Generate Metabolic Map Poster
Total Page:16
File Type:pdf, Size:1020Kb

Load more
Recommended publications
-

| Hai Lui a Un Acutul Luniit Moonhiti
|HAI LUI AUN ACUTULUS010006055B2 LUNIIT MOONHITI (12 ) United States Patent (10 ) Patent No. : US 10 , 006 , 055 B2 Burk et al. (45 ) Date of Patent: Jun . 26 , 2018 ( 54 ) MICROORGANISMS FOR PRODUCING 2002/ 0168654 A1 11/ 2002 Maranas et al. 2003 / 0059792 Al 3 /2003 Palsson et al . BUTADIENE AND METHODS RELATED 2003 /0087381 A1 5 / 2003 Gokarn THERETO 2003 / 0224363 Al 12 /2003 Park et al . 2003 / 0233218 Al 12 /2003 Schilling (71 ) Applicant: Genomatica , Inc. , San Diego , CA (US ) 2004 / 0009466 AL 1 /2004 Maranas et al. 2004 / 0029149 Al 2 /2004 Palsson et al. ( 72 ) Inventors : Mark J . Burk , San Diego , CA (US ) ; 2004 / 0072723 A1 4 /2004 Palsson et al. Anthony P . Burgard , Bellefonte , PA 2004 / 0152159 Al 8 / 2004 Causey et al . 2005 /0042736 A1 2 / 2005 San et al . (US ) ; Robin E . Osterhout , San Diego , 2005 / 0079482 A1 4 / 2005 Maranas et al . CA (US ) ; Jun Sun , San Diego , CA 2006 / 0046288 Al 3 / 2006 Ka - Yiu et al. ( US ) ; Priti Pharkya , San Diego , CA 2006 / 0073577 A1 4 / 2006 Ka - Yiu et al . (US ) 2007 /0184539 Al 8 / 2007 San et al . 2009 / 0047718 Al 2 / 2009 Blaschek et al . 2009 / 0047719 Al 2 / 2009 Burgard et al . (73 ) Assignee : Genomatica , Inc ., San Diego , CA (US ) 2009 /0191593 A1 7 / 2009 Burk et al . 2010 / 0003716 A1 1 / 2010 Cervin et al. ( * ) Notice : Subject to any disclaimer , the term of this 2010 /0184171 Al 7 /2010 Jantama et al. patent is extended or adjusted under 35 2010 /0304453 Al 12 / 2010 Trawick et al . -

(12) Patent Application Publication (10) Pub. No.: US 2014/0155567 A1 Burk Et Al
US 2014O155567A1 (19) United States (12) Patent Application Publication (10) Pub. No.: US 2014/0155567 A1 Burk et al. (43) Pub. Date: Jun. 5, 2014 (54) MICROORGANISMS AND METHODS FOR (60) Provisional application No. 61/331,812, filed on May THE BIOSYNTHESIS OF BUTADENE 5, 2010. (71) Applicant: Genomatica, Inc., San Diego, CA (US) Publication Classification (72) Inventors: Mark J. Burk, San Diego, CA (US); (51) Int. Cl. Anthony P. Burgard, Bellefonte, PA CI2P 5/02 (2006.01) (US); Jun Sun, San Diego, CA (US); CSF 36/06 (2006.01) Robin E. Osterhout, San Diego, CA CD7C II/6 (2006.01) (US); Priti Pharkya, San Diego, CA (52) U.S. Cl. (US) CPC ................. CI2P5/026 (2013.01); C07C II/I6 (2013.01); C08F 136/06 (2013.01) (73) Assignee: Genomatica, Inc., San Diego, CA (US) USPC ... 526/335; 435/252.3:435/167; 435/254.2: (21) Appl. No.: 14/059,131 435/254.11: 435/252.33: 435/254.21:585/16 (22) Filed: Oct. 21, 2013 (57) ABSTRACT O O The invention provides non-naturally occurring microbial Related U.S. Application Data organisms having a butadiene pathway. The invention addi (63) Continuation of application No. 13/101,046, filed on tionally provides methods of using Such organisms to produce May 4, 2011, now Pat. No. 8,580,543. butadiene. Patent Application Publication Jun. 5, 2014 Sheet 1 of 4 US 2014/O155567 A1 ?ueudos!SMS |?un61– Patent Application Publication Jun. 5, 2014 Sheet 2 of 4 US 2014/O155567 A1 VOJ OO O Z?un61– Patent Application Publication US 2014/O155567 A1 {}}} Hººso Patent Application Publication Jun. -

