Tnik Are Novel Interaction Partners of Cnk2
Total Page:16
File Type:pdf, Size:1020Kb

Load more
Recommended publications
-

Mouse Tnik Conditional Knockout Project (CRISPR/Cas9)
https://www.alphaknockout.com Mouse Tnik Conditional Knockout Project (CRISPR/Cas9) Objective: To create a Tnik conditional knockout Mouse model (C57BL/6J) by CRISPR/Cas-mediated genome engineering. Strategy summary: The Tnik gene (NCBI Reference Sequence: NM_026910 ; Ensembl: ENSMUSG00000027692 ) is located on Mouse chromosome 3. 33 exons are identified, with the ATG start codon in exon 1 and the TAA stop codon in exon 33 (Transcript: ENSMUST00000160307). Exon 7 will be selected as conditional knockout region (cKO region). Deletion of this region should result in the loss of function of the Mouse Tnik gene. To engineer the targeting vector, homologous arms and cKO region will be generated by PCR using BAC clone RP23-433L13 as template. Cas9, gRNA and targeting vector will be co-injected into fertilized eggs for cKO Mouse production. The pups will be genotyped by PCR followed by sequencing analysis. Note: Mice homozygous for a knock-out allele exhibit impaired postsynaptic signaling and cognitive function. Exon 7 starts from about 12.48% of the coding region. The knockout of Exon 7 will result in frameshift of the gene. The size of intron 6 for 5'-loxP site insertion: 2463 bp, and the size of intron 7 for 3'-loxP site insertion: 12829 bp. The size of effective cKO region: ~631 bp. The cKO region does not have any other known gene. Page 1 of 8 https://www.alphaknockout.com Overview of the Targeting Strategy Wildtype allele gRNA region 5' gRNA region 3' 1 7 33 Targeting vector Targeted allele Constitutive KO allele (After Cre recombination) Legends Exon of mouse Tnik Homology arm cKO region loxP site Page 2 of 8 https://www.alphaknockout.com Overview of the Dot Plot Window size: 10 bp Forward Reverse Complement Sequence 12 Note: The sequence of homologous arms and cKO region is aligned with itself to determine if there are tandem repeats. -

