Oct4-Mediated Reprogramming Induces Embryonic-Like Microrna
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Network Assessment of Demethylation Treatment in Melanoma: Differential Transcriptome-Methylome and Antigen Profile Signatures
RESEARCH ARTICLE Network assessment of demethylation treatment in melanoma: Differential transcriptome-methylome and antigen profile signatures Zhijie Jiang1☯, Caterina Cinti2☯, Monia Taranta2, Elisabetta Mattioli3,4, Elisa Schena3,5, Sakshi Singh2, Rimpi Khurana1, Giovanna Lattanzi3,4, Nicholas F. Tsinoremas1,6, 1 Enrico CapobiancoID * a1111111111 1 Center for Computational Science, University of Miami, Miami, FL, United States of America, 2 Institute of Clinical Physiology, CNR, Siena, Italy, 3 CNR Institute of Molecular Genetics, Bologna, Italy, 4 IRCCS Rizzoli a1111111111 Orthopedic Institute, Bologna, Italy, 5 Endocrinology Unit, Department of Medical & Surgical Sciences, Alma a1111111111 Mater Studiorum University of Bologna, S Orsola-Malpighi Hospital, Bologna, Italy, 6 Department of a1111111111 Medicine, University of Miami, Miami, FL, United States of America a1111111111 ☯ These authors contributed equally to this work. * [email protected] OPEN ACCESS Abstract Citation: Jiang Z, Cinti C, Taranta M, Mattioli E, Schena E, Singh S, et al. (2018) Network assessment of demethylation treatment in Background melanoma: Differential transcriptome-methylome and antigen profile signatures. PLoS ONE 13(11): In melanoma, like in other cancers, both genetic alterations and epigenetic underlie the met- e0206686. https://doi.org/10.1371/journal. astatic process. These effects are usually measured by changes in both methylome and pone.0206686 transcriptome profiles, whose cross-correlation remains uncertain. We aimed to assess at Editor: Roger Chammas, Universidade de Sao systems scale the significance of epigenetic treatment in melanoma cells with different met- Paulo, BRAZIL astatic potential. Received: June 20, 2018 Accepted: October 17, 2018 Methods and findings Published: November 28, 2018 Treatment by DAC demethylation with 5-Aza-2'-deoxycytidine of two melanoma cell lines Copyright: © 2018 Jiang et al. -

CRB3 Antibody (Center) Blocking Peptide Synthetic Peptide Catalog # Bp17560c
10320 Camino Santa Fe, Suite G San Diego, CA 92121 Tel: 858.875.1900 Fax: 858.622.0609 CRB3 Antibody (Center) Blocking Peptide Synthetic peptide Catalog # BP17560c Specification CRB3 Antibody (Center) Blocking Peptide - CRB3 Antibody (Center) Blocking Peptide - Background Product Information This gene encodes a member of the Crumbs Primary Accession Q9BUF7 family ofproteins. This protein may play a role in epithelial cell polarityand is associated with tight junctions at the apical surface ofepithelial CRB3 Antibody (Center) Blocking Peptide - Additional Information cells. Alternate transcriptional splice variants,encoding different isoforms, have been characterized. [provided byRefSeq]. Gene ID 92359 CRB3 Antibody (Center) Blocking Peptide - Other Names References Protein crumbs homolog 3, CRB3 Pardossi-Piquard, R., et al. Biochemistry Format 46(48):13704-13710(2007)Fan, S., et al. J. Cell Peptides are lyophilized in a solid powder Biol. 178(3):387-398(2007)Fogg, V.C., et al. J. format. Peptides can be reconstituted in solution using the appropriate buffer as Cell. Sci. 118 (PT 13), 2859-2869 (2005) needed. :Lemmers, C., et al. Mol. Biol. Cell 15(3):1324-1333(2004)Roh, M.H., et al. J. Cell. Storage Sci. 116 (PT 14), 2895-2906 (2003) : Maintain refrigerated at 2-8°C for up to 6 months. For long term storage store at -20°C. Precautions This product is for research use only. Not for use in diagnostic or therapeutic procedures. CRB3 Antibody (Center) Blocking Peptide - Protein Information Name CRB3 (HGNC:20237) Function Involved in the establishment of cell polarity in mammalian epithelial cells (PubMed:<a href="http://www.uniprot.org/c itations/12771187" target="_blank">12771187</a>, PubMed:<a href="http://www.uniprot.org/ci tations/14718572" target="_blank">14718572</a>). -