Serine Proteases with Altered Sensitivity to Activity-Modulating
(19) & (11) EP 2 045 321 A2 (12) EUROPEAN PATENT APPLICATION (43) Date of publication: (51) Int Cl.: 08.04.2009 Bulletin 2009/15 C12N 9/00 (2006.01) C12N 15/00 (2006.01) C12Q 1/37 (2006.01) (21) Application number: 09150549.5 (22) Date of filing: 26.05.2006 (84) Designated Contracting States: • Haupts, Ulrich AT BE BG CH CY CZ DE DK EE ES FI FR GB GR 51519 Odenthal (DE) HU IE IS IT LI LT LU LV MC NL PL PT RO SE SI • Coco, Wayne SK TR 50737 Köln (DE) •Tebbe, Jan (30) Priority: 27.05.2005 EP 05104543 50733 Köln (DE) • Votsmeier, Christian (62) Document number(s) of the earlier application(s) in 50259 Pulheim (DE) accordance with Art. 76 EPC: • Scheidig, Andreas 06763303.2 / 1 883 696 50823 Köln (DE) (71) Applicant: Direvo Biotech AG (74) Representative: von Kreisler Selting Werner 50829 Köln (DE) Patentanwälte P.O. Box 10 22 41 (72) Inventors: 50462 Köln (DE) • Koltermann, André 82057 Icking (DE) Remarks: • Kettling, Ulrich This application was filed on 14-01-2009 as a 81477 München (DE) divisional application to the application mentioned under INID code 62. (54) Serine proteases with altered sensitivity to activity-modulating substances (57) The present invention provides variants of ser- screening of the library in the presence of one or several ine proteases of the S1 class with altered sensitivity to activity-modulating substances, selection of variants with one or more activity-modulating substances. A method altered sensitivity to one or several activity-modulating for the generation of such proteases is disclosed, com- substances and isolation of those polynucleotide se- prising the provision of a protease library encoding poly- quences that encode for the selected variants. -

Supplemental Methods
Supplemental Methods: Sample Collection Duplicate surface samples were collected from the Amazon River plume aboard the R/V Knorr in June 2010 (4 52.71’N, 51 21.59’W) during a period of high river discharge. The collection site (Station 10, 4° 52.71’N, 51° 21.59’W; S = 21.0; T = 29.6°C), located ~ 500 Km to the north of the Amazon River mouth, was characterized by the presence of coastal diatoms in the top 8 m of the water column. Sampling was conducted between 0700 and 0900 local time by gently impeller pumping (modified Rule 1800 submersible sump pump) surface water through 10 m of tygon tubing (3 cm) to the ship's deck where it then flowed through a 156 µm mesh into 20 L carboys. In the lab, cells were partitioned into two size fractions by sequential filtration (using a Masterflex peristaltic pump) of the pre-filtered seawater through a 2.0 µm pore-size, 142 mm diameter polycarbonate (PCTE) membrane filter (Sterlitech Corporation, Kent, CWA) and a 0.22 µm pore-size, 142 mm diameter Supor membrane filter (Pall, Port Washington, NY). Metagenomic and non-selective metatranscriptomic analyses were conducted on both pore-size filters; poly(A)-selected (eukaryote-dominated) metatranscriptomic analyses were conducted only on the larger pore-size filter (2.0 µm pore-size). All filters were immediately submerged in RNAlater (Applied Biosystems, Austin, TX) in sterile 50 mL conical tubes, incubated at room temperature overnight and then stored at -80oC until extraction. Filtration and stabilization of each sample was completed within 30 min of water collection. -