The Essential Role of TNIK Gene Amplification in Gastric Cancer Growth
OPEN Citation: Oncogenesis (2014) 3, e89; doi:10.1038/oncsis.2014.2 & 2014 Macmillan Publishers Limited All rights reserved 2157-9024/14 www.nature.com/oncsis ORIGINAL ARTICLE The essential role of TNIK gene amplification in gastric cancer growth This article has been corrected since Online Publication and an erratum has also been published D-H Yu1,5, X Zhang1,5, H Wang1,2,5, L Zhang3, H Chen4,MHu1, Z Dong1, G Zhu1, Z Qian1,JFan1,XSu1,YXu1, L Zheng1, H Dong1, X Yin1,QJi1 and J Ji3 Traf2- and Nck-interacting kinase (TNIK) is one of the germinal center kinase family members involved in cytoskeleton organization and neuronal dendrite extension. Emerging evidence supports that TNIK is essential for activation of WNT signaling pathway in colon cancer growth. To search for novel genetic aberrations that drive carcinogenesis, we performed microarray-based comparative hybridization assay for gene copy number variations in primary tumor samples. Our data showed that TNIK gene was amplified in 7% (8/106) of Chinese gastric cancer patients. Theses amplifications were confirmed by fluorescence in situ hybridization analysis. PAMC82 human gastric cancer and T47D human breast cancer cell lines with TNIK amplification were identified to further understand the function of TNIK gene amplification. RNA-interference-mediated silencing of TNIK resulted in significant inhibition of cell growth and induction of cell death in TNIK-amplified, but not in TNIK-non-amplified, cell lines tested. This selective sensitivity to the TNIK inhibition was also observed under the effect of a small-molecule TNIK inhibitor. Furthermore, our data indicated that TNIK’s role in gastric cancer growth was not dependent on Wnt signaling but rather was involved in AKT activation and cell autophagy. -
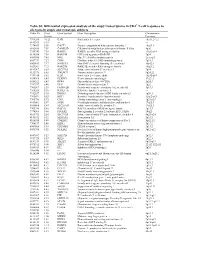
Table S1| Differential Expression Analysis of the Atopy Transcriptome
Table S1| Differential expression analysis of the atopy transcriptome in CD4+ T-cell responses to allergens in atopic and nonatopic subjects Probe ID S.test Gene Symbol Gene Description Chromosome Statistic Location 7994280 10.32 IL4R Interleukin 4 receptor 16p11.2-12.1 8143383 8.95 --- --- --- 7974689 8.50 DACT1 Dapper, antagonist of beta-catenin, homolog 1 14q23.1 8102415 7.59 CAMK2D Calcium/calmodulin-dependent protein kinase II delta 4q26 7950743 7.58 RAB30 RAB30, member RAS oncogene family 11q12-q14 8136580 7.54 RAB19B GTP-binding protein RAB19B 7q34 8043504 7.45 MAL Mal, T-cell differentiation protein 2cen-q13 8087739 7.27 CISH Cytokine inducible SH2-containing protein 3p21.3 8000413 7.17 NSMCE1 Non-SMC element 1 homolog (S. cerevisiae) 16p12.1 8021301 7.15 RAB27B RAB27B, member RAS oncogene family 18q21.2 8143367 6.83 SLC37A3 Solute carrier family 37 member 3 7q34 8152976 6.65 TMEM71 Transmembrane protein 71 8q24.22 7931914 6.56 IL2R Interleukin 2 receptor, alpha 10p15-p14 8014768 6.43 PLXDC1 Plexin domain containing 1 17q21.1 8056222 6.43 DPP4 Dipeptidyl-peptidase 4 (CD26) 2q24.3 7917697 6.40 GFI1 Growth factor independent 1 1p22 7903507 6.39 FAM102B Family with sequence similarity 102, member B 1p13.3 7968236 5.96 RASL11A RAS-like, family 11, member A --- 7912537 5.95 DHRS3 Dehydrogenase/reductase (SDR family) member 3 1p36.1 7963491 5.83 KRT1 Keratin 1 (epidermolytic hyperkeratosis) 12q12-q13 7903786 5.72 CSF1 Colony stimulating factor 1 (macrophage) 1p21-p13 8019061 5.67 SGSH N-sulfoglucosamine sulfohydrolase (sulfamidase) 17q25.3 -

Convergent Functional Genomics of Schizophrenia: from Comprehensive Understanding to Genetic Risk Prediction
Molecular Psychiatry (2012) 17, 887 -- 905 & 2012 Macmillan Publishers Limited All rights reserved 1359-4184/12 www.nature.com/mp IMMEDIATE COMMUNICATION Convergent functional genomics of schizophrenia: from comprehensive understanding to genetic risk prediction M Ayalew1,2,9, H Le-Niculescu1,9, DF Levey1, N Jain1, B Changala1, SD Patel1, E Winiger1, A Breier1, A Shekhar1, R Amdur3, D Koller4, JI Nurnberger1, A Corvin5, M Geyer6, MT Tsuang6, D Salomon7, NJ Schork7, AH Fanous3, MC O’Donovan8 and AB Niculescu1,2 We have used a translational convergent functional genomics (CFG) approach to identify and prioritize genes involved in schizophrenia, by gene-level integration of genome-wide association study data with other genetic and gene expression studies in humans and animal models. Using this polyevidence scoring and pathway analyses, we identify top genes (DISC1, TCF4, MBP, MOBP, NCAM1, NRCAM, NDUFV2, RAB18, as well as ADCYAP1, BDNF, CNR1, COMT, DRD2, DTNBP1, GAD1, GRIA1, GRIN2B, HTR2A, NRG1, RELN, SNAP-25, TNIK), brain development, myelination, cell adhesion, glutamate receptor signaling, G-protein-- coupled receptor signaling and cAMP-mediated signaling as key to pathophysiology and as targets for therapeutic intervention. Overall, the data are consistent with a model of disrupted connectivity in schizophrenia, resulting from the effects of neurodevelopmental environmental stress on a background of genetic vulnerability. In addition, we show how the top candidate genes identified by CFG can be used to generate a genetic risk prediction score (GRPS) to aid schizophrenia diagnostics, with predictive ability in independent cohorts. The GRPS also differentiates classic age of onset schizophrenia from early onset and late-onset disease. -