CRB3 Polyclonal Antibody Purified Rabbit Polyclonal Antibody (Pab) Catalog # AP55396
10320 Camino Santa Fe, Suite G San Diego, CA 92121 Tel: 858.875.1900 Fax: 858.622.0609 CRB3 Polyclonal Antibody Purified Rabbit Polyclonal Antibody (Pab) Catalog # AP55396 Specification CRB3 Polyclonal Antibody - Product Information Application WB, IHC-P, IHC-F, IF, ICC Primary Accession Q9BUF7 Host Rabbit Clonality Polyclonal Calculated MW 12854 CRB3 Polyclonal Antibody - Additional Information Gene ID 92359 Other Names Protein crumbs homolog 3, CRB3 (<a href= "http://www.genenames.org/cgi-bin/gene_sy mbol_report?hgnc_id=20237" target="_blank">HGNC:20237</a>) Format 0.01M TBS(pH7.4) with 1% BSA, 0.09% (W/V) sodium azide and 50% Glyce Storage Store at -20 ℃ for one year. Avoid repeated freeze/thaw cycles. When reconstituted in sterile pH 7.4 0.01M PBS or diluent of antibody the antibody is stable for at least two weeks at 2-4 ℃. CRB3 Polyclonal Antibody - Protein Information Name CRB3 (HGNC:20237) Function Involved in the establishment of cell polarity in mammalian epithelial cells (PubMed:<a href="http://www.uniprot.org/c itations/12771187" target="_blank">12771187</a>, PubMed:<a href="http://www.uniprot.org/ci tations/14718572" target="_blank">14718572</a>). Regulates the morphogenesis of tight Page 1/2 10320 Camino Santa Fe, Suite G San Diego, CA 92121 Tel: 858.875.1900 Fax: 858.622.0609 junctions (PubMed:<a href="http://www.uni prot.org/citations/12771187" target="_blank">12771187</a>, PubMed:<a href="http://www.uniprot.org/ci tations/14718572" target="_blank">14718572</a>). Involved in promoting phosphorylation and cytoplasmic retention of transcriptional coactivators YAP1 and WWTR1/TAZ which leads to suppression of TGFB1-dependent transcription of target genes such as CCN2/CTGF, SERPINE1/PAI1, SNAI1/SNAIL1 and SMAD7 (By similarity). -

Positive Biofeedback of Crb3 Protein Favouring Notch Mediated Tumour
International Journal of Applied Research 2016; 2(2): 370-381 ISSN Print: 2394-7500 ISSN Online: 2394-5869 Positive biofeedback of Crb3 protein favouring notch Impact Factor: 5.2 IJAR 2016; 2(2): 370-381 mediated tumour suppression in human body: A www.allresearchjournal.com Received: 16-12-2015 glimpse for better understanding of cancer biology Accepted: 18-01-2016 and to identify potential therapeutic targets Partha Majumder Biomedical Scientist, Former Head, Department of Partha Majumder, Anjana Mazumdar Applied Biotechnology & Bioinformatics, Sikkim Manipal University, CC: 1637, Abstract Kolkata, India. Our study is a comprehensive approach of a biofeedback paying regards to homeostasis of human body. A biofeedback system involves a sensory organ and an appropriate stimulus. The stimulus is mediated Anjana Mazumdar through organs derived from specific biosensors. Homeostasis is the set of processes by which constant Professor, or static conditions are maintained within the internal environment of a subject and therefore a Department of Oral Pathology, homeostat is a controller involved in maintaining homeostasis. Paying regards to extensive study on Dr. R. Ahmed Govt. Dental Cytology, we found that, Cell polarity is essential in many biological processes and required for College, Kolkata, India. development as well as maintenance of tissue integrity. Loss of polarity is considered both a hallmark and precondition for human cancer. According to our knowledge, three conserved polarity protein complexes regulate different modes of polarity that are conserved throughout numerous cell types and species. These complexes are the Crumbs, Par and Scribble complex. Defects in apical-basal polarity regulation are associated with tissue overgrowth and tumorigenesis. -