Computational and Systems Biology Themed Issue
View Article Online / Journal Homepage / Table of Contents for this issue Molecular BioSystems This article was published as part of the Computational and Systems Biology themed issue Please take a look at the full table of contents to access the other papers in this issue. Open Access Article. Published on 13 October 2009. Downloaded 9/23/2021 6:41:00 PM. View Article Online PAPER www.rsc.org/molecularbiosystems | Molecular BioSystems Amidoligases with ATP-grasp, glutamine synthetase-like and acetyltransferase-like domains: synthesis of novel metabolites and peptide modifications of proteinswz Lakshminarayan M. Iyer,a Saraswathi Abhiman,a A. Maxwell Burroughsb and L. Aravind*a Received 28th August 2009, Accepted 28th August 2009 First published as an Advance Article on the web 13th October 2009 DOI: 10.1039/b917682a Recent studies have shown that the ubiquitin system had its origins in ancient cofactor/amino acid biosynthesis pathways. Preliminary studies also indicated that conjugation systems for other peptide tags on proteins, such as pupylation, have evolutionary links to cofactor/amino acid biosynthesis pathways. Following up on these observations, we systematically investigated the non-ribosomal amidoligases of the ATP-grasp, glutamine synthetase-like and acetyltransferase folds by classifying the known members and identifying novel versions. We then established their contextual connections using information from domain architectures and conserved gene neighborhoods. This showed remarkable, previously uncharacterized functional links between diverse peptide ligases, several peptidases of unrelated folds and enzymes involved in synthesis of modified amino acids. Using the network of contextual connections we were able to predict numerous novel pathways for peptide synthesis and modification, amine-utilization, secondary metabolite synthesis and potential peptide-tagging systems. -

Comparative Genomics of the Mycobacterium Ulcerans And
MPhil Thesis Ken Doig (197821835) MPhil Thesis – 2012 Candidate Kenneth Douglas Doig Student No. 197821835 Title Comparative genomics of the Mycobacterium ulcerans and Mycobacterium marinum complex Status Submitted in total fulfilment of the requirements of the degree of Master of Philosophy Date September 2012 Supervisors A/Prof. Tim Stinear1 Dr. Torsten Seemann2 Prof. Richard Strugnell1 Department Microbiology and Immunology Faculty Medicine, Dentistry and Health Sciences The University of Melbourne 1. Department of Microbiology and Immunology, University of Melbourne, Parkville, Australia 2. Victorian Bioinformatics Consortium, Monash University, Clayton, Australia MPhil_Thesis-22.doc Page 1 of 1 MPhil Thesis Ken Doig (197821835) Abstract Buruli ulcer is a neglected tropical disease which is prevalent in Western Africa. Its etiological agent, Mycobacteria ulcerans occurs in a wide range of hosts and environments across the world. In contrast to its progenitor Mycobacteria marinum, the bacteria and related strains produce a toxin mycolactone, an immunosuppressive polyketide which gives rise to its pathogenicity. Bacterial pathogenesis has a number of hallmarks such as; an evolutionary bottleneck, insertion sequence (IS) expansion, pseudogene increase, genome reduction, horizontal gene transfers (HGT) and adaption to a new niche. M. ulcerans shows all of these characteristics and is a fine model for gaining a deeper understanding of the mechanics of bacterial pathogenesis in general and specifically for M. ulcerans and related strains. This research analyses the genomic makeup of 35 isolates that produce mycolactone and five M. marinum isolates. Such analysis have recently become possible through the rapid technological development of high throughput sequencing (HTS) allowing whole genome sequences of all isolates to be compared and contrasted. -

