Intensity and Duration of TCR Signaling Is Limited by P38 Phosphorylation of ZAP-70T293 and Destabilization of the Signalosome
Total Page:16
File Type:pdf, Size:1020Kb

Load more
Recommended publications
-

Promoter Janus Kinase 3 Proximal Characterization and Analysis Of
The Journal of Immunology Characterization and Analysis of the Proximal Janus Kinase 3 Promoter1 Martin Aringer,2*† Sigrun R. Hofmann,2* David M. Frucht,* Min Chen,* Michael Centola,* Akio Morinobu,* Roberta Visconti,* Daniel L. Kastner,* Josef S. Smolen,† and John J. O’Shea3* Janus kinase 3 (Jak3) is a nonreceptor tyrosine kinase essential for signaling via cytokine receptors that comprise the common ␥-chain (␥c), i.e., the receptors for IL-2, IL-4, IL-7, IL-9, IL-15, and IL-21. Jak3 is preferentially expressed in hemopoietic cells and is up-regulated upon cell differentiation and activation. Despite the importance of Jak3 in lymphoid development and immune function, the mechanisms that govern its expression have not been defined. To gain insight into this issue, we set out to characterize the Jak3 promoter. The 5-untranslated region of the Jak3 gene is interrupted by a 3515-bp intron. Upstream of this intron and the transcription initiation site, we identified an ϳ1-kb segment that exhibited lymphoid-specific promoter activity and was responsive to TCR signals. Truncation of this fragment revealed that core promoter activity resided in a 267-bp fragment that contains putative Sp-1, AP-1, Ets, Stat, and other binding sites. Mutation of the AP-1 sites significantly diminished, whereas mutation of the Ets sites abolished, the inducibility of the promoter construct. Chromatin immunoprecipitation assays showed that histone acetylation correlates with mRNA expression and that Ets-1/2 binds this region. Thus, transcription factors that bind these sites, especially Ets family members, are likely to be important regulators of Jak3 expression. -

Regulation of Calcium/Calmodulin-Dependent Protein Kinase II Activation by Intramolecular and Intermolecular Interactions
8394 • The Journal of Neuroscience, September 29, 2004 • 24(39):8394–8398 Mini-Review Regulation of Calcium/Calmodulin-Dependent Protein Kinase II Activation by Intramolecular and Intermolecular Interactions Leslie C. Griffith Department of Biology and Volen Center for Complex Systems, Brandeis University, Waltham, Massachusetts 02454-9110 Key words: calcium; calmodulin; learning; localization; NMDA; phosphatase; protein kinase As its name implies, calcium/calmodulin-dependent protein ki- The overlap of these subdomains is no accident. Binding of nase II (CaMKII) is calcium dependent. In its basal state, the Ca 2ϩ/CaM is the primary signal for release of autoinhibition. activity of CaMKII is extremely low. Regulation of intracellular Current models of activation posit that the binding of Ca 2ϩ/CaM calcium levels allows the neuron to link activity with phosphor- serves to disrupt the interactions of specific residues within the ylation by CaMKII. This review will briefly summarize our cur- autoinhibitory domain with the catalytic domain (Smith et al., rent understanding of the intramolecular mechanisms of activity 1992). Because there is no crystal structure for the catalytic and regulation and their modulation by Ca 2ϩ/CaM and will then regulatory parts of CaMKII, the interaction face of these two focus on the growing number of other modes of intermolecular domains has been inferred using the effects of charge-reversal regulation of CaMKII activity by substrate and scaffolding mutagenesis on activity and molecular modeling (Yang and molecules. Schulman, 1999). This study confirmed the role of Arg 297 at the P-3 position of the pseudosubstrate ligand (Mukherji and Soder- Regulation of CaMKII by its autoinhibitory domain ling, 1995) and identified residues in the catalytic domain that All members of the CaMKII family (␣, , ␥, and ␦ isozymes) share may have direct interactions with the regulatory region. -