Cystatin-B Negatively Regulates the Malignant Characteristics of Oral Squamous Cell Carcinoma Possibly Via the Epithelium Proliferation/ Differentiation Program
ORIGINAL RESEARCH published: 24 August 2021 doi: 10.3389/fonc.2021.707066 Cystatin-B Negatively Regulates the Malignant Characteristics of Oral Squamous Cell Carcinoma Possibly Via the Epithelium Proliferation/ Differentiation Program Tian-Tian Xu 1, Xiao-Wen Zeng 1, Xin-Hong Wang 2, Lu-Xi Yang 1, Gang Luo 1* and Ting Yu 1* 1 Department of Periodontics, Affiliated Stomatology Hospital of Guangzhou Medical University, Guangzhou Key Laboratory of Basic and Applied Research of Oral Regenerative Medicine, Guangzhou, China, 2 Department of Oral Pathology and Medicine, Affiliated Stomatology Hospital of Guangzhou Medical University, Guangzhou Key Laboratory of Basic and Applied Research of Oral Regenerative Medicine, Guangzhou, China Disturbance in the proteolytic process is one of the malignant signs of tumors. Proteolysis Edited by: Eva Csosz, is highly orchestrated by cysteine cathepsin and its inhibitors. Cystatin-B (CSTB) is a University of Debrecen, Hungary general cysteine cathepsin inhibitor that prevents cysteine cathepsin from leaking from Reviewed by: lysosomes and causing inappropriate proteolysis. Our study found that CSTB was Csongor Kiss, downregulated in both oral squamous cell carcinoma (OSCC) tissues and cells University of Debrecen, Hungary Gergely Nagy, compared with normal controls. Immunohistochemical analysis showed that CSTB was University of Debrecen, Hungary mainly distributed in the epithelial structure of OSCC tissues, and its expression intensity *Correspondence: was related to the grade classification. A correlation analysis between CSTB and clinical Gang Luo [email protected] prognosis was performed using gene expression data and clinical information acquired Ting Yu from The Cancer Genome Atlas (TCGA) database. Patients with lower expression levels of [email protected] CSTB had shorter disease-free survival times and poorer clinicopathological features Specialty section: (e.g., lymph node metastases, perineural invasion, low degree of differentiation, and This article was submitted to advanced tumor stage). -

393LN V 393P 344SQ V 393P Probe Set Entrez Gene
393LN v 393P 344SQ v 393P Entrez fold fold probe set Gene Gene Symbol Gene cluster Gene Title p-value change p-value change chemokine (C-C motif) ligand 21b /// chemokine (C-C motif) ligand 21a /// chemokine (C-C motif) ligand 21c 1419426_s_at 18829 /// Ccl21b /// Ccl2 1 - up 393 LN only (leucine) 0.0047 9.199837 0.45212 6.847887 nuclear factor of activated T-cells, cytoplasmic, calcineurin- 1447085_s_at 18018 Nfatc1 1 - up 393 LN only dependent 1 0.009048 12.065 0.13718 4.81 RIKEN cDNA 1453647_at 78668 9530059J11Rik1 - up 393 LN only 9530059J11 gene 0.002208 5.482897 0.27642 3.45171 transient receptor potential cation channel, subfamily 1457164_at 277328 Trpa1 1 - up 393 LN only A, member 1 0.000111 9.180344 0.01771 3.048114 regulating synaptic membrane 1422809_at 116838 Rims2 1 - up 393 LN only exocytosis 2 0.001891 8.560424 0.13159 2.980501 glial cell line derived neurotrophic factor family receptor alpha 1433716_x_at 14586 Gfra2 1 - up 393 LN only 2 0.006868 30.88736 0.01066 2.811211 1446936_at --- --- 1 - up 393 LN only --- 0.007695 6.373955 0.11733 2.480287 zinc finger protein 1438742_at 320683 Zfp629 1 - up 393 LN only 629 0.002644 5.231855 0.38124 2.377016 phospholipase A2, 1426019_at 18786 Plaa 1 - up 393 LN only activating protein 0.008657 6.2364 0.12336 2.262117 1445314_at 14009 Etv1 1 - up 393 LN only ets variant gene 1 0.007224 3.643646 0.36434 2.01989 ciliary rootlet coiled- 1427338_at 230872 Crocc 1 - up 393 LN only coil, rootletin 0.002482 7.783242 0.49977 1.794171 expressed sequence 1436585_at 99463 BB182297 1 - up 393 -

