DNA CONCENTRATION MEASUREMENT at 260 Nm USING PHOTOPETTE® BIO
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

White Paper Absorbance Or Fluorescence: Which Is the Best
WHITE PAPER No. 40 Absorbance or Fluorescence: Which Is the Best Way to Quantify Nucleic Acids? Natascha Weiß1, Martin Armbrecht1 ¹Eppendorf AG, Hamburg, Germany Executive Summary When performing molecular experiments on nucleic acids, it is a basic requirement to determine the concentration as well as the quality of the sample. The standard method which serves this purpose is UV-Vis spectrophotometry – but is it always the right choice? In the present White Paper, the advantages and disadvantages of this technique will be described and it will be compared with nucleic acid quantification via fluorescence. In this context, it will be discussed which situations warrant the use of which one of the two methods. The decision will generally depend on the condition of the sample as well as on the requirements of downstream applications. Since the advantages of both methods complement each other well, it is most practical to have the ability to perform both methods on a single instrument in a flexible manner. Introduction Experiments involving nucleic acids are a mainstay in any evaluated by measuring the sample at additional wave molecular laboratory. DNA and RNA are isolated from micro- lengths (230 nm, 280 nm) and calculating the purity ratios, organisms and from cells of higher order organisms in order i.e. the ratios of the values obtained at 260/230 nm and at to be employed in a broad variety of processing steps and 260/280 nm, respectively. In this way, it will be evident analyses. It is crucial for any type of downstream applica- whether cellular debris or remainders of reagent used during tion that a defined amount of nucleic acid is used and that purification such as proteins, sugar molecules, certain salts the sample is free from contaminations that may impact the or phenols, are present in the solution, as these will generate experiments. -

On Food, Spectrophotometry, and Measurement Data Processing (Keynote Lecture)
12th IMEKO TC1 & TC7 Joint Symposium on Man Science & Measurement September, 3–5, 2008, Annecy, France ON FOOD, SPECTROPHOTOMETRY, AND MEASUREMENT DATA PROCESSING (KEYNOTE LECTURE) Roman Z. Morawski Warsaw University of Technology, Faculty of Electronics and Information Technology, Institute of Radioelectronics Warsaw, Poland [email protected] Abstract: Spectrophotometry is getting more and more Despite philosophical abnegation of food issues, the often the method of choice not only in laboratory analysis of practice of food preparation and refinement has flourished (bio)chemical substances, but also in the off-laboratory for centuries, and inspired research-and-invention-oriented identification and testing of physical properties of various minds. Today, we may speak about a fully developed products, in particular – of various organic mixtures discipline of science and technology. Enough to say that the including food products and ingredients. Specialized Institute of Food Technologists – the largest international, spectrophotometers, called spectrophotometric analyzers are non-profit professional organization involved in the designed for such applications. This keynote lecture is on advancement of food science and technology – is the state of the art and developmental trends in the domain encompassing 23 000 members worldwide. Its Committee of spectrophotometric analyzers of food with particular on Higher Education provided the following definition of emphasis on wine analyzers. The following issues are food science: "Food science is the discipline in which the covered: philosophical and methodological background of engineering, biological, and physical sciences are used to food analysis, physical and metrological principles of study the nature of foods, the causes of deterioration, the spectrophotometry, the role of measurement data processing principles underlying food processing, and the improvement in spectrophotometry, food analyzers on the market and of foods for the consuming public" [1]. -

