A Disease Spectrum for ITPA Variation: Advances in Biochemical and Clinical Research Nicholas E
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Inosine in Biology and Disease
G C A T T A C G G C A T genes Review Inosine in Biology and Disease Sundaramoorthy Srinivasan 1, Adrian Gabriel Torres 1 and Lluís Ribas de Pouplana 1,2,* 1 Institute for Research in Biomedicine, Barcelona Institute of Science and Technology, 08028 Barcelona, Catalonia, Spain; [email protected] (S.S.); [email protected] (A.G.T.) 2 Catalan Institution for Research and Advanced Studies, 08010 Barcelona, Catalonia, Spain * Correspondence: [email protected]; Tel.: +34-934034868; Fax: +34-934034870 Abstract: The nucleoside inosine plays an important role in purine biosynthesis, gene translation, and modulation of the fate of RNAs. The editing of adenosine to inosine is a widespread post- transcriptional modification in transfer RNAs (tRNAs) and messenger RNAs (mRNAs). At the wobble position of tRNA anticodons, inosine profoundly modifies codon recognition, while in mRNA, inosines can modify the sequence of the translated polypeptide or modulate the stability, localization, and splicing of transcripts. Inosine is also found in non-coding and exogenous RNAs, where it plays key structural and functional roles. In addition, molecular inosine is an important secondary metabolite in purine metabolism that also acts as a molecular messenger in cell signaling pathways. Here, we review the functional roles of inosine in biology and their connections to human health. Keywords: inosine; deamination; adenosine deaminase acting on RNAs; RNA modification; translation Citation: Srinivasan, S.; Torres, A.G.; Ribas de Pouplana, L. Inosine in 1. Introduction Biology and Disease. Genes 2021, 12, 600. https://doi.org/10.3390/ Inosine was one of the first nucleobase modifications discovered in nucleic acids, genes12040600 having been identified in 1965 as a component of the first sequenced transfer RNA (tRNA), tRNAAla [1]. -

Standard Abbreviations
Journal of CancerJCP Prevention Standard Abbreviations Journal of Cancer Prevention provides a list of standard abbreviations. Standard Abbreviations are defined as those that may be used without explanation (e.g., DNA). Abbreviations not on the Standard Abbreviations list should be spelled out at first mention in both the abstract and the text. Abbreviations should not be used in titles; however, running titles may carry abbreviations for brevity. ▌Abbreviations monophosphate ADP, dADP adenosine diphosphate, deoxyadenosine IR infrared diphosphate ITP, dITP inosine triphosphate, deoxyinosine AMP, dAMP adenosine monophosphate, deoxyadenosine triphosphate monophosphate LOH loss of heterozygosity ANOVA analysis of variance MDR multiple drug resistance AP-1 activator protein-1 MHC major histocompatibility complex ATP, dATP adenosine triphosphate, deoxyadenosine MRI magnetic resonance imaging trip hosphate mRNA messenger RNA bp base pair(s) MTS 3-(4,5-dimethylthiazol-2-yl)-5-(3- CDP, dCDP cytidine diphosphate, deoxycytidine diphosphate carboxymethoxyphenyl)-2-(4-sulfophenyl)- CMP, dCMP cytidine monophosphate, deoxycytidine mono- 2H-tetrazolium phosphate mTOR mammalian target of rapamycin CNBr cyanogen bromide MTT 3-(4,5-Dimethylthiazol-2-yl)-2,5- cDNA complementary DNA diphenyltetrazolium bromide CoA coenzyme A NAD, NADH nicotinamide adenine dinucleotide, reduced COOH a functional group consisting of a carbonyl and nicotinamide adenine dinucleotide a hydroxyl, which has the formula –C(=O)OH, NADP, NADPH nicotinamide adnine dinucleotide -

Epistasis-Driven Identification of SLC25A51 As a Regulator of Human