Predict AID Targeting in Non-Ig Genes Multiple Transcription Factor
Downloaded from http://www.jimmunol.org/ by guest on September 26, 2021 is online at: average * The Journal of Immunology published online 20 March 2013 from submission to initial decision 4 weeks from acceptance to publication Multiple Transcription Factor Binding Sites Predict AID Targeting in Non-Ig Genes Jamie L. Duke, Man Liu, Gur Yaari, Ashraf M. Khalil, Mary M. Tomayko, Mark J. Shlomchik, David G. Schatz and Steven H. Kleinstein J Immunol http://www.jimmunol.org/content/early/2013/03/20/jimmun ol.1202547 Submit online. Every submission reviewed by practicing scientists ? is published twice each month by http://jimmunol.org/subscription Submit copyright permission requests at: http://www.aai.org/About/Publications/JI/copyright.html Receive free email-alerts when new articles cite this article. Sign up at: http://jimmunol.org/alerts http://www.jimmunol.org/content/suppl/2013/03/20/jimmunol.120254 7.DC1 Information about subscribing to The JI No Triage! Fast Publication! Rapid Reviews! 30 days* Why • • • Material Permissions Email Alerts Subscription Supplementary The Journal of Immunology The American Association of Immunologists, Inc., 1451 Rockville Pike, Suite 650, Rockville, MD 20852 Copyright © 2013 by The American Association of Immunologists, Inc. All rights reserved. Print ISSN: 0022-1767 Online ISSN: 1550-6606. This information is current as of September 26, 2021. Published March 20, 2013, doi:10.4049/jimmunol.1202547 The Journal of Immunology Multiple Transcription Factor Binding Sites Predict AID Targeting in Non-Ig Genes Jamie L. Duke,* Man Liu,†,1 Gur Yaari,‡ Ashraf M. Khalil,x Mary M. Tomayko,{ Mark J. Shlomchik,†,x David G. -

Whole-Genome DNA Methylation Profiling Identifies Epigenetic Signatures of Uterine Carcinosarcoma Jing Li Washington University School of Medicine in St
Washington University School of Medicine Digital Commons@Becker Open Access Publications 2017 Whole-genome DNA methylation profiling identifies epigenetic signatures of uterine carcinosarcoma Jing Li Washington University School of Medicine in St. Louis Xiaoyun Xing Washington University School of Medicine in St. Louis Daofeng Li Washington University School of Medicine in St. Louis Bo Zhang Washington University School of Medicine in St. Louis David G. Mutch Washington University School of Medicine in St. Louis See next page for additional authors Follow this and additional works at: https://digitalcommons.wustl.edu/open_access_pubs Recommended Citation Li, Jing; Xing, Xiaoyun; Li, Daofeng; Zhang, Bo; Mutch, David G.; Hagemann, Ian S.; and Wang, Ting, ,"Whole-genome DNA methylation profiling identifies epigenetic signatures of uterine carcinosarcoma." Neoplasia.19,2. 100-111. (2017). https://digitalcommons.wustl.edu/open_access_pubs/5534 This Open Access Publication is brought to you for free and open access by Digital Commons@Becker. It has been accepted for inclusion in Open Access Publications by an authorized administrator of Digital Commons@Becker. For more information, please contact [email protected]. Authors Jing Li, Xiaoyun Xing, Daofeng Li, Bo Zhang, David G. Mutch, Ian S. Hagemann, and Ting Wang This open access publication is available at Digital Commons@Becker: https://digitalcommons.wustl.edu/open_access_pubs/5534 Volume 19 Number xx Month 2016 pp. 100–111 100 www.neoplasia.com Jing Li*, †, ‡, Xiaoyun Xing*, †, Daofeng Li*, †, Whole-Genome DNA Methylation † Bo Zhang*, , David G. Mutch§, Ian S. Hagemann§, ¶ Profiling Identifies Epigenetic and Ting Wang*, † Signatures of Uterine *Department of Genetics, Washington University School of 1,2 † Carcinosarcoma Medicine, St. -

