Genome-Wide Analysis of Long Non-Coding RNA Expression Profile in Lung Adenocarcinoma Compared to Spinal Metastasis
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Systematic Detection of Divergent Brain Proteins in Human Evolution and Their Roles in Cognition
bioRxiv preprint doi: https://doi.org/10.1101/658658; this version posted June 3, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY 4.0 International license. Systematic detection of divergent brain proteins in human evolution and their roles in cognition Guillaume Dumas1,*, Simon Malesys1 and Thomas Bourgeron1 Affiliations: 1 Human Genetics and Cognitive Functions, Institut Pasteur, UMR3571 CNRS, Université de Paris, Paris, (75015) France * Corresponding author: [email protected] 1 bioRxiv preprint doi: https://doi.org/10.1101/658658; this version posted June 3, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY 4.0 International license. Abstract The human brain differs from that of other primates, but the underlying genetic mechanisms remain unclear. Here we measured the evolutionary pressures acting on all human protein- coding genes (N=17,808) based on their divergence from early hominins such as Neanderthal, and non-human primates. We confirm that genes encoding brain-related proteins are among the most conserved of the human proteome. Conversely, several of the most divergent proteins in humans compared to other primates are associated with brain-associated diseases such as micro/macrocephaly, dyslexia, and autism. We identified specific eXpression profiles of a set of divergent genes in ciliated cells of the cerebellum, that might have contributed to the emergence of fine motor skills and social cognition in humans. -

Supplementary Note
1 SUPPLEMENTARY NOTE 2 Table of Contents 3 1 Studies contributing to systolic (SBP) and diastolic blood pressure (DBP) analyses ............ 4 4 2 Consortia and studies providing association results for cardiovascular outcomes ............. 4 5 2.1 CHARGE - Heart Failure Working Group .................................................................................. 4 6 2.2 EchoGen (LM mass and LV weight) .......................................................................................... 4 7 2.3 NEURO-CHARGE (stroke) ........................................................................................................ 4 8 2.4 MetaStroke (stroke) ................................................................................................................ 5 9 2.5 CARDIoGRAMplusC4D (CAD) ................................................................................................... 5 10 2.6 CHARGE CKDgen (CKD, eGFR, microalbuminuria, UACR) ......................................................... 5 11 2.7 KidneyGen (creatinine) ........................................................................................................... 6 12 2.8 CHARGE (cIMT) ....................................................................................................................... 6 13 2.9 CHARGE (mild retinopathy, central retinal artery caliber) ....................................................... 6 14 2.10 SEED (mild retinopathy, central retinal artery caliber) .......................................................... 6 15 -

Noncoding Rnas As Novel Pancreatic Cancer Targets
NONCODING RNAS AS NOVEL PANCREATIC CANCER TARGETS by Amy Makler A Thesis Submitted to the Faculty of The Charles E. Schmidt College of Science In Partial Fulfillment of the Requirements for the Degree of Master of Science Florida Atlantic University Boca Raton, FL August 2018 Copyright 2018 by Amy Makler ii ACKNOWLEDGEMENTS I would first like to thank Dr. Narayanan for his continuous support, constant encouragement, and his gentle, but sometimes critical, guidance throughout the past two years of my master’s education. His faith in my abilities and his belief in my future success ensured I continue down this path of research. Working in Dr. Narayanan’s lab has truly been an unforgettable experience as well as a critical step in my future endeavors. I would also like to extend my gratitude to my committee members, Dr. Binninger and Dr. Jia, for their support and suggestions regarding my thesis. Their recommendations added a fresh perspective that enriched our initial hypothesis. They have been indispensable as members of my committee, and I thank them for their contributions. My parents have been integral to my successes in life and their support throughout my education has been crucial. They taught me to push through difficulties and encouraged me to pursue my interests. Thank you, mom and dad! I would like to thank my boyfriend, Joshua Disatham, for his assistance in ensuring my writing maintained a logical progression and flow as well as his unwavering support. He was my rock when the stress grew unbearable and his encouraging words kept me pushing along. -