Table S3. Bootstrapped Frequency of COG Occurrences for Specific Metabolic Processes Encoded in the Antarctic Bacterioplankton Environmental Genomes
Table S3. Bootstrapped frequency of COG occurrences for specific metabolic processes encoded in the Antarctic bacterioplankton environmental genomes. Category COG Predicted Protein WEG SEG Inorganic carbon COG1850 Ribulose 1,5-bisphosphate carboxylase, large subunit 4.34 0.85 COG4451 Ribulose bisphosphate carboxylase small subunit 1.19 0.83 COG0574 Phosphoenolpyruvate synthase/pyruvate phosphate dikinase 13.7 6.63 COG4770 Acetyl/propionyl-CoA carboxylase, alpha subunit 7.91 4.85 Sulfur Metabolism (inorganic) COG2897 Rhodanese-related sulfurtransferase 5.64 3.25 COG0607 Rhodanese-related sulfurtransferase 6.02 5.68 COG2895 GTPases - Sulfate adenylate transferase subunit 1 2.28 4.88 COG1054 Predicted sulfurtransferase 2.53 4.17 COG0306 Phosphate/sulphate permeases 7.56 3.27 COG0659 Sulfate permease and related transporters (MFS superfamily) 15.72 15.22 COG0529 Adenylylsulfate kinase and related kinases 1.98 4.16 COG0225 Peptide methionine sulfoxide reductase 7.03 3.22 COG0229 Conserved domain frequently associated with peptide methionine sulfoxide reductase 3.77 2.47 COG0641 Arylsulfatase regulator (Fe-S oxidoreductase) 0.67 0 COG0526 Thiol-disulfide isomerase and thioredoxins 6.99 4.12 COG1651 Protein-disulfide isomerase 10.75 1.65 COG2041 Sulfite oxidase and related enzymes 5.51 3.05 COG2046 ATP sulfurylase (sulfate adenylyltransferase) 4.34 3.46 COG2221 Dissimilatory sulfite reductase (desulfoviridin), alpha and beta subunits 1.21 0 COG2920 Dissimilatory sulfite reductase (desulfoviridin), gamma subunit 1.9 0 COG0155 Sulfite reductase, -