Discerning the Role of Krüppel-Like Factor 4 in Breast Cancer
DISCERNING THE ROLE OF KRÜPPEL-LIKE FACTOR 4 IN BREAST CANCER by JENNIFER L. YORI Submitted in partial fulfillment of the requirements For the degree of Doctor of Philosophy Dissertation Advisor: Dr. Ruth A. Keri Department of Pharmacology CASE WESTERN RESERVE UNIVERSITY May, 2011 CASE WESTERN RESERVE UNIVERSITY SCHOOL OF GRADUATE STUDIES We hereby approve the thesis/dissertation of ______________________________________________________ candidate for the ________________________________degree *. (signed)_______________________________________________ (chair of the committee) ________________________________________________ ________________________________________________ ________________________________________________ ________________________________________________ ________________________________________________ (date) _______________________ *We also certify that written approval has been obtained for any proprietary material contained therein. DEDICATION This work is dedicatied to Beverly Ann McMillan for all that she taught me and Fiona Campbell Yori for all that she teaches me everyday. iii TABLE OF CONTENTS List of Tables vii List of Figures viii Acknowledgements x Abbreviations xii Abstract 1 Chapter I INTRODUCTION, REVIEW OF THE LITERTURE, AND 3 STATEMENT OF PURPOSE 1.1 Molecular/Histopathological subtypes and cellular origin of 3 breast cancer 1.1.1 Luminal breast cancer 5 1.1.2 HER2/ERRB2 breast cancer 6 1.1.3 Basal-like breast cancer 8 1.1.4 Comparison of mouse models and human tumors generate a 10 new molecular subtype 1.2 -
Primepcr™Assay Validation Report
PrimePCR™Assay Validation Report Gene Information Gene Name crumbs homolog 3 (Drosophila) Gene Symbol CRB3 Organism Human Gene Summary This gene encodes a member of the Crumbs family of proteins. This protein may play a role in epithelial cell polarity and is associated with tight junctions at the apical surface of epithelial cells. Alternate transcriptional splice variants encoding different isoforms have been characterized. Gene Aliases Not Available RefSeq Accession No. NC_000019.9, NT_011255.14 UniGene ID Hs.150319 Ensembl Gene ID ENSG00000130545 Entrez Gene ID 92359 Assay Information Unique Assay ID qHsaCIP0029329 Assay Type Probe - Validation information is for the primer pair using SYBR® Green detection Detected Coding Transcript(s) ENST00000356762, ENST00000308243 Amplicon Context Sequence GGCCGAGCCTGGGGGCAAATACAGACCACTTCTGCAAATGAGAATAGCACTGTT TTGCCTTCATCCACCAGCTCCAGCTCCGATGGCAACCTGCGTCCAGAAGCCATC ACTGCTATCATCGTGGTCTTCTCCCTCTTGGCTGCCT Amplicon Length (bp) 115 Chromosome Location 19:6464776-6466528 Assay Design Intron-spanning Purification Desalted Validation Results Efficiency (%) 96 R2 0.9987 cDNA Cq 24.4 cDNA Tm (Celsius) 85 gDNA Cq 35.32 Page 1/5 PrimePCR™Assay Validation Report Specificity (%) 100 Information to assist with data interpretation is provided at the end of this report. Page 2/5 PrimePCR™Assay Validation Report CRB3, Human Amplification Plot Amplification of cDNA generated from 25 ng of universal reference RNA Melt Peak Melt curve analysis of above amplification Standard Curve Standard curve generated using 20 million copies of template diluted 10-fold to 20 copies Page 3/5 PrimePCR™Assay Validation Report Products used to generate validation data Real-Time PCR Instrument CFX384 Real-Time PCR Detection System Reverse Transcription Reagent iScript™ Advanced cDNA Synthesis Kit for RT-qPCR Real-Time PCR Supermix SsoAdvanced™ SYBR® Green Supermix Experimental Sample qPCR Human Reference Total RNA Data Interpretation Unique Assay ID This is a unique identifier that can be used to identify the assay in the literature and online. -