ARTICLE https://doi.org/10.1038/s41467-020-19871-x OPEN Epistasis-driven identification of SLC25A51 as a regulator of human mitochondrial NAD import Enrico Girardi 1, Gennaro Agrimi 2, Ulrich Goldmann 1, Giuseppe Fiume1, Sabrina Lindinger1, Vitaly Sedlyarov1, Ismet Srndic1, Bettina Gürtl1, Benedikt Agerer 1, Felix Kartnig1, Pasquale Scarcia 2, Maria Antonietta Di Noia2, Eva Liñeiro1, Manuele Rebsamen1, Tabea Wiedmer 1, Andreas Bergthaler1, ✉ Luigi Palmieri2,3 & Giulio Superti-Furga 1,4 1234567890():,; About a thousand genes in the human genome encode for membrane transporters. Among these, several solute carrier proteins (SLCs), representing the largest group of transporters, are still orphan and lack functional characterization. We reasoned that assessing genetic interactions among SLCs may be an efficient way to obtain functional information allowing their deorphanization. Here we describe a network of strong genetic interactions indicating a contribution to mitochondrial respiration and redox metabolism for SLC25A51/MCART1, an uncharacterized member of the SLC25 family of transporters. Through a combination of metabolomics, genomics and genetics approaches, we demonstrate a role for SLC25A51 as enabler of mitochondrial import of NAD, showcasing the potential of genetic interaction- driven functional gene deorphanization. 1 CeMM Research Center for Molecular Medicine of the Austrian Academy of Sciences, Vienna, Austria. 2 Laboratory of Biochemistry and Molecular Biology, Department of Biosciences, Biotechnologies and Biopharmaceutics, -

High-Purity Rebaudioside D and Applications Hochreines Rebaudiosid-D Und Anwendungen Rébaudioside D De Grande Pureté Et Applications
(19) TZZ Z_T (11) EP 2 708 548 B1 (12) EUROPEAN PATENT SPECIFICATION (45) Date of publication and mention (51) Int Cl.: of the grant of the patent: C07H 1/08 (2006.01) A21D 2/36 (2006.01) 06.12.2017 Bulletin 2017/49 A23G 1/42 (2006.01) (21) Application number: 13196410.8 (22) Date of filing: 13.10.2010 (54) High-Purity Rebaudioside D and Applications Hochreines Rebaudiosid-D und Anwendungen Rébaudioside D de grande pureté et applications (84) Designated Contracting States: (72) Inventors: AL AT BE BG CH CY CZ DE DK EE ES FI FR GB • Abelyan, Varuzhan GR HR HU IE IS IT LI LT LU LV MC MK MT NL NO 50480 Kuala Lumpur (MY) PL PT RO RS SE SI SK SM TR • Markosyan, Avetik 59200 Kuala Lumpur (MY) (30) Priority: 15.10.2009 US 580233 • Abelyan, Lidia 24.05.2010 US 785501 50480 Kuala Lumpur (MY) 24.05.2010 US 785504 24.05.2010 US 785506 (74) Representative: Hocking, Adrian Niall et al 24.05.2010 US 785507 Albright IP Limited 24.05.2010 US 785508 County House 24.05.2010 US 786392 Bayshill Road 24.05.2010 US 786402 Cheltenham, Glos. GL50 3BA (GB) 24.05.2010 US 786413 24.05.2010 US 786416 (56) References cited: 24.05.2010 US 786427 WO-A1-2009/071277 24.05.2010 US 786430 24.05.2010 US 786419 • I. SAKAMOTO ET AL: "Application of 13C NMR spectroscopy to chemistry of natural glycosices: (43) Date of publication of application: rebaudioside-C,a newsweet diterpeneglycoside 19.03.2014 Bulletin 2014/12 of Stevia rebaudiana", CHEM. -

Utilization of L-Methionine and S-Adenosyl-L-Methionine for Methylation of Soluble RNA by Mouse Liver and Hepatoma Extracts1