Supplementary Informations SI2. Supplementary Table 1
Supplementary Informations SI2. Supplementary Table 1. M9, soil, and rhizosphere media composition. LB in Compound Name Exchange Reaction LB in soil LBin M9 rhizosphere H2O EX_cpd00001_e0 -15 -15 -10 O2 EX_cpd00007_e0 -15 -15 -10 Phosphate EX_cpd00009_e0 -15 -15 -10 CO2 EX_cpd00011_e0 -15 -15 0 Ammonia EX_cpd00013_e0 -7.5 -7.5 -10 L-glutamate EX_cpd00023_e0 0 -0.0283302 0 D-glucose EX_cpd00027_e0 -0.61972444 -0.04098397 0 Mn2 EX_cpd00030_e0 -15 -15 -10 Glycine EX_cpd00033_e0 -0.0068175 -0.00693094 0 Zn2 EX_cpd00034_e0 -15 -15 -10 L-alanine EX_cpd00035_e0 -0.02780553 -0.00823049 0 Succinate EX_cpd00036_e0 -0.0056245 -0.12240603 0 L-lysine EX_cpd00039_e0 0 -10 0 L-aspartate EX_cpd00041_e0 0 -0.03205557 0 Sulfate EX_cpd00048_e0 -15 -15 -10 L-arginine EX_cpd00051_e0 -0.0068175 -0.00948672 0 L-serine EX_cpd00054_e0 0 -0.01004986 0 Cu2+ EX_cpd00058_e0 -15 -15 -10 Ca2+ EX_cpd00063_e0 -15 -100 -10 L-ornithine EX_cpd00064_e0 -0.0068175 -0.00831712 0 H+ EX_cpd00067_e0 -15 -15 -10 L-tyrosine EX_cpd00069_e0 -0.0068175 -0.00233919 0 Sucrose EX_cpd00076_e0 0 -0.02049199 0 L-cysteine EX_cpd00084_e0 -0.0068175 0 0 Cl- EX_cpd00099_e0 -15 -15 -10 Glycerol EX_cpd00100_e0 0 0 -10 Biotin EX_cpd00104_e0 -15 -15 0 D-ribose EX_cpd00105_e0 -0.01862144 0 0 L-leucine EX_cpd00107_e0 -0.03596182 -0.00303228 0 D-galactose EX_cpd00108_e0 -0.25290619 -0.18317325 0 L-histidine EX_cpd00119_e0 -0.0068175 -0.00506825 0 L-proline EX_cpd00129_e0 -0.01102953 0 0 L-malate EX_cpd00130_e0 -0.03649016 -0.79413596 0 D-mannose EX_cpd00138_e0 -0.2540567 -0.05436649 0 Co2 EX_cpd00149_e0 -
Generated by SRI International Pathway Tools Version 25.0, Authors S
Authors: Pallavi Subhraveti Ron Caspi Peter Midford Peter D Karp An online version of this diagram is available at BioCyc.org. Biosynthetic pathways are positioned in the left of the cytoplasm, degradative pathways on the right, and reactions not assigned to any pathway are in the far right of the cytoplasm. Transporters and membrane proteins are shown on the membrane. Ingrid Keseler Periplasmic (where appropriate) and extracellular reactions and proteins may also be shown. Pathways are colored according to their cellular function. Gcf_000753795Cyc: Shewanella mangrovi YQH10 Cellular Overview Connections between pathways are omitted for legibility. Anamika Kothari molybdate phosphate phosphate PstC RS04785 RS08110 FtsX GspE RS18465 FliL RS02695 RS05740 ModC RsxB RS12030 SohB FliG FliM FliQ FlhF RS02410 RS02250 RS10010 RS15620 LptG RS01680 RS01605 MurJ RS02400 RS01330 RS11775 RS06520 RS11750 RS08505 FlhB RS00580 RS09545 LptC CrcB RS06405 RS14070 RS14040 FliG RS08485 RS08495 RS00745 RS15365 RS01080 MotA RS11785 RS09655 CcmD RS19200 YajC BrnQ TrkA ArsB RS02180 RS05465 NrfE FlhB FliO FliJ FliF TolR RS10365 ZipA RS06935 RS05785 RS12390 GspL RS03340 RS03595 RS06070 RS07340 RS16385 UbiB PstB PstB molybdate phosphate phosphate LolA RS14075 6 Tetrapyrrole Biosynthesis N -(3-methylbut- ditrans,octacis- a peptidoglycan with 3-phenyl-2- ditrans,octacis- ditrans,octacis- a [bis(guanylyl UDP-N-acetyl- (4R)-4-hydroxy- a [protein]-L- a [protein]-L- a [protein] Cofactor, Carrier, and Vitamin Biosynthesis a sulfurated a lipid II MoO - a uracil in DNA a -

ABSTRACT BOZDAG, AHMET. Investigation of Methanol and Formaldehyde Metabolism in Bacillus Methanolicus
ABSTRACT BOZDAG, AHMET. Investigation of Methanol and Formaldehyde Metabolism in Bacillus methanolicus.(Under the direction of Prof. Michael C. Flickinger). Bacillus methanolicus is a Gram-positive aerobic methylotroph growing optimally at 50-53 °C. Wild-type strains of B. methanolicus have been reported to secrete 58 g/l of L- glutamate in fed-batch cultures. Mutants of B. methanolicus created via classical mutangenesis can secrete 37 g/l of L-lysine, at 50 °C. The genes required for methylotrophyin B. methanolicus are encoded by an endogenous plasmid, pBM19 in strain MGA3, except for hexulose phosphate synthase (hps) and phosphohexuloisomerase (phi) which are encoded on the chromosome.It is a promising candidate for industrial production of chemical intermediates or amino acids from methanol. B. methanolicus employs the ribulose monophospate (RuMP) pathway to assimilate the carbon derived from the methanol, but enzymes that dissimilate carbon are not identified, although formaldehyde and formate were identified as intermediates by 13C NMR. It is important to understand how methanol is oxidized to formaldehyde and then, to formate and carbon dioxide. This study aims to elucidate the methanol dissimilation pathway of B. methanolicus. Growth rates of B. methanolicus MGA3 were assessed on methanol, mannitol, and glucose as a substrate. B. methanolicus achieved maximum growth rate, µmax, when growing on 25 mM methanol, 0.65±0.007 h-1, and it gradually decreased to 0.231±0.004 h-1 at 2Mmethanol concentration which demonstrates substrate inhibition. The maximum growth rates (µmax) of B. methanolicus MGA3 on mannitol and glucose are 0.532±0.002 and 0.336±0.003 h-1, respectively. -