ELISA Plate Reader
applications guide to microplate systems applications guide to microplate systems GETTING THE MOST FROM YOUR MOLECULAR DEVICES MICROPLATE SYSTEMS SALES OFFICES United States Molecular Devices Corp. Tel. 800-635-5577 Fax 408-747-3601 United Kingdom Molecular Devices Ltd. Tel. +44-118-944-8000 Fax +44-118-944-8001 Germany Molecular Devices GMBH Tel. +49-89-9620-2340 Fax +49-89-9620-2345 Japan Nihon Molecular Devices Tel. +06-6399-8211 Fax +06-6399-8212 www.moleculardevices.com ©2002 Molecular Devices Corporation. Printed in U.S.A. #0120-1293A SpectraMax, SoftMax Pro, Vmax and Emax are registered trademarks and VersaMax, Lmax, CatchPoint and Stoplight Red are trademarks of Molecular Devices Corporation. All other trademarks are proprty of their respective companies. complete solutions for signal transduction assays AN EXAMPLE USING THE CATCHPOINT CYCLIC-AMP FLUORESCENT ASSAY KIT AND THE GEMINI XS MICROPLATE READER The Molecular Devices family of products typical applications for Molecular Devices microplate readers offers complete solutions for your signal transduction assays. Our integrated systems γ α β s include readers, washers, software and reagents. GDP αs AC absorbance fluorescence luminescence GTP PRINCIPLE OF CATCHPOINT CYCLIC-AMP ASSAY readers readers readers > Cell lysate is incubated with anti-cAMP assay type SpectraMax® SpectraMax® SpectraMax® VersaMax™ VMax® EMax® Gemini XS LMax™ ATP Plus384 190 340PC384 antibody and cAMP-HRP conjugate ELISA/IMMUNOASSAYS > nucleus Single addition step PROTEIN QUANTITATION cAMP > λEX 530 nm/λEM 590 nm, λCO 570 nm UV (280) Bradford, BCA, Lowry For more information on CatchPoint™ assay NanoOrange™, CBQCA kits, including the complete procedure for this NUCLEIC ACID QUANTITATION assay (MaxLine Application Note #46), visit UV (260) our web site at www.moleculardevices.com. -
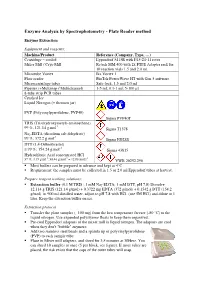
Enzyme Analysis by Spectrophotometry - Plate Reader Method
Enzyme Analysis by Spectrophotometry - Plate Reader method Enzyme Extraction Equipment and reagents: Machine/Product Reference (Company, Type, …) Centrifuge – cooled Eppendorf 5415R with F45-24-11 rotor Mixer Mill / Cryo Mill Retsch MM 400 with 2x PTFE Adapter rack for 10 reaction vials 1.5 and 2.0 ml. Microtube Vortex Ika Vortex 1 Plate reader BioTek PowerWave HT with Gen 5 software Microcentrifuge tubes Safe-lock, 1.5 and 2.0 ml Pipettes (+Multistep / Multichannel) 1-5 ml; 0.1-1 ml; 5-100 µl 8-tube strip PCR tubes Crushed Ice Liquid Nitrogen (+ thermos jar) PVP (Polyvinylpyrrolidone, PVP40) Sigma PVP40T TRIS (Tris(hydroxymetyl)-aminoethane) -1 99 %, 121.14 g.mol Sigma T1378 Na 2-EDTA (disodium salt dehydrate) -1 99 %; 372.2 g.mol Sigma ED2SS DTT (1,4-Dithiothreitol) -1 ≥ 99 %; 154.24 g.mol Sigma 43815 Hydrochloric Acid concentrated HCl -1 -1 -1 37 %; 1,19 g.ml ; 36.46 g.mol = 12.08 mol.l VWR 20252.290 Most buffers can be prepared in advance and kept at 4°C Requirement: the samples must be collected in 1.5 or 2.0 ml Eppendorf tubes at harvest. Prepare reagent working solutions: Extraction buffer (0.1 M TRIS ; 1 mM Na 2-EDTA; 1 mM DTT, pH 7.8) Dissolve 12.114 g TRIS (121.14 g/mol) + 0.3722 mg EDTA (372 g/mol) + 0.1542 g DTT (154.2 g/mol) in 900 ml distilled water, adjust to pH 7.8 with HCl (use 5M HCl) and dilute to 1 liter. Keep the extraction buffer on ice. -