Comprehensive Analysis Reveals Novel Gene Signature in Head and Neck Squamous Cell Carcinoma: Predicting Is Associated with Poor Prognosis in Patients
5892 Original Article Comprehensive analysis reveals novel gene signature in head and neck squamous cell carcinoma: predicting is associated with poor prognosis in patients Yixin Sun1,2#, Quan Zhang1,2#, Lanlin Yao2#, Shuai Wang3, Zhiming Zhang1,2 1Department of Breast Surgery, The First Affiliated Hospital of Xiamen University, School of Medicine, Xiamen University, Xiamen, China; 2School of Medicine, Xiamen University, Xiamen, China; 3State Key Laboratory of Cellular Stress Biology, School of Life Sciences, Xiamen University, Xiamen, China Contributions: (I) Conception and design: Y Sun, Q Zhang; (II) Administrative support: Z Zhang; (III) Provision of study materials or patients: Y Sun, Q Zhang; (IV) Collection and assembly of data: Y Sun, L Yao; (V) Data analysis and interpretation: Y Sun, S Wang; (VI) Manuscript writing: All authors; (VII) Final approval of manuscript: All authors. #These authors contributed equally to this work. Correspondence to: Zhiming Zhang. Department of Surgery, The First Affiliated Hospital of Xiamen University, Xiamen, China. Email: [email protected]. Background: Head and neck squamous cell carcinoma (HNSC) remains an important public health problem, with classic risk factors being smoking and excessive alcohol consumption and usually has a poor prognosis. Therefore, it is important to explore the underlying mechanisms of tumorigenesis and screen the genes and pathways identified from such studies and their role in pathogenesis. The purpose of this study was to identify genes or signal pathways associated with the development of HNSC. Methods: In this study, we downloaded gene expression profiles of GSE53819 from the Gene Expression Omnibus (GEO) database, including 18 HNSC tissues and 18 normal tissues. -

Bioinformatics Mining of the Dark Matter Proteome For
BIOINFORMATICS MINING OF THE DARK MATTER PROTEOME FOR CANCER TARGETS DISCOVERY by Ana Paula Delgado A Thesis Submitted to the Faculty of The Charles E. Schmidt College of Science In Partial Fulfillment of the Requirements for the Degree of Master of Science Florida Atlantic University Boca Raton, Florida May 2015 Copyright 2015 by Ana Paula Delgado ii ACKNOWLEDGEMENTS I would first like to thank Dr. Narayanan for his continuous encouragement, guidance, and support during the past two years of my graduate education. It has truly been an unforgettable experience working in his laboratory. I also want to express gratitude to my external advisor Professor Van de Ven from the University of Leuven, Belgium for his constant involvement and assistance on my project. Moreover, I would like to thank Dr. Binninger and Dr. Dawson-Scully for their advice and for agreeing to serve on my thesis committee. I also thank provost Dr. Perry for his involvement in my project. I thank Jeanine Narayanan for editorial assistance with the publications and with this dissertation. It has been a pleasure working with various undergraduate students some of whom became lab mates including Pamela Brandao, Maria Julia Chapado and Sheilin Hamid. I thank them for their expert help in the projects we were involved in. Lastly, I want to express my profound thanks to my parents and brother for their unconditional love, support and guidance over the last couple of years. They were my rock when I was in doubt and never let me give up. I would also like to thank my boyfriend Spencer Daniel and best friends for being part of an incredible support system. -