X-Linked Diseases: Susceptible Females
REVIEW ARTICLE X-linked diseases: susceptible females Barbara R. Migeon, MD 1 The role of X-inactivation is often ignored as a prime cause of sex data include reasons why women are often protected from the differences in disease. Yet, the way males and females express their deleterious variants carried on their X chromosome, and the factors X-linked genes has a major role in the dissimilar phenotypes that that render women susceptible in some instances. underlie many rare and common disorders, such as intellectual deficiency, epilepsy, congenital abnormalities, and diseases of the Genetics in Medicine (2020) 22:1156–1174; https://doi.org/10.1038/s41436- heart, blood, skin, muscle, and bones. Summarized here are many 020-0779-4 examples of the different presentations in males and females. Other INTRODUCTION SEX DIFFERENCES ARE DUE TO X-INACTIVATION Sex differences in human disease are usually attributed to The sex differences in the effect of X-linked pathologic variants sex specific life experiences, and sex hormones that is due to our method of X chromosome dosage compensation, influence the function of susceptible genes throughout the called X-inactivation;9 humans and most placental mammals – genome.1 5 Such factors do account for some dissimilarities. compensate for the sex difference in number of X chromosomes However, a major cause of sex-determined expression of (that is, XX females versus XY males) by transcribing only one disease has to do with differences in how males and females of the two female X chromosomes. X-inactivation silences all X transcribe their gene-rich human X chromosomes, which is chromosomes but one; therefore, both males and females have a often underappreciated as a cause of sex differences in single active X.10,11 disease.6 Males are the usual ones affected by X-linked For 46 XY males, that X is the only one they have; it always pathogenic variants.6 Females are biologically superior; a comes from their mother, as fathers contribute their Y female usually has no disease, or much less severe disease chromosome. -

Supplementary Table 2
Supplementary Table 2. Differentially Expressed Genes following Sham treatment relative to Untreated Controls Fold Change Accession Name Symbol 3 h 12 h NM_013121 CD28 antigen Cd28 12.82 BG665360 FMS-like tyrosine kinase 1 Flt1 9.63 NM_012701 Adrenergic receptor, beta 1 Adrb1 8.24 0.46 U20796 Nuclear receptor subfamily 1, group D, member 2 Nr1d2 7.22 NM_017116 Calpain 2 Capn2 6.41 BE097282 Guanine nucleotide binding protein, alpha 12 Gna12 6.21 NM_053328 Basic helix-loop-helix domain containing, class B2 Bhlhb2 5.79 NM_053831 Guanylate cyclase 2f Gucy2f 5.71 AW251703 Tumor necrosis factor receptor superfamily, member 12a Tnfrsf12a 5.57 NM_021691 Twist homolog 2 (Drosophila) Twist2 5.42 NM_133550 Fc receptor, IgE, low affinity II, alpha polypeptide Fcer2a 4.93 NM_031120 Signal sequence receptor, gamma Ssr3 4.84 NM_053544 Secreted frizzled-related protein 4 Sfrp4 4.73 NM_053910 Pleckstrin homology, Sec7 and coiled/coil domains 1 Pscd1 4.69 BE113233 Suppressor of cytokine signaling 2 Socs2 4.68 NM_053949 Potassium voltage-gated channel, subfamily H (eag- Kcnh2 4.60 related), member 2 NM_017305 Glutamate cysteine ligase, modifier subunit Gclm 4.59 NM_017309 Protein phospatase 3, regulatory subunit B, alpha Ppp3r1 4.54 isoform,type 1 NM_012765 5-hydroxytryptamine (serotonin) receptor 2C Htr2c 4.46 NM_017218 V-erb-b2 erythroblastic leukemia viral oncogene homolog Erbb3 4.42 3 (avian) AW918369 Zinc finger protein 191 Zfp191 4.38 NM_031034 Guanine nucleotide binding protein, alpha 12 Gna12 4.38 NM_017020 Interleukin 6 receptor Il6r 4.37 AJ002942 -