Microlab 500-Series
MicroLab 500-series Getting Started 2 Contents CHAPTER 1: Getting Started Connecting the Hardware…………………………………………………………………………….…...6 Installing the USB driver…………………….…………………………………………………...………..6 Installing the Software…………………………………………………………………………..………...8 Starting a new Experiment………………………………………………………………………..……….8 CHAPTER 2: Setting up a MicroLab Experiment Adding Input Sources Sensor Variables……………………………………………………………………………….…………10 Adding a Sensor ………………………………………………………………………………………….10 Calibrating a Sensor………………………………………………………………………………………11 The Calibration Module………………………………………………………………………………......11 Adding a Calibration Point……………………………………………………………………………….11 Correlating the Data……………………………………………………………………………………....12 Adding More Variables………………………………………………………………………………...…14 Formula Variables………………………………………………………………………………………...15 Designing an Experiment The Programming Steps………………………………………………………………………..………..17 Displaying the Data…………………………………………………………………………..…….……..17 Controlling an Experiment Starting the Experiment…………………………………………………………………….………..…..18 Stopping the Experiment…………………………………………………………………….………......18 Repeating an Experiment………………………………………………………………………………..18 Switches A & B……………………………………………………………………………………………19 Analyzing the Data Graph Analysis…………………………………………………………………………..………..………19 Add a Curve Fit………………………………………………………………………….....……19 Data Analysis…………………….…………………………………………………………………..……22 CHAPTER 3 - Spectrophotometry (522/516) Calibrating the Spectrophotometer Reading a Blank………...…………………………………………………………………………..…….23 3 Reading -

Validation of Spectrophotometric Microplate
a Food Science and Technology ISSN 0101-2061 DDII: httpI://dx.doi.org/10.1590/1678-457X.36216 Validation of spectrophotometric microplate methods for polyphenol oxidase and peroxidase activities analysis in fruits and vegetables Érica Sayuri SIGUEMDTD1, Jorge Andrey Wilhelms GUT1,2* Abstract Enzymes polyphenol oxidase (PPD) and peroxidase (PDD) play important roles in the processing of fruits and vegetables, since they can produce undesirable changes in color, texture and flavor. Classical methods of activity assessment are based on cuvette spectrophotometric readings. This work aims to propose, to validate and to test microplate spectrophotometric methods. Samples of apple juice and lyophilized enzymes from mushroom and horseradish were analyzed by the cuvette and microplate methods and it was possible to validate the microplate assays with satisfactory results regarding linearity, repeatability, accuracy along with quantitation and detection limits. The proposed microplate methods proved to be reliable and reproducible as the classical methods besides having the advantages of allowing simultaneous analysis and requiring a reduced amount of samples and reactants, which can beneficial to the study of enzyme inactivation in the processing of fruits and vegetables. Keywords: microplate; polyphenol oxidase; peroxidase; enzymatic assay; apple juice. Practical Application: Rapid assessment of PDD/PPD activities in fruits and vegetables with reduced amounts of reactants. 1 Introduction Monitoring the activity of enzymes polyphenol oxidase 2002). The selection of the appropriate substrate is also important, (PPD, E.C.1.14.18.1) and peroxidase (PDD, E.C.1.11.1.7) is an seeing that the colored compounds formed from the oxidation important control point for harvesting, storing and processing products have their maximum absorption at different wavelengths of fruits and vegetables. -

Chemistry 2A Lab Manual Standard Operating Procedures Winter Quarter 2018
Chemistry 2A Lab Manual Standard Operating Procedures Winter Quarter 2018 Department of Chemistry University of California - Davis Davis, CA 95616 Student Name Locker # Laboratory Information Teaching Assistant’s Name Laboratory Section Number Laboratory Room Number Dispensary Room Number 1060 Sciences Lab Building Location of Safety Equipment Nearest to Your Laboratory Safety Shower Eye Wash Fountain Fire Extinguisher Fire Alarm Safety Chemicals Revision Date 12/1/2017 Preface Chemistry is an experimental science. Thus, it is important that students of chemistry do experiments in the laboratory to more fully understand that the theories they study in lecture and in their textbook are developed from the critical evaluation of experimental data. The laboratory can also aid the student in the study of the science by clearly illustrating the principles and concepts involved. Finally, laboratory experimentation allows students the opportunity to develop techniques and other manipulative skills that students of science must master. The faculty of the Chemistry Department at UC Davis clearly understands the importance of laboratory work in the study of chemistry. The Department is committed to this component of your education and hopes that you will take full advantage of this opportunity to explore the science of chemistry. A unique aspect of this laboratory program is that a concerted effort has been made to use environmentally less toxic or non-toxic materials in these experiments. This was not only done to protect students but also to lessen the impact of this program upon the environment. This commitment to the environment has presented an enormous challenge, as many traditional experiments could not be used due to the negative impact of the chemicals involved. -