REVIEW Signal Transduction, Cell Cycle Regulatory, and Anti
Leukemia (1999) 13, 1109–1166 1999 Stockton Press All rights reserved 0887-6924/99 $12.00 http://www.stockton-press.co.uk/leu REVIEW Signal transduction, cell cycle regulatory, and anti-apoptotic pathways regulated by IL-3 in hematopoietic cells: possible sites for intervention with anti-neoplastic drugs WL Blalock1, C Weinstein-Oppenheimer1,2, F Chang1, PE Hoyle1, X-Y Wang3, PA Algate4, RA Franklin1,5, SM Oberhaus1,5, LS Steelman1 and JA McCubrey1,5 1Department of Microbiology and Immunology, 5Leo Jenkins Cancer Center, East Carolina University School of Medicine Greenville, NC, USA; 2Escuela de Quı´mica y Farmacia, Facultad de Medicina, Universidad de Valparaiso, Valparaiso, Chile; 3Department of Laboratory Medicine and Pathology, Mayo Clinic and Foundation, Rochester, MN, USA; and 4Division of Basic Sciences, Fred Hutchinson Cancer Research Center, Seattle, WA, USA Over the past decade, there has been an exponential increase growth factor), Flt-L (the ligand for the flt2/3 receptor), erythro- in our knowledge of how cytokines regulate signal transduc- poietin (EPO), and others affect the growth and differentiation tion, cell cycle progression, differentiation and apoptosis. Research has focused on different biochemical and genetic of these early hematopoietic precursor cells into cells of the 1–4 aspects of these processes. Initially, cytokines were identified myeloid, lymphoid and erythroid lineages (Table 1). This by clonogenic assays and purified by biochemical techniques. review will concentrate on IL-3 since much of the knowledge This soon led to the molecular cloning of the genes encoding of how cytokines affect cell growth, signal transduction, and the cytokines and their cognate receptors. -

Human and Mouse CD Marker Handbook Human and Mouse CD Marker Key Markers - Human Key Markers - Mouse
Welcome to More Choice CD Marker Handbook For more information, please visit: Human bdbiosciences.com/eu/go/humancdmarkers Mouse bdbiosciences.com/eu/go/mousecdmarkers Human and Mouse CD Marker Handbook Human and Mouse CD Marker Key Markers - Human Key Markers - Mouse CD3 CD3 CD (cluster of differentiation) molecules are cell surface markers T Cell CD4 CD4 useful for the identification and characterization of leukocytes. The CD CD8 CD8 nomenclature was developed and is maintained through the HLDA (Human Leukocyte Differentiation Antigens) workshop started in 1982. CD45R/B220 CD19 CD19 The goal is to provide standardization of monoclonal antibodies to B Cell CD20 CD22 (B cell activation marker) human antigens across laboratories. To characterize or “workshop” the antibodies, multiple laboratories carry out blind analyses of antibodies. These results independently validate antibody specificity. CD11c CD11c Dendritic Cell CD123 CD123 While the CD nomenclature has been developed for use with human antigens, it is applied to corresponding mouse antigens as well as antigens from other species. However, the mouse and other species NK Cell CD56 CD335 (NKp46) antibodies are not tested by HLDA. Human CD markers were reviewed by the HLDA. New CD markers Stem Cell/ CD34 CD34 were established at the HLDA9 meeting held in Barcelona in 2010. For Precursor hematopoetic stem cell only hematopoetic stem cell only additional information and CD markers please visit www.hcdm.org. Macrophage/ CD14 CD11b/ Mac-1 Monocyte CD33 Ly-71 (F4/80) CD66b Granulocyte CD66b Gr-1/Ly6G Ly6C CD41 CD41 CD61 (Integrin b3) CD61 Platelet CD9 CD62 CD62P (activated platelets) CD235a CD235a Erythrocyte Ter-119 CD146 MECA-32 CD106 CD146 Endothelial Cell CD31 CD62E (activated endothelial cells) Epithelial Cell CD236 CD326 (EPCAM1) For Research Use Only. -