[CANCER RESEARCH 27, 646-«S3,April 1967] Utilization of L-Methionine and S-Adenosyl-L-methionine for Methylation of Soluble RNA by Mouse Liver and Hepatoma Extracts1 R. L. HANCOCK The Jackson Laboratory, Bar Harbor, Maine SUMMARY following reaction: ATP + L-methionine = SAM + pyrophos The methyl group of L-methionine or S-adenosyl-L-methionine phate + monophosphate (7). SAM is used in the methylation of sRNA by sRNA methylases in the following reaction: sRNA + is used by nonparticulate mouse liver and hepatoma prepara SAM = methylated sRNA + S-adenosyl-L-homocysteine (10). tions to methylate soluble ribonucleic acid (sRNA). Esclierichia The two enzyme reactions are presented along with a representa coli K12 sRNA was a more active substrate than yeast sRNA. tion of the origin of the unmethylated sRNA (tp-RNA) in Chart Four different lots of E. coli B sRNA gave similar incorporations between lots. "Stripped" and "nonstripped" sRNA had similar 1. An array of RNA methylases exhibiting specificity for par ticular bases and for sRNA from different biologic sources, along amounts of methyl incorporation. Other nucleoside triphosphates with other, less si^ecific RNA methylases, has been described besides adenosine triphosphate were capable of sup]x>rting the incorjioration of methyl groups from L-methionine into sRNA. (11, 15, 16, 17, 22). The most extensively methylated sRNA molecules known have been found in mammary adenocarcinoma The nonspecificity of the nucleoside triphosphate reaction was tissue (3), and recently Mittelman et al. (18) have found RNA shown to be due to an adenosine diphosphokinase reaction and methylase activity to be extremely high in certain viral-induced not to nucleoside tri phosphate: L-methionine nucleosidyl trans- tumor cells. -

Role of Inosine Triphosphate Pyrophosphatase Gene Variant on Fever Incidence During Zidovudine Antiretroviral Therapy
Role of inosine triphosphate pyrophosphatase gene variant on fever incidence during zidovudine antiretroviral therapy A.V.C. Coelho1, S.P.S. Silva2, L. Zandonà3, G. Stocco3, G. Decorti3 and S. Crovella1,4 1Departamento de Genética, Universidade Federal de Pernambuco Federal de Pernambuco, Recife, PE, Brasil 2Programa de Pós-Graduação em Inovação Terapêutica, Universidade Federal de Pernambuco Federal de Pernambuco, Recife, PE, Brasil 3Department of Life Sciences, University of Trieste, Trieste, Italy 4Institute for Maternal and Child Health, Scientific Institute for Research, Hospitalization and Care Burlo Garofolo, Trieste, Italy Corresponding author: A.V.C. Coelho E-mail: [email protected] / [email protected] Genet. Mol. Res. 16 (1): gmr16019373 Received September 22, 2016 Accepted November 18, 2016 Published January 23, 2017 DOI http://dx.doi.org/10.4238/gmr16019373 Copyright © 2017 The Authors. This is an open-access article distributed under the terms of the Creative Commons Attribution ShareAlike (CC BY-SA) 4.0 License. ABSTRACT. Zidovudine, the antiretroviral drug used to treat HIV infection, commonly causes adverse effects, such as systemic fever and gastrointestinal alterations. In the present study, the potential role of inosine triphosphate pyrophosphatase (ITPA) gene variant on the incidence of adverse events during antiretroviral therapy (ART) of HIV with zidovudine was discussed. Individuals from Northeastern Brazil (N = 204) receiving treatment for HIV-1 infection were recruited. Zidovudine-related adverse effects developed during the treatment were registered. The rs1127354 polymorphism in the ITPA gene was Genetics and Molecular Research 16 (1): gmr16019373 A.V.C. Coelho et al. 2 genotyped using real-time PCR to assess whether this single nucleotide polymorphism was associated with the occurrence of zidovudine- related adverse effects. -
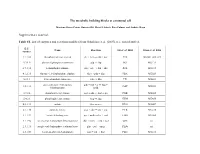
The Metabolic Building Blocks of a Minimal Cell Supplementary