Content Based Search in Gene Expression Databases and a Meta-Analysis of Host Responses to Infection
Content Based Search in Gene Expression Databases and a Meta-analysis of Host Responses to Infection A Thesis Submitted to the Faculty of Drexel University by Francis X. Bell in partial fulfillment of the requirements for the degree of Doctor of Philosophy November 2015 c Copyright 2015 Francis X. Bell. All Rights Reserved. ii Acknowledgments I would like to acknowledge and thank my advisor, Dr. Ahmet Sacan. Without his advice, support, and patience I would not have been able to accomplish all that I have. I would also like to thank my committee members and the Biomed Faculty that have guided me. I would like to give a special thanks for the members of the bioinformatics lab, in particular the members of the Sacan lab: Rehman Qureshi, Daisy Heng Yang, April Chunyu Zhao, and Yiqian Zhou. Thank you for creating a pleasant and friendly environment in the lab. I give the members of my family my sincerest gratitude for all that they have done for me. I cannot begin to repay my parents for their sacrifices. I am eternally grateful for everything they have done. The support of my sisters and their encouragement gave me the strength to persevere to the end. iii Table of Contents LIST OF TABLES.......................................................................... vii LIST OF FIGURES ........................................................................ xiv ABSTRACT ................................................................................ xvii 1. A BRIEF INTRODUCTION TO GENE EXPRESSION............................. 1 1.1 Central Dogma of Molecular Biology........................................... 1 1.1.1 Basic Transfers .......................................................... 1 1.1.2 Uncommon Transfers ................................................... 3 1.2 Gene Expression ................................................................. 4 1.2.1 Estimating Gene Expression ............................................ 4 1.2.2 DNA Microarrays ...................................................... -

University of Dundee a Comprehensive 1000 Genomes
University of Dundee A comprehensive 1000 Genomes-based genome-wide association meta-analysis of coronary artery disease Nikpay, Majid; Goel, Anuj; Won, Hong-Hee; Hall, Leanne M.; Willenborg, Christina; Kanoni, Stavroula Published in: Nature Genetics DOI: 10.1038/ng.3396 Publication date: 2015 Document Version Publisher's PDF, also known as Version of record Link to publication in Discovery Research Portal Citation for published version (APA): Nikpay, M., Goel, A., Won, H-H., Hall, L. M., Willenborg, C., Kanoni, S., Saleheen, D., Kyriakou, T., Nelson, C. P., Hopewell, J. C., Webb, T. R., Zeng, L., Dehghan, A., Elver, M., Armasu, S. M., Auro, K., Bjonnes, A., Chasman, D. I., Chen, S., ... Farrall, M. (2015). A comprehensive 1000 Genomes-based genome-wide association meta-analysis of coronary artery disease. Nature Genetics, 47(10), 1121-1130. https://doi.org/10.1038/ng.3396 General rights Copyright and moral rights for the publications made accessible in Discovery Research Portal are retained by the authors and/or other copyright owners and it is a condition of accessing publications that users recognise and abide by the legal requirements associated with these rights. • Users may download and print one copy of any publication from Discovery Research Portal for the purpose of private study or research. • You may not further distribute the material or use it for any profit-making activity or commercial gain. • You may freely distribute the URL identifying the publication in the public portal. Take down policy If you believe that this document breaches copyright please contact us providing details, and we will remove access to the work immediately and investigate your claim. -

C11orf16 (D-16): Sc-163799
SAN TA C RUZ BI OTEC HNOL OG Y, INC . C11orf16 (D-16): sc-163799 BACKGROUND PRODUCT C11orf16 (chromosome 11 open reading frame 16) is a 404 amino acid protein Each vial contains 200 µg IgG in 1.0 ml of PBS with < 0.1% sodium azide that is encoded by a gene located on human chromosome 11. With approxi - and 0.1% gelatin. mately 135 million base pairs and 1,400 genes, chromosome 11 makes up Blocking peptide available for competition studies, sc-163799 P, (100 µg around 4% of human genomic DNA and is considered a gene and disease pep tide in 0.5 ml PBS containing < 0.1% sodium azide and 0.2% BSA). association dense chromosome. The chromosome 11 encoded Atm gene is important for regulation of cell cycle arrest and apoptosis following double APPLICATIONS strand DNA breaks. Atm mutation leads to the disorder known as ataxia- C11orf16 (D-16) is recommended for detection of C11orf16 of human origin telangiectasia. The blood disorders Sickle cell anemia and β thalassemia are caused by HBB gene mutations. Wilms' tumors, WAGR syndrome and Denys- and BC051019 of mouse origin by Western Blotting (starting dilution 1:200, Drash syndrome are associated with mutations of the WT1 gene. Jervell and dilution range 1:100-1:1000), immunofluorescence (starting dilution 1:50, Lange-Nielsen syndrome, Jacobsen syndrome, Niemann-Pick disease, heredi - dilution range 1:50-1:500) and solid phase ELISA (starting dilution 1:30, tary angioedema and Smith-Lemli-Opitz syndrome are also associated with dilution range 1:30-1:3000); non cross-reactive with other C11orf family defects in chromosome 11. -