Circulating Supar in Two Cohorts of Primary FSGS
CLINICAL RESEARCH www.jasn.org Circulating suPAR in Two Cohorts of Primary FSGS † ‡ Changli Wei,* Howard Trachtman, Jing Li,* Chuanhui Dong, Aaron L. Friedman,§ | | | Jennifer J. Gassman, June L. McMahan, Milena Radeva, Karsten M. Heil,¶ †† ‡‡ Agnes Trautmann,¶ Ali Anarat,** Sevinc Emre, Gian M. Ghiggeri, Fatih Ozaltin,§§ || ††† Dieter Haffner, Debbie S. Gipson,¶¶ Frederick Kaskel,*** Dagmar-Christiane Fischer, ‡‡‡ Franz Schaefer,¶ and Jochen Reiser, for the PodoNet and FSGS CT Study Consortia Departments of *Medicine and ‡Neurology, University of Miami Miller School of Medicine, Miami, Florida; †Department of Pediatrics, NYU Langone Medical Center, New York, New York; §Department of Pediatrics, University of Minnesota, Minneapolis, Minnesota; |Department of Quantitative Health Sciences, Cleveland Clinic, Cleveland, Ohio; ¶Center for Pediatric and Adolescent Medicine, University of Heidelberg, Heidelberg, Germany; **Department of Pediatric Nephrology, Cukurova University School of Medicine, Adana, Turkey; ††Department of Pediatrics, Istanbul Medical Faculty, University of Istanbul, Istanbul, Turkey; ‡‡Division of Nephrology, Dialysis, and Transplantation, Laboratory on Pathophysiology of Uremia, G. Gaslini Children’s Hospital, Genoa, Italy; §§Pediatric Nephrology Unit, Department of Pediatrics, Faculty of Medicine, Hacettepe University, Ankara, Turkey; ||Department of Pediatric Kidney, Liver, and Metabolic Diseases, Hannover Medical School, Hannover, Germany; ¶¶Department of Pediatrics, University of Michigan, Ann Arbor, Michigan; ***Children’s Hospital at Montefiore, Albert Einstein College of Medicine, Bronx, New York; †††Department of Pediatrics, Rostock University Hospital, Rostock, Germany; and ‡‡‡Department of Medicine, Rush University Medical Center, Chicago, Illinois ABSTRACT Overexpression of soluble urokinase receptor (suPAR) causes pathology in animal models similar to pri- mary FSGS, and one recent study demonstrated elevated levels of serum suPAR in patients with the disease. -

Hras Intracellular Trafficking and Signal Transduction Jodi Ho-Jung Mckay Iowa State University
Iowa State University Capstones, Theses and Retrospective Theses and Dissertations Dissertations 2007 HRas intracellular trafficking and signal transduction Jodi Ho-Jung McKay Iowa State University Follow this and additional works at: https://lib.dr.iastate.edu/rtd Part of the Biological Phenomena, Cell Phenomena, and Immunity Commons, Cancer Biology Commons, Cell Biology Commons, Genetics and Genomics Commons, and the Medical Cell Biology Commons Recommended Citation McKay, Jodi Ho-Jung, "HRas intracellular trafficking and signal transduction" (2007). Retrospective Theses and Dissertations. 13946. https://lib.dr.iastate.edu/rtd/13946 This Dissertation is brought to you for free and open access by the Iowa State University Capstones, Theses and Dissertations at Iowa State University Digital Repository. It has been accepted for inclusion in Retrospective Theses and Dissertations by an authorized administrator of Iowa State University Digital Repository. For more information, please contact [email protected]. HRas intracellular trafficking and signal transduction by Jodi Ho-Jung McKay A dissertation submitted to the graduate faculty in partial fulfillment of the requirements for the degree of DOCTOR OF PHILOSOPHY Major: Genetics Program of Study Committee: Janice E. Buss, Co-major Professor Linda Ambrosio, Co-major Professor Diane Bassham Drena Dobbs Ted Huiatt Iowa State University Ames, Iowa 2007 Copyright © Jodi Ho-Jung McKay, 2007. All rights reserved. UMI Number: 3274881 Copyright 2007 by McKay, Jodi Ho-Jung All rights reserved. UMI Microform 3274881 Copyright 2008 by ProQuest Information and Learning Company. All rights reserved. This microform edition is protected against unauthorized copying under Title 17, United States Code. ProQuest Information and Learning Company 300 North Zeeb Road P.O. -