The metabolic building blocks of a minimal cell Mariana Reyes-Prieto, Rosario Gil, Mercè Llabrés, Pere Palmer and Andrés Moya Supplementary material. Table S1. List of enzymes and reactions modified from Gabaldon et. al. (2007). n.i.: non identified. E.C. Name Reaction Gil et. al. 2004 Glass et. al. 2006 number 2.7.1.69 phosphotransferase system glc + pep → g6p + pyr PTS MG041, 069, 429 5.3.1.9 glucose-6-phosphate isomerase g6p ↔ f6p PGI MG111 2.7.1.11 6-phosphofructokinase f6p + atp → fbp + adp PFK MG215 4.1.2.13 fructose-1,6-bisphosphate aldolase fbp ↔ gdp + dhp FBA MG023 5.3.1.1 triose-phosphate isomerase gdp ↔ dhp TPI MG431 glyceraldehyde-3-phosphate gdp + nad + p ↔ bpg + 1.2.1.12 GAP MG301 dehydrogenase nadh 2.7.2.3 phosphoglycerate kinase bpg + adp ↔ 3pg + atp PGK MG300 5.4.2.1 phosphoglycerate mutase 3pg ↔ 2pg GPM MG430 4.2.1.11 enolase 2pg ↔ pep ENO MG407 2.7.1.40 pyruvate kinase pep + adp → pyr + atp PYK MG216 1.1.1.27 lactate dehydrogenase pyr + nadh ↔ lac + nad LDH MG460 1.1.1.94 sn-glycerol-3-phosphate dehydrogenase dhp + nadh → g3p + nad GPS n.i. 2.3.1.15 sn-glycerol-3-phosphate acyltransferase g3p + pal → mag PLSb n.i. 2.3.1.51 1-acyl-sn-glycerol-3-phosphate mag + pal → dag PLSc MG212 acyltransferase 2.7.7.41 phosphatidate cytidyltransferase dag + ctp → cdp-dag + pp CDS MG437 cdp-dag + ser → pser + 2.7.8.8 phosphatidylserine synthase PSS n.i. cmp 4.1.1.65 phosphatidylserine decarboxylase pser → peta PSD n.i. -

Coupled Nucleoside Phosphorylase Reactions in Escherichia Coli John Lewis Ott Iowa State College
Iowa State University Capstones, Theses and Retrospective Theses and Dissertations Dissertations 1956 Coupled nucleoside phosphorylase reactions in Escherichia coli John Lewis Ott Iowa State College Follow this and additional works at: https://lib.dr.iastate.edu/rtd Part of the Biochemistry Commons, and the Microbiology Commons Recommended Citation Ott, John Lewis, "Coupled nucleoside phosphorylase reactions in Escherichia coli " (1956). Retrospective Theses and Dissertations. 13758. https://lib.dr.iastate.edu/rtd/13758 This Dissertation is brought to you for free and open access by the Iowa State University Capstones, Theses and Dissertations at Iowa State University Digital Repository. It has been accepted for inclusion in Retrospective Theses and Dissertations by an authorized administrator of Iowa State University Digital Repository. For more information, please contact [email protected]. NOTE TO USERS This reproduction is the best copy available. UMI COUPLED NUCLEOSIDE PHOSPHORYLASE REACTIONS IN ESCHERICHIA COLI / by John Lewis Ott A Dissertation Submitted to the Graduate Faculty in Partial Fulfillment of The Requirements for the Degree of DOCTOR OF PHILOSOPHY Major Subject: Physlolgglcal Bacteriology Approved: Signature was redacted for privacy. In Charge of Major Work Signature was redacted for privacy. Head of Major Department Signature was redacted for privacy. Dean of Graduate College Iowa State College 1956 UMI Number: DP12892 INFORMATION TO USERS The quality of this reproduction is dependent upon the quality of the copy submitted. Broken or indistinct print, colored or poor quality illustrations and photographs, print bleed-through, substandard margins, and improper alignment can adversely affect reproduction. In the unlikely event that the author did not send a complete manuscript and there are missing pages, these will be noted. -

Original Article Identification of B Cells Participated in the Mechanism of Postmenopausal Women Osteoporosis Using Microarray Analysis