Supplementary Data
Gene expression profile of the A549 human non-small cell lung carcinoma cell line following treatment with the seeds of Descurainia sophia, a potential anticancer drug Bu-Yeo Kim, Jun Lee, Sung Joon Park, Ok-Sun Bang, and No Soo Kim Supplementary Data Figure Legends Supplementary Figure 1. Electropherogram of total RNA samples used for gene expression profiles. The A549 cells were treated with increasing concentrations of EEDS (0-20 mg/mL). After 24 h drug treatment, total RNAs were extracted from the cells, and their integrities were determined by electropherogram. RIN values range from 10 (intact) to 1(totally degraded). Supplementary Figure 2. Pathways enriched (FDR<0.01) in the Up- and Down-patterns. The genes that are shown in Table 5 and Figure 4, and involved in each pathway are highlighted in green Supplementary Figure 3. The level of similarity is represented in red with a scale bar. The color in the box of a diagonal line or “Activity” on the right panel represents the activity of the pathway. The positions and names of signaling-related pathways are colored red and the metabolism-related pathways are colored blue. Supplementary Table 1. Quality control of total RNA samples used for gene expression analysis. * EEDS (mg/mL) OD260/280 OD260/230 Ratio (28s/18s) RIN Result 0 2.04 2.22 2.3 9.8 Pass 1.25 2.03 2.28 2.2 10.0 Pass 5 2.03 2.28 2.1 10.0 Pass 20 2.04 2.23 2.0 10.0 Pass *RIN values, 10 (intact) to 1 (completely degraded). -
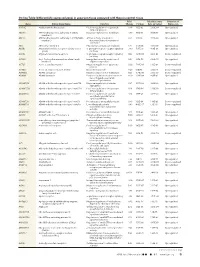
Differentially Expressed Genes in Aneurysm Tissue Compared With
On-line Table: Differentially expressed genes in aneurysm tissue compared with those in control tissue Fold False Discovery Direction of Gene Entrez Gene Name Function Change P Value Rate (q Value) Expression AADAC Arylacetamide deacetylase Positive regulation of triglyceride 4.46 1.33E-05 2.60E-04 Up-regulated catabolic process ABCA6 ATP-binding cassette, subfamily A (ABC1), Integral component of membrane 3.79 9.15E-14 8.88E-12 Up-regulated member 6 ABCC3 ATP-binding cassette, subfamily C (CFTR/MRP), ATPase activity, coupled to 6.63 1.21E-10 7.33E-09 Up-regulated member 3 transmembrane movement of substances ABI3 ABI family, member 3 Peptidyl-tyrosine phosphorylation 6.47 2.47E-05 4.56E-04 Up-regulated ACKR1 Atypical chemokine receptor 1 (Duffy blood G-protein–coupled receptor signaling 3.80 7.95E-10 4.18E-08 Up-regulated group) pathway ACKR2 Atypical chemokine receptor 2 G-protein–coupled receptor signaling 0.42 3.29E-04 4.41E-03 Down-regulated pathway ACSM1 Acyl-CoA synthetase medium-chain family Energy derivation by oxidation of 9.87 1.70E-08 6.52E-07 Up-regulated member 1 organic compounds ACTC1 Actin, ␣, cardiac muscle 1 Negative regulation of apoptotic 0.30 7.96E-06 1.65E-04 Down-regulated process ACTG2 Actin, ␥2, smooth muscle, enteric Blood microparticle 0.29 1.61E-16 2.36E-14 Down-regulated ADAM33 ADAM domain 33 Integral component of membrane 0.23 9.74E-09 3.95E-07 Down-regulated ADAM8 ADAM domain 8 Positive regulation of tumor necrosis 4.69 2.93E-04 4.01E-03 Up-regulated factor (ligand) superfamily member 11 production ADAMTS18 -