Regulation of Calmodulin-Stimulated Cyclic Nucleotide Phosphodiesterase (PDE1): Review
95-105 5/6/06 13:44 Page 95 INTERNATIONAL JOURNAL OF MOLECULAR MEDICINE 18: 95-105, 2006 95 Regulation of calmodulin-stimulated cyclic nucleotide phosphodiesterase (PDE1): Review RAJENDRA K. SHARMA, SHANKAR B. DAS, ASHAKUMARY LAKSHMIKUTTYAMMA, PONNIAH SELVAKUMAR and ANURAAG SHRIVASTAV Department of Pathology and Laboratory Medicine, College of Medicine, University of Saskatchewan, Cancer Research Division, Saskatchewan Cancer Agency, 20 Campus Drive, Saskatoon SK S7N 4H4, Canada Received January 16, 2006; Accepted March 13, 2006 Abstract. The response of living cells to change in cell 6. Differential inhibition of PDE1 isozymes and its environment depends on the action of second messenger therapeutic applications molecules. The two second messenger molecules cAMP and 7. Role of proteolysis in regulating PDE1A2 Ca2+ regulate a large number of eukaryotic cellular events. 8. Role of PDE1A1 in ischemic-reperfused heart Calmodulin-stimulated cyclic nucleotide phosphodiesterase 9. Conclusion (PDE1) is one of the key enzymes involved in the complex interaction between cAMP and Ca2+ second messenger systems. Some PDE1 isozymes have similar kinetic and 1. Introduction immunological properties but are differentially regulated by Ca2+ and calmodulin. Accumulating evidence suggests that the A variety of cellular activities are regulated through mech- activity of PDE1 is selectively regulated by cross-talk between anisms controlling the level of cyclic nucleotides. These Ca2+ and cAMP signalling pathways. These isozymes are mechanisms include synthesis, degradation, efflux and seque- also further distinguished by various pharmacological agents. stration of cyclic adenosine 3':5'-monophosphate (cAMP) and We have demonstrated a potentially novel regulation of PDE1 cyclic guanosine 3':5'- monophosphate (cGMP) within the by calpain. -