Chemistry 50 and 51 Laboratory Manual General Information
Chemistry 50 and 51 Laboratory Manual General Information Mt. San Antonio College Chemistry Department 2019 - 2020 TABLE OF CONTENTS PREFACE………………………………………………………………………..………..… 1 GENERAL INFORMATION Safety………………..………………………..…………………………………INFORMATION….……… 3 Equipment……………………………………..……………………………….………….. 9 Techniques………………………………………………………………………………… 13 Heating……………………………………...……………………………….…..….… 13 Cleaning and Labeling Glassware……….……………………….........……........…... 14 Reading Analog Scales…………………………………………………..……….…... 14 Volumetric Flasks………………………………………………………..……….…... 15 Graduated Cylinders.………………………………………………………………..… 15 Volumetric Pipets……………………………………..…………………………..…... 16 Graduated Pipets……………….……………………..………………………….…… 16 Burets………………………….………………..………………..………..….………. 18 Analytical Balances…………………………………………………….……….…….. 19 Solution Preparation…………………………………………..……………….….…... 20 Percent Concentration....……………………………………………….……….…….. 20 Molarity……………………………………………………………………………….. 21 Dilution……………………………………………………………...………………… 22 Titration ………………………………………………………………....…………..... 23 Vacuum Filtration……………………….…………………………………..…….…… 24 Spectrophotometry and Beer’s Law…………………………………..…………..…... 25 Measurement of pH………………………………………………….….….……..…... 27 Pasco Spectrometer……………………………………..…………………...….…… 28 Vernier Go Direct Sensors ………………………..…………………...….………….. 31 Notebook…………………………………………………………….….………..……...... 35 Precision and Accuracy……………………………………………………….………....... 37 Spreadsheet and Graphing with Excel…..…………………..…………..……………....... 46 EXPERIMENTS PREFACE The laboratory -

UV-Vis Spectrophotometry: Monochromators Vs Photodiode Arrays
Technical Note UV-Vis spectrophotometry: monochromators vs photodiode arrays Comparing absorbance measurements between the Quad4 Monochromators™-based Infinite® M200 PRO and a multimode reader using photodiode array technology Introduction Materials and methods The most widely established technology for UV-Vis Infinite M200 PRO multimode microplate reader absorbance measurement is a monochromator-based Multimode microplate reader with a PDA microplate reader. Spectrophotometers have undergone a Herring sperm DNA standard great deal of development since their introduction in the Tris-EDTA early 1950s (1) and, in recent years, multimode readers using Orange G (OG) linear photodiode array (PDA) technology for absorbance ddH2O measurements have become available. PDA-based readers 96-well, transparent UV-Star® plates incorporate an optical grating and a solid state array detector, enabling measurement of light intensity throughout the UV and Experiment 1 visible regions of the spectrum. Similar to a monochromator, Linearity in the visible spectrum using Orange G but much faster, they allow the entire UV-Vis spectrum of a In the first experiment, the OD (optical density) linearity of the sample to be captured within a few seconds per well. PDA spectrophotometer in the visible wavelength range was However, this technology suffers from a number of drawbacks, compared with the OD linearity of the monochromator-based mainly due to high levels of stray light. This results in a Infinite M200 PRO. An OG dilution series was prepared in dramatically limited dynamic measurement range (2). ddH20 (200, 150, 112.5, 84.4, 63.3, 47.5 and 35.6 mg/ml) and This technical note compares the results of basic absorbance 200 µl of each concentration was pipetted into a 96-well measurements performed on an Infinite M200 PRO multimode UV-Star plate in triplicate. -