Deciphering Common and Rare Genetic E Ects on Reading Ability
Deciphering common and rare genetic eects on reading ability © 2016, Amaia Carrion´ Castillo ISBN: 978-90-76203-78-2 Printed and bound by Ipskamp Drukkers Deciphering common and rare genetic eects on reading ability Proefschri ter verkrijging van de graad van doctor aan de Radboud Universiteit Nijmegen op gezag van de rector magnicus prof. dr. J.H.J.M. van Krieken, volgens besluit van het college van decanen in het openbaar te verdedigen op maandag 7 november 2016 om 12.30 uur precies door Amaia Carrion´ Castillo geboren op 22 november 1988 te Donostia (Spanje) Promotor Prof. dr. Simon E. Fisher Copromotor Dr. Clyde Francks (MPI) Manuscriptcommissie Prof. dr. Anneke I. den Hollander Prof. dr. Juha Kere (Karolinska Institutet, Zweden) Prof. dr. Gerd Schulte-Korne¨ (Ludwig-Maximilians-University Munich, Duitsland) Deciphering common and rare genetic eects on reading ability Doctoral esis to obtain the degree of doctor from Radboud University Nijmegen on the authority of the Rector Magnicus prof. dr. J.H.J.M. van Krieken, according to the decision of the Council of Deans to be defended in public on Monday, November 7, 2016 at 12.30 hours by Amaia Carrion´ Castillo Born on 22 November 1988 in Donostia (Spain) Supervisor Prof. dr. Simon E. Fisher Co-supervisor Dr. Clyde Francks (MPI) Doctoral esis Committee Prof. dr. Anneke I. den Hollander Prof. dr. Juha Kere (Karolinska Institutet, Sweden) Prof. dr. Gerd Schulte-Korne¨ (Ludwig-Maximilians-University Munich, Germany) Etxekoei CONTENTS 1 General introduction 11 1.1 e learning-to-read brain ........................ 11 1.2 Variation in reading ability........................ 13 1.3 Complex and multifactorial aetiology................. -

Comprehensive Analysis of Gene Expression and DNA Methylation for Human Nasopharyngeal Carcinoma
European Archives of Oto-Rhino-Laryngology (2019) 276:2565–2576 https://doi.org/10.1007/s00405-019-05525-2 HEAD AND NECK Comprehensive analysis of gene expression and DNA methylation for human nasopharyngeal carcinoma Hu Li1 · Fu‑Ling Wang2 · Liang‑peng Shan3 · Jun An1 · Ming‑lei Liu1 · Wei Li1 · Jing‑E. Zhang1 · Ping‑ping Wu1 Received: 7 March 2019 / Accepted: 18 June 2019 / Published online: 25 June 2019 © Springer-Verlag GmbH Germany, part of Springer Nature 2019 Abstract Purpose Nasopharyngeal carcinoma (NPC) is one of the most malignant head and neck carcinomas with unique epidemio- logical features. In this study, we aimed to identify the novel NPC-related genes and biological pathways, shedding light on the potential molecular mechanisms of NPC. Methods Based on Gene Expression Omnibus (GEO) database, an integrated analysis of microarrays studies was performed to identify diferentially expressed genes (DEGs) and diferentially methylated genes (DMGs) in NPC compared to nor- mal control. The genes which were both diferentially expressed and diferentially methylated were identifed. Functional annotation and protein–protein interaction (PPI) network construction were used to uncover biological functions of DEGs. Results Two DNA methylation and fve gene expression datasets were incorporated. A total of 1074 genes were up-regulated and 939 genes were down-regulated in NPC were identifed. A total of 719 diferential methylation CpG sites (DMCs) includ- ing 1 hypermethylated sites and 718 hypomethylated sites were identifed. Among which, 11 genes were both DEGs and DMGs in NPC. Pathways in cancer, p53 signaling pathway and Epstein–Barr virus infection were three pathways signifcantly enriched pathways in DEmRNAs of NPC.