Int J Clin Exp Med 2015;8(1):1027-1034 www.ijcem.com /ISSN:1940-5901/IJCEM0003564 Original Article Identification of B cells participated in the mechanism of postmenopausal women osteoporosis using microarray analysis Bing Yan*, Jie Li*, Li Zhang Department of Orthopedics, Shandong Provincial Traditional Chinise Medical Hospital, Jinan 250014, Shandong Province, China. *Equal contributors. Received November 3, 2014; Accepted January 7, 2015; Epub January 15, 2015; Published January 30, 2015 Abstract: To further understand the molecular mechanism of lymphocytes B cells in postmenopausal women os- teoporosis. Microarray data (GSE7429) were downloaded from Gene Expression Omnibus, in which B cells were separated from the whole blood of postmenopausal women, including 10 with high bone mineral density (BMD) and 10 with low BMD. Differentially expressed genes (DEGs) between high and low BMD women were identified by Student’s t-test, and P < 0.01 was used as the significant criterion. Functional enrichment analysis was performed for up- and down-regulated DEGs using KEGG, REACTOME, and Gene Ontology (GO) databases. Protein-protein inter- action network (PPI) of up- and down-regulated DEGs was respectively constructed by Cytoscape software using the STRING data. Total of 169 up-regulated and 69 down-regulated DEGs were identified. Functional enrichment analy- sis indicated that the genes (ITPA, ATIC, UMPS, HPRT1, COX10 and COX15) might participate in metabolic pathways, MAP3K10 and MAP3K9 might participate in the activation of JNKK activity, COX10 and COX15 might involve in mitochondrial electron transport, and ATIC, UMPS and HPRT1 might involve in transferase activity. MAPK3, ITPA, ATIC, UMPS and HPRT1 with a higher degree in PPI network were identified. -

By Cx- and 3-Adrenoceptors W
Br. J. Pharmac. (1970), 38, 399-415. J'hibition of rabbit intestine mediated by cx- and 3-adrenoceptors W. C. BOWMAN AND MOIRA T. HALL Department of Pharmacology, University of Strathclyde, Glasgow, C.1 Summary 1. The effects of some a- and /8-adrenoceptor agonists and antagonists were studied on isolated segments of rabbit intestine in an attempt to characterize the two types of inhibitory response produced by sympathomimetic amines. 2. Phenylephrine, an a-adrenoceptor agonist, produced an inhibition of rapid onset, from which recovery occurred despite the continued presence of the drug. On washout there was an overshoot in contraction height. Isoprenaline, a /3- adrenoceptor agonist, produced an inhibition of slow onset which was main- tained throughout the presence of the drug and there was no overshoot on washout. 3. Adrenaline resembled phenylephrine more closely than it resembled iso- prenaline, in that it showed more affinity for a-adrenoceptors, whereas nor- adrenaline, and the transmitter released on periarterial nerve stimulation, behaved more like isoprenaline, although both types of receptor were affected. 4. Adenosine-5'-triphosphate produced an inhibition resembling that produced by an a-adrenoceptor agonist, whereas the dibutyryl analogue of cyclic adeno- sine 3',5'-monophosphate (cyclic 3',5'-AMP) produced an inhibition resembling that produced by a /3-adrenoceptor agonist. 5. In critical concentrations theophylline augmented and imidazole inhibited /3-adrenoceptor mediated responses, as well as responses to dibutyryl cyclic AMP. However, additional actions of theophylline and imidazole were also demonstrated. 6. Responses mediated by a-adrenoceptors, but not those mediated by ,8-adrenoceptors, were blocked by membrane stabilizers, quinidine being the most potent of those studied. -

Sheep Gene Mapping by Somatic Cell Hybridization