The Proteomic Landscape of Resting and Activated CD4+ T Cells Reveal Insights Into Cell Differentiation and Function
International Journal of Molecular Sciences Article The Proteomic Landscape of Resting and Activated CD4+ T Cells Reveal Insights into Cell Differentiation and Function Yashwanth Subbannayya 1 , Markus Haug 1, Sneha M. Pinto 1, Varshasnata Mohanty 2, Hany Zakaria Meås 1, Trude Helen Flo 1, T.S. Keshava Prasad 2 and Richard K. Kandasamy 1,* 1 Centre of Molecular Inflammation Research (CEMIR), Department of Clinical and Molecular Medicine (IKOM), Norwegian University of Science and Technology, 7491 Trondheim, Norway; [email protected] (Y.S.); [email protected] (M.H.); [email protected] (S.M.P.); [email protected] (H.Z.M.); trude.fl[email protected] (T.H.F.) 2 Center for Systems Biology and Molecular Medicine, Yenepoya (Deemed to be University), Mangalore 575018, India; [email protected] (V.M.); [email protected] (T.S.K.P.) * Correspondence: [email protected] Abstract: CD4+ T cells (T helper cells) are cytokine-producing adaptive immune cells that activate or regulate the responses of various immune cells. The activation and functional status of CD4+ T cells is important for adequate responses to pathogen infections but has also been associated with auto-immune disorders and survival in several cancers. In the current study, we carried out a label-free high-resolution FTMS-based proteomic profiling of resting and T cell receptor-activated (72 h) primary human CD4+ T cells from peripheral blood of healthy donors as well as SUP-T1 cells. We identified 5237 proteins, of which significant alterations in the levels of 1119 proteins were observed between resting and activated CD4+ T cells. -

Exons Deletion of CNKSR2 Gene Identified in X-Linked Syndromic
Daoqi et al. BMC Medical Genetics (2020) 21:69 https://doi.org/10.1186/s12881-020-01004-2 CASE REPORT Open Access Exons deletion of CNKSR2 gene identified in X-linked syndromic intellectual disability Mei Daoqi1, Chen Guohong1, Wang Yuan1, Yang Zhixiao1, Xu Kaili1 and Mei Shiyue2* Abstract Background: The Houge type of X-linked syndromic mental retardation is an X-linked intellectual disability (XLID) recently recorded in the Online Mendelian Inheritance in Man (OMIM) and only 8 cases have been reported in literature thus far. Case presentation: We present two brothers with intractable seizures and syndromic intellectual disability with symptoms consisting of delayed development, intellectual disability, and speech and language delay. The mother was a symptomatic carrier with milder clinical phenotype. Whole exome sequencing identified a small fragment deletion spanning four exons, about 9.5 kilobases (kb) in length in the CNKSR2 gene in the patients. The mutation co-segregation revealed that exon deletions occurred de novo in the proband’s mother. Conclusion: Although large deletions have been reported, no small deletions have yet been identified. In this case report, we identified a small deletion in the CNKSR2 gene. This study enhances our knowledge of the CNKSR2 gene mutation spectrum and provides further information about the phenotypic characteristics of X-linked syndromic intellectual disability. Keywords: X-linked syndromic mental retardation, Intractable seizures, CNKSR2 gene, Exons deletion Background (#301008) in 2017. MRXSHG is caused by a deficiency X-linked intellectual disability (XLID), also known as of the CNKSR2 gene and only 8 cases have been X-linked mental retardation (XLMR), refers to a gener- reported in literature. -

