]-visited the nectar-rich flowers of Heliconia stilesii Daniels at a lowland study site on the Osa Peninsula of Costa Rica. Unlike H. pogonanthaCufodontis, a related Caribbean lowland specieswith a less specialized flower, H. stilesii may not realize its full reproductive potential at this site, becauseit cannot retain the services of alternative pollinators such as Phaethornis.The flowers of H. stilesii appear adapted for pollination by Eutoxeres, but this hummingbird rarely visited them at this site. Lek male Phaethornisvisited the flowers frequently in late May and early June, but then abandonedthis nectar sourcein favor of other flowers offering more accessiblenectar. The strong curvature of the perianth prevents accessby Phaethornis to the main nectar chamber; instead they obtain only small amounts of nectar that leaks anteriorly into the belly of the flower. Key words: Hummingbird; pollination; mutualism;foraging; Heliconia stilesii; nectar. INTRODUCTION Ultimately affectedare the hummingbird’s choice Species that expand their distribution following of flowers and patterns of competition among speciationenter novel ecologicalassociations un- hummingbird species for nectar (Stiles 1975, related to previous evolutionary history and face 1978; Wolf et al. 1976; Feinsinger 1978; Gill the challenges of adjustment to new settings, 1978). Use of specificHeliconia flowers as sources called “ecological fitting” (Janzen 1985a). In the of nectar by particular species of hermit hum- case of mutualistic species, such as plants and mingbirds, however, varies seasonally and geo- their pollinators, new ecological settingsmay in- graphically (Stiles 1975).