Multi-Volume Based Protein Quantification Methods
Application Guide Multi-Volume Based Protein Quantification Methods Why Quantify Proteins? Proteins are central to our understanding of biology. In cells, they are multipurpose: from actin providing structural support to proteins acting as enzymes for modulating signal transduction pathways, such as kinases, proteases and phosphatases; to transmembrane proteins that allow for extracellular interactions, such as GPCRs and ion channels. Although almost all proteins are made from the same set of 20 amino acids, their structures and functions are incredibly diverse through the various interactions that can occur through the amino acids that comprise them, their ability to assume quaternary structure and post-translational modifications that can modulate their activity. Many of the studies that probe protein structure and function rely on upstream quantification to ensure suitable results. These downstream studies include mass spectrometry to elucidate primary structure and post-translational modifications, circular dichroism and x-ray crystallography to probe secondary, tertiary and quaternary structure, 2-D gel electrophoresis and western blots to monitor relative protein expression and protein arrays for highly multiplexed protein expression studies. Fluorescence and absorbance spectroscopy are both common protein quantification methods, although the former is preferred when dilute protein samples are used (i.e. typically <100 µg/mL). In general, absorbance quantification methods provide sufficient analytical performance and are convenient for many studies and laboratories due to the widespread availability of absorbance spectrophotometers. Convenience via increased sample throughput is realized when these methodologies are performed in microplates; and via simplified work flows using small amounts of sample in micro-volume formats. In this application guide, absorbance-based protein quantification methods appropriate for microplate and micro-volume analysis are discussed in detail. -

Lab 2 Spectrophotometric Measurement of Glucose
Lab 2 Spectrophotometric Measurement of Glucose Objectives 1. Learn how to use a spectrophotometer. 2. Produce a glucose standard curve. 3. Perform a glucose assay. Safety Precautions Glucose Color Reagent and the Glucose Standard are irritants. Hydrochloric acid is a corrosive. Use gloves and goggles. Materials Spectrophotometer (340-600 nm) 0.1, 1.0, and 10 mL serological pipettes 15 x 125 mixing tubes cuvettes 0.1 N Hydrochloric acid Glucose Kit (Sigma 115-A) 500 mg/dl Glucose standard (Sigma G3761) Grape Kool-Aid test solution Blank 1 Blank 2 Grape Kool-Aid Glucose Introduction Diabetes mellitus is a serious incurable disease that affects approximately 16 million U.S. citizens, although only 10 million have been diagnosed. It is a major cause of death in the United States, and its complications include kidney failure, blindness, and lower limb amputations. Diabetes mellitus is caused by the inability of body cells to uptake glucose. This inability can be caused either by low levels of insulin, a hormone necessary for the movement of glucose across cell membranes, or by a defect in the insulin-binding receptors on cell membranes. Glucose is a monosaccharide that is used by cells as an energy source, although when it is not available, most cells in the body use fatty acids as their fuel. After a meal, glucose is absorbed into the bloodstream, elevating the concentration of glucose found in the blood. Insulin is released as a result, and all cells switch to burning glucose. Diabetes is diagnosed by measuring the amount of glucose in blood after an 8-hour fast from food. -

Glass Filters As a Standard Reference Material for Spectrophotometry-- Selection, Preparation, Certification, Use Srm 930
NBS SPECIAL PUBLICATION 260-51 y*£AU of U.S. DEPARTMENT OF COMMERCE / National Bureau of Standards GLASS FILTERS AS A STANDARD REFERENCE MATERIAL FOR SPECTROPHOTOMETRY-- SELECTION, PREPARATION, CERTIFICATION, USE SRM 930 C.3L NATIONAL BUREAU OF STANDARDS 1 The National Bureau of Standards was established by an act of Congress March 3, 1901. The Bureau's overall goal is to strengthen and advance the Nation's science and technology and facilitate their effective application for public benefit. To this end, the Bureau conducts research and provides: (1) a basis for the Nation's physical measurement system, (2) scientific and technological services for industry and government, (3) a technical basis for equity in trade, and (4) technical services to promote public safety. The Bureau consists of the Institute for Basic Standards, the Institute for Materials Research, the Institute for Applied Technology, the Institute for Computer Sciences and Technology, and the Office for Information Programs. THE INSTITUTE FOR BASIC STANDARDS provides the central basis within the United States of a complete and consistent system of physical measurement; coordinates that system with measurement systems of other nations; and furnishes essential services leading to accurate and uniform physical measurements throughout the Nation's scientific community, industry, and commerce. The