Sheep gene mapping by somatic cell hybridization N Saïdi-Mehtar M Imam-Ghali1 S Heuertz2 MC Hors-Cayla2 1 Université d’Oran Es-Sénia, Laboratoire de Biologie M oléculaire et Génétique, Unite de Recherches de Biologie, Algeria; z 2 INSERM U12, Unit6 de Recherches de Genetique M6dicale, Paris, France (Proceedings of the 9th European Colloquium on Cytogenetics of Domestic Animals; Toulouse-Auzeville, 10-13 July 1990) sheep / ovine / gene mapping / synteny / cell hybrid INTRODUCTION Twenty-five hamster x sheep hybrid cell lines were previously obtained (Saidi- Mehtar et al, 1981b). Analysis of these hybrids has already enabled the identification of 5 syntenic groups and 10 independent markers (Saidi-Mehtar et al, 1981a, b, 1987; Millot et al, 1981; Imam-Ghali et al, 1987). In this paper, we report results obtained with 3 other enzyme markers: aconitase 1 (AC01), inosine triphosphatase (ITPA) and glyceraldehyde-3-phosphate dehydrogenase (GAPD); and 6 DNA markers: ornithine transcarbamylase (OTC), color vision red pigment (RCB), raf oncogene (ARAF1) and /3-gene locus DR of human lymphocyte antigen region (HLA-DR!). ENZYME MARKERS AC01, ITPA and GAPD were studied by cellulose acetate electrophoresis using modified versions of the methods described by Womack and Moll (1986). The cytoplasmic (AC01) and mitochondrial (AC02) forms of aconitase were identified by successive freezing and thawing of cells followed by electrophoresis of the supernatants. AC01 was obtained first and corresponded to the most anodal band. A different electrophoretic migration between sheep and hamster was only observed for ACO1. Ovine ITPA migrated more anodally than hamster ITPA. Ovine GAPD is a trimeric enzyme: positive hybrids for sheep GAPD showed 2 intermediate bands (fig 1). -

Inosine Triphosphate Pyrophosphatase (ITPA) (NM 181493) Human Tagged ORF Clone Product Data
OriGene Technologies, Inc. 9620 Medical Center Drive, Ste 200 Rockville, MD 20850, US Phone: +1-888-267-4436 [email protected] EU: [email protected] CN: [email protected] Product datasheet for RC215516L4 Inosine triphosphate pyrophosphatase (ITPA) (NM_181493) Human Tagged ORF Clone Product data: Product Type: Expression Plasmids Product Name: Inosine triphosphate pyrophosphatase (ITPA) (NM_181493) Human Tagged ORF Clone Tag: mGFP Symbol: ITPA Synonyms: C20orf37; DEE35; dJ794I6.3; HLC14-06-P; ITPase; My049; NTPase Vector: pLenti-C-mGFP-P2A-Puro (PS100093) E. coli Selection: Chloramphenicol (34 ug/mL) Cell Selection: Puromycin ORF Nucleotide The ORF insert of this clone is exactly the same as(RC215516). Sequence: Restriction Sites: SgfI-MluI Cloning Scheme: ACCN: NM_181493 ORF Size: 531 bp This product is to be used for laboratory only. Not for diagnostic or therapeutic use. View online » ©2021 OriGene Technologies, Inc., 9620 Medical Center Drive, Ste 200, Rockville, MD 20850, US 1 / 2 Inosine triphosphate pyrophosphatase (ITPA) (NM_181493) Human Tagged ORF Clone – RC215516L4 OTI Disclaimer: The molecular sequence of this clone aligns with the gene accession number as a point of reference only. However, individual transcript sequences of the same gene can differ through naturally occurring variations (e.g. polymorphisms), each with its own valid existence. This clone is substantially in agreement with the reference, but a complete review of all prevailing variants is recommended prior to use. More info OTI Annotation: This clone was engineered to express the complete ORF with an expression tag. Expression varies depending on the nature of the gene. RefSeq: NM_181493.1 RefSeq Size: 1155 bp RefSeq ORF: 534 bp Locus ID: 3704 UniProt ID: Q9BY32 Protein Families: Druggable Genome Protein Pathways: Drug metabolism - other enzymes, Metabolic pathways, Purine metabolism, Pyrimidine metabolism MW: 19.4 kDa Gene Summary: This gene encodes an inosine triphosphate pyrophosphohydrolase.