Molecular Signatures Differentiate Immune States in Type 1 Diabetes Families
Page 1 of 65 Diabetes Molecular signatures differentiate immune states in Type 1 diabetes families Yi-Guang Chen1, Susanne M. Cabrera1, Shuang Jia1, Mary L. Kaldunski1, Joanna Kramer1, Sami Cheong2, Rhonda Geoffrey1, Mark F. Roethle1, Jeffrey E. Woodliff3, Carla J. Greenbaum4, Xujing Wang5, and Martin J. Hessner1 1The Max McGee National Research Center for Juvenile Diabetes, Children's Research Institute of Children's Hospital of Wisconsin, and Department of Pediatrics at the Medical College of Wisconsin Milwaukee, WI 53226, USA. 2The Department of Mathematical Sciences, University of Wisconsin-Milwaukee, Milwaukee, WI 53211, USA. 3Flow Cytometry & Cell Separation Facility, Bindley Bioscience Center, Purdue University, West Lafayette, IN 47907, USA. 4Diabetes Research Program, Benaroya Research Institute, Seattle, WA, 98101, USA. 5Systems Biology Center, the National Heart, Lung, and Blood Institute, the National Institutes of Health, Bethesda, MD 20824, USA. Corresponding author: Martin J. Hessner, Ph.D., The Department of Pediatrics, The Medical College of Wisconsin, Milwaukee, WI 53226, USA Tel: 011-1-414-955-4496; Fax: 011-1-414-955-6663; E-mail: [email protected]. Running title: Innate Inflammation in T1D Families Word count: 3999 Number of Tables: 1 Number of Figures: 7 1 For Peer Review Only Diabetes Publish Ahead of Print, published online April 23, 2014 Diabetes Page 2 of 65 ABSTRACT Mechanisms associated with Type 1 diabetes (T1D) development remain incompletely defined. Employing a sensitive array-based bioassay where patient plasma is used to induce transcriptional responses in healthy leukocytes, we previously reported disease-specific, partially IL-1 dependent, signatures associated with pre and recent onset (RO) T1D relative to unrelated healthy controls (uHC). -

Stargazin and Other Transmembrane AMPA Receptor Regulating Proteins Interact with Synaptic Scaffolding Protein MAGI-2 in Brain
The Journal of Neuroscience, July 26, 2006 • 26(30):7875–7884 • 7875 Cellular/Molecular Stargazin and Other Transmembrane AMPA Receptor Regulating Proteins Interact with Synaptic Scaffolding Protein MAGI-2 in Brain Fang Deng, Maureen G. Price, Caleb F. Davis, Mayra Mori, and Daniel L. Burgess Department of Neurology, Baylor College of Medicine, Houston, Texas 77030 The spatial coordination of neurotransmitter receptors with other postsynaptic signaling and structural molecules is regulated by a diverse array of cell-specific scaffolding proteins. The synaptic trafficking of AMPA receptors by the stargazin protein in some neurons, for example, depends on specific interactions between the C terminus of stargazin and the PDZ [postsynaptic density-95 (PSD-95)/Discs large/zona occludens-1] domains of membrane-associated guanylate kinase scaffolding proteins PSD-93 or PSD-95. Stargazin [Cacng2 (Ca 2ϩ channel ␥2 subunit)] is one of four closely related proteins recently categorized as transmembrane AMPA receptor regulating proteins (TARPs) that appear to share similar functions but exhibit distinct expression patterns in the CNS. We used yeast two-hybrid screeningtoidentifyMAGI-2(membraneassociatedguanylatekinase,WWandPDZdomaincontaining2)asanovelcandidateinteractor with the cytoplasmic C termini of the TARPs. MAGI-2 [also known as S-SCAM (synaptic scaffolding molecule)] is a multi-PDZ domain scaffolding protein that interacts with several different ligands in brain, including PTEN (phosphatase and tensin homolog), dasm1 (dendrite arborization and synapse maturation 1), dendrin, axin, - and ␦-catenin, neuroligin, hyperpolarization-activated cation channels, 1-adrenergic receptors, and NMDA receptors. We confirmed that MAGI-2 coimmunoprecipitated with stargazin in vivo from mouse cerebral cortex and used in vitro assays to localize the interaction to the C-terminal -TTPV amino acid motif of stargazin and the PDZ1, PDZ3, and PDZ5 domains of MAGI-2. -