Planctomycetes Attached to Algal Surfaces: Insight Into Their Genomes
MSc 2.º CICLO FCUP 2015 into Planctomycetes their Planctomycetes genomes attached attached to algal surfaces: Insight into to algal their genomes surfaces Mafalda Seabra Faria : Insight : Dissertação de Mestrado apresentada à Faculdade de Ciências da Universidade do Porto Laboratório de Ecofisiologia Microbiana da Universidade do Porto Biologia Celular e Molecular 2014/2015 Mafalda Seabra Faria Seabra Mafalda I FCUP Planctomycetes attached to algal surfaces: Insight into their genomes Planctomycetes attached to algal surfaces: Insight into their genomes Mafalda Seabra Faria Mestrado em Biologia Celular e Molecular Biologia 2015 Orientador Olga Maria Oliveira da Silva Lage, Professora Auxiliar, Faculdade de Ciências da Universidade do Porto Co-orientador Jens Harder, Senior Scientist and Professor, Max Planck Institute for Marine Microbiology FCUP II Planctomycetes attached to algal surfaces: Insight into their genomes Todas as correções determinadas pelo júri, e só essas, foram efetuadas. O Presidente do Júri, Porto, ______/______/_________ FCUP III Planctomycetes attached to algal surfaces: Insight into their genomes FCUP IV Planctomycetes attached to algal surfaces: Insight into their genomes “Tell me and I forget, teach me and I may remember, involve me and I learn.” Benjamin Franklin FCUP V Planctomycetes attached to algal surfaces: Insight into their genomes FCUP VI Planctomycetes attached to algal surfaces: Insight into their genomes Acknowledgements Foremost, I would like to express my sincere gratitude to my supervisor Professor -
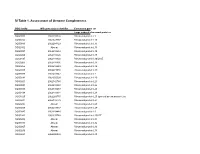
SI Table 1. Assessment of Genome Completeness
SI Table 1. Assessment of Genome Completeness COG family IMG gene object identifier Conserved gene set Large subunit ribosomal proteins COG0081 2062288324 Ribosomal protein L1 COG0244 2062347387 Ribosomal protein L10 COG0080 2062288323 Ribosomal protein L11 COG0102 Absent Ribosomal protein L13 COG0093 2062418832 Ribosomal protein L14 COG0200 2062418826 Ribosomal protein L15 COG0197 2062418838 Ribosomal protein L16/L10E COG0203 2062418836 Ribosomal protein L17 COG0256 2062418829 Ribosomal protein L18 COG0335 2062273558 Ribosomal protein L19 COG0090 2062418842 Ribosomal protein L2 COG0292 2062350539 Ribosomal protein L20 COG0261 2062142780 Ribosomal protein L21 COG0091 2062418840 Ribosomal protein L22 COG0089 2062138283 Ribosomal protein L23 COG0198 2062418834 Ribosomal protein L24 COG1825 2062269715 Ribosomal protein L25 (general stress protein Ctc) COG0211 2062142779 Ribosomal protein L27 COG0227 Absent Ribosomal protein L28 COG0255 2062418837 Ribosomal protein L29 COG0087 2062154483 Ribosomal protein L3 COG1841 2062335748 Ribosomal protein L30/L7E COG0254 Absent Ribosomal protein L31 COG0333 Absent Ribosomal protein L32 COG0267 Absent Ribosomal protein L33 COG0230 Absent Ribosomal protein L34 COG0291 2062350538 Ribosomal protein L35 COG0257 Absent Ribosomal protein L36 COG0088 2062138282 Ribosomal protein L4 COG0094 2062418833 Ribosomal protein L5 COG0097 2062418830 Ribosomal protein L6P/L9E COG0222 2062288326 Ribosomal protein L7/L12 COG0359 2062209880 Ribosomal protein L9 Small subunit ribosomal proteins COG0539 Absent Ribosomal protein