G-Protein ␥-Complex Is Crucial for Efficient Signal Amplification in Vision
The Journal of Neuroscience, June 1, 2011 • 31(22):8067–8077 • 8067 Cellular/Molecular G-Protein ␥-Complex Is Crucial for Efficient Signal Amplification in Vision Alexander V. Kolesnikov,1 Loryn Rikimaru,2 Anne K. Hennig,1 Peter D. Lukasiewicz,1 Steven J. Fliesler,4,5,6,7 Victor I. Govardovskii,8 Vladimir J. Kefalov,1 and Oleg G. Kisselev2,3 1Department of Ophthalmology and Visual Sciences, Washington University School of Medicine, St. Louis, Missouri 63110, Departments of 2Ophthalmology and 3Biochemistry and Molecular Biology, Saint Louis University School of Medicine, Saint Louis, Missouri 63104, 4Research Service, Veterans Administration Western New York Healthcare System, and Departments of 5Ophthalmology (Ross Eye Institute) and 6Biochemistry, University at Buffalo/The State University of New York (SUNY), and 7SUNY Eye Institute, Buffalo, New York 14215, and 8Sechenov Institute for Evolutionary Physiology and Biochemistry, Russian Academy of Sciences, Saint Petersburg 194223, Russia A fundamental question of cell signaling biology is how faint external signals produce robust physiological responses. One universal mechanism relies on signal amplification via intracellular cascades mediated by heterotrimeric G-proteins. This high amplification system allows retinal rod photoreceptors to detect single photons of light. Although much is now known about the role of the ␣-subunit of the rod-specific G-protein transducin in phototransduction, the physiological function of the auxiliary ␥-complex in this process remains a mystery. Here, we show that elimination of the transducin ␥-subunit drastically reduces signal amplification in intact mouse rods. The consequence is a striking decline in rod visual sensitivity and severe impairment of nocturnal vision. Our findings demonstrate that transducin ␥-complex controls signal amplification of the rod phototransduction cascade and is critical for the ability of rod photoreceptors to function in low light conditions. -
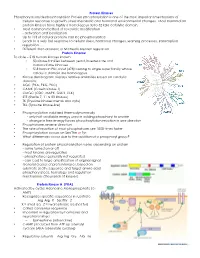
Protein Kinases Phosphorylation/Dephosphorylation Protein Phosphorylation Is One of the Most Important Mechanisms of Cellular Re
Protein Kinases Phosphorylation/dephosphorylation Protein phosphorylation is one of the most important mechanisms of cellular responses to growth, stress metabolic and hormonal environmental changes. Most mammalian protein kinases have highly a homologous 30 to 32 kDa catalytic domain. • Most common method of reversible modification - activation and localization • Up to 1/3 of cellular proteins can be phosphorylated • Leads to a very fast response to cellular stress, hormonal changes, learning processes, transcription regulation .... • Different than allosteric or Michealis Menten regulation Protein Kinome To date – 518 human kinases known • 50 kinase families between yeast, invertebrate and mammaliane kinomes • 518 human PKs, most (478) belong to single super family whose catalytic domain are homologous. • Kinase dendrogram displays relative similarities based on catalytic domains. • AGC (PKA, PKG, PKC) • CAMK (Casein kinase 1) • CMGC (CDC, MAPK, GSK3, CLK) • STE (Sterile 7, 11 & 20 kinases) • TK (Tryosine kinases memb and cyto) • TKL (Tyrosine kinase-like) • Phosphorylation stabilized thermodynamically - only half available energy used in adding phosphoryl to protein - change in free energy forces phosphorylation reaction in one direction • Phosphatases reverse direction • The rate of reaction of most phosphatases are 1000 times faster • Phosphorylation occurs on Ser/The or Tyr • What differences occur due to the addition of a phosphoryl group? • Regulation of protein phosphorylation varies depending on protein - some turned on or off -

Inhibition of Src Family Kinases and Receptor Tyrosine Kinases by Dasatinib: Possible Combinations in Solid Tumors
Published OnlineFirst June 13, 2011; DOI: 10.1158/1078-0432.CCR-10-2616 Clinical Cancer Molecular Pathways Research Inhibition of Src Family Kinases and Receptor Tyrosine Kinases by Dasatinib: Possible Combinations in Solid Tumors Juan Carlos Montero1, Samuel Seoane1, Alberto Ocaña2,3, and Atanasio Pandiella1 Abstract Dasatinib is a small molecule tyrosine kinase inhibitor that targets a wide variety of tyrosine kinases implicated in the pathophysiology of several neoplasias. Among the most sensitive dasatinib targets are ABL, the SRC family kinases (SRC, LCK, HCK, FYN, YES, FGR, BLK, LYN, and FRK), and the receptor tyrosine kinases c-KIT, platelet-derived growth factor receptor (PDGFR) a and b, discoidin domain receptor 1 (DDR1), c-FMS, and ephrin receptors. Dasatinib inhibits cell duplication, migration, and invasion, and it triggers apoptosis of tumoral cells. As a consequence, dasatinib reduces tumoral mass and decreases the metastatic dissemination of tumoral cells. Dasatinib also acts on the tumoral microenvironment, which is particularly important in the bone, where dasatinib inhibits osteoclastic activity and favors osteogenesis, exerting a bone-protecting effect. Several preclinical studies have shown that dasatinib potentiates the antitumoral action of various drugs used in the oncology clinic, paving the way for the initiation of clinical trials of dasatinib in combination with standard-of-care treatments for the therapy of various neoplasias. Trials using combinations of dasatinib with ErbB/HER receptor antagonists are being explored in breast, head and neck, and colorectal cancers. In hormone receptor–positive breast cancer, trials using combina- tions of dasatinib with antihormonal therapies are ongoing. Dasatinib combinations with chemother- apeutic agents are also under development in prostate cancer (dasatinib plus docetaxel), melanoma (dasatinib plus dacarbazine), and colorectal cancer (dasatinib plus oxaliplatin plus capecitabine). -