Downloaded from Ftp://Ftp.Uniprot.Org/ on July 3, 2019) Using Maxquant (V1.6.10.43) Search Algorithm
bioRxiv preprint doi: https://doi.org/10.1101/2020.11.17.385096; this version posted November 17, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-ND 4.0 International license. The proteomic landscape of resting and activated CD4+ T cells reveal insights into cell differentiation and function Yashwanth Subbannayya1, Markus Haug1, Sneha M. Pinto1, Varshasnata Mohanty2, Hany Zakaria Meås1, Trude Helen Flo1, T.S. Keshava Prasad2 and Richard K. Kandasamy1,* 1Centre of Molecular Inflammation Research (CEMIR), and Department of Clinical and Molecular Medicine (IKOM), Norwegian University of Science and Technology, N-7491 Trondheim, Norway 2Center for Systems Biology and Molecular Medicine, Yenepoya (Deemed to be University), Mangalore, India *Correspondence to: Professor Richard Kumaran Kandasamy Norwegian University of Science and Technology (NTNU) Centre of Molecular Inflammation Research (CEMIR) PO Box 8905 MTFS Trondheim 7491 Norway E-mail: [email protected] (Kandasamy R K) Tel.: +47-7282-4511 1 bioRxiv preprint doi: https://doi.org/10.1101/2020.11.17.385096; this version posted November 17, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-ND 4.0 International license. Abstract CD4+ T cells (T helper cells) are cytokine-producing adaptive immune cells that activate or regulate the responses of various immune cells. -

Centre for Arab Genomic Studies a Division of Sheikh Hamdan Award for Medical Sciences
Centre for Arab Genomic Studies A Division of Sheikh Hamdan Award for Medical Sciences The atalogue for ransmission enetics in rabs C T G A CTGA Database TRAF2- and NCK-Interacting Kinase Alternative Names spanning about 399 kb within the genomic DNA. TNIK The encoded protein comprises 1360 amino acids KIAA0551 with a molecular weight of 155 kDa. Like other members of the germinal center kinase (GCK) Record Category proteins family, TNIK is also characterized by the Gene locus presence of an N-terminal kinase domain and a regulatory C-terminal GCK domain. The protein is WHO-ICD expressed ubiquitously, but maximum expression N.B.: Classification not applicable to gene loci. levels are seen in heart, brain and skeletal muscle cells, as well as in colorectal cancers. Incidence per 100,000 Live Births N/A to gene loci Epidemiology in the Arab World OMIM Number Saudi Arabia 610005 Anazi et al., (2016) reported on two multiplex consanguineous Saudi family siblings affected with Mode of Inheritance autosomal recessive intellectual disability. The first N/A family consisted of two affected and two normal siblings, while the second family had three affected Gene Map Locus and four normal siblings. Autozygome analysis on 3q26.2-q26.3 the first family revealed six autozygous intervals that were exclusive to the two affected brothers. Description Within the second family, only one autozygous The TNIK gene codes for a member of the block exclusive to the affected patients was serine/threonine kinase of the Ste20 family of It identified. Interestingly, this block on chromosome belongs to the germinal center kinase subgroup of 3 was also one of the three identified in the first this family that is involved in cytoskeleton family.