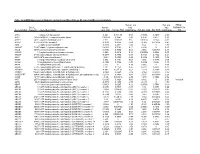
Mrna Expression in Human Leiomyoma and Eker Rats As Measured by Microarray Analysis
Table 3S: mRNA Expression in Human Leiomyoma and Eker Rats as Measured by Microarray Analysis Human_avg Rat_avg_ PENG_ Entrez. Human_ log2_ log2_ RAPAMYCIN Gene.Symbol Gene.ID Gene Description avg_tstat Human_FDR foldChange Rat_avg_tstat Rat_FDR foldChange _DN A1BG 1 alpha-1-B glycoprotein 4.982 9.52E-05 0.68 -0.8346 0.4639 -0.38 A1CF 29974 APOBEC1 complementation factor -0.08024 0.9541 -0.02 0.9141 0.421 0.10 A2BP1 54715 ataxin 2-binding protein 1 2.811 0.01093 0.65 0.07114 0.954 -0.01 A2LD1 87769 AIG2-like domain 1 -0.3033 0.8056 -0.09 -3.365 0.005704 -0.42 A2M 2 alpha-2-macroglobulin -0.8113 0.4691 -0.03 6.02 0 1.75 A4GALT 53947 alpha 1,4-galactosyltransferase 0.4383 0.7128 0.11 6.304 0 2.30 AACS 65985 acetoacetyl-CoA synthetase 0.3595 0.7664 0.03 3.534 0.00388 0.38 AADAC 13 arylacetamide deacetylase (esterase) 0.569 0.6216 0.16 0.005588 0.9968 0.00 AADAT 51166 aminoadipate aminotransferase -0.9577 0.3876 -0.11 0.8123 0.4752 0.24 AAK1 22848 AP2 associated kinase 1 -1.261 0.2505 -0.25 0.8232 0.4689 0.12 AAMP 14 angio-associated, migratory cell protein 0.873 0.4351 0.07 1.656 0.1476 0.06 AANAT 15 arylalkylamine N-acetyltransferase -0.3998 0.7394 -0.08 0.8486 0.456 0.18 AARS 16 alanyl-tRNA synthetase 5.517 0 0.34 8.616 0 0.69 AARS2 57505 alanyl-tRNA synthetase 2, mitochondrial (putative) 1.701 0.1158 0.35 0.5011 0.6622 0.07 AARSD1 80755 alanyl-tRNA synthetase domain containing 1 4.403 9.52E-05 0.52 1.279 0.2609 0.13 AASDH 132949 aminoadipate-semialdehyde dehydrogenase -0.8921 0.4247 -0.12 -2.564 0.02993 -0.32 AASDHPPT 60496 aminoadipate-semialdehyde -

Table S1. 103 Ferroptosis-Related Genes Retrieved from the Genecards
Table S1. 103 ferroptosis-related genes retrieved from the GeneCards. Gene Symbol Description Category GPX4 Glutathione Peroxidase 4 Protein Coding AIFM2 Apoptosis Inducing Factor Mitochondria Associated 2 Protein Coding TP53 Tumor Protein P53 Protein Coding ACSL4 Acyl-CoA Synthetase Long Chain Family Member 4 Protein Coding SLC7A11 Solute Carrier Family 7 Member 11 Protein Coding VDAC2 Voltage Dependent Anion Channel 2 Protein Coding VDAC3 Voltage Dependent Anion Channel 3 Protein Coding ATG5 Autophagy Related 5 Protein Coding ATG7 Autophagy Related 7 Protein Coding NCOA4 Nuclear Receptor Coactivator 4 Protein Coding HMOX1 Heme Oxygenase 1 Protein Coding SLC3A2 Solute Carrier Family 3 Member 2 Protein Coding ALOX15 Arachidonate 15-Lipoxygenase Protein Coding BECN1 Beclin 1 Protein Coding PRKAA1 Protein Kinase AMP-Activated Catalytic Subunit Alpha 1 Protein Coding SAT1 Spermidine/Spermine N1-Acetyltransferase 1 Protein Coding NF2 Neurofibromin 2 Protein Coding YAP1 Yes1 Associated Transcriptional Regulator Protein Coding FTH1 Ferritin Heavy Chain 1 Protein Coding TF Transferrin Protein Coding TFRC Transferrin Receptor Protein Coding FTL Ferritin Light Chain Protein Coding CYBB Cytochrome B-245 Beta Chain Protein Coding GSS Glutathione Synthetase Protein Coding CP Ceruloplasmin Protein Coding PRNP Prion Protein Protein Coding SLC11A2 Solute Carrier Family 11 Member 2 Protein Coding SLC40A1 Solute Carrier Family 40 Member 1 Protein Coding STEAP3 STEAP3 Metalloreductase Protein Coding ACSL1 Acyl-CoA Synthetase Long Chain Family Member 1 Protein