Brief Definitive Report ZAP- 70 Gene
Brief Definitive Report Reconstitution of T Cell Receptor Signaling in ZAP-70-deficient Cells by Retroviral Transduction of the ZAP- 70 Gene By Naomi Taylor,* Kevin B. Bacon,¢ Susan Smith,* Thomas Jahn,* Theresa A. Kadlecekfl Lisa Uribe,* Donald B. Kohn,* Erwin W. Gelfand,IIArthur Weiss,~ and Kenneth Weinberg* From the *Division of Research Immunology and Bone Marrow Transplantation, Children's Hospital Los Angeles, Los Angeles, California 90027; :~DNAX Research Institute, Palo Alto, California 94304; gHoward Hughes Medical Institute, Department of Medicine and of Microbiology and Immunology, University of California, San Francisco, California 94143; and IIDivision of Basic Sciences and Molecular Signal Transduction Program, Department of Pediatrics, National Jewish Centerfor Immunology and Respiratory Diseases, Denver, Colorado 80206 Summal-y A variant of severe combined lmmunodeficiency syndrome (SCID) with a selective inability to produce CD8 single positive T cells and a signal transduction defect in peripheral CD4 + cells has recently been shown to be the result of mutations in the ZAP-70 gene. T cell receptor (TCR) signaling requires the association of the ZAP-70 protein tyrosine kinase with the TCR complex. Human T cell leukemia virus type I-transformed CD4 + T cell lines w.ere established from ZAP-70-deficient patients and normal controls. ZAP-70 was expressed and appropriately phosphorylated in normal T cell lines after TCR engagement, but was not detected in T cell lines from ZAP-70-deficient patients. To determine whether signaling could be reconstituted, wild-type ZAP-70 was introduced into deficient cells with a ZAP-70 retroviral vector. High titer producer clones expressing ZAP-70 were generated in the Gibbon ape leukemia virus packaging line PG13. -

Wrangling Phosphoproteomic Data to Elucidate Cancer Signaling Pathways
University of Montana ScholarWorks at University of Montana Biological Sciences Faculty Publications Biological Sciences 1-3-2013 Wrangling Phosphoproteomic Data to Elucidate Cancer Signaling Pathways Mark L. Grimes University of Montana - Missoula, [email protected] Wan-Jui Lee Laurens van der Maaten Paul Shannon Follow this and additional works at: https://scholarworks.umt.edu/biosci_pubs Part of the Biology Commons Let us know how access to this document benefits ou.y Recommended Citation Grimes, Mark L.; Lee, Wan-Jui; van der Maaten, Laurens; and Shannon, Paul, "Wrangling Phosphoproteomic Data to Elucidate Cancer Signaling Pathways" (2013). Biological Sciences Faculty Publications. 178. https://scholarworks.umt.edu/biosci_pubs/178 This Article is brought to you for free and open access by the Biological Sciences at ScholarWorks at University of Montana. It has been accepted for inclusion in Biological Sciences Faculty Publications by an authorized administrator of ScholarWorks at University of Montana. For more information, please contact [email protected]. Wrangling Phosphoproteomic Data to Elucidate Cancer Signaling Pathways Mark L. Grimes1*, Wan-Jui Lee2, Laurens van der Maaten2, Paul Shannon3 1 Division of Biological Sciences, University of Montana, Missoula, Montana, United States of America, 2 Pattern Recognition and Bioinformatics Group, Delft University of Technology, CD Delft, The Netherlands, 3 Fred Hutchison Cancer Research Institute, Seattle, Washington, United States of America Abstract The interpretation of biological data sets is essential for generating hypotheses that guide research, yet modern methods of global analysis challenge our ability to discern meaningful patterns and then convey results in a way that can be easily appreciated. Proteomic data is especially challenging because mass spectrometry detectors often miss peptides in complex samples, resulting in sparsely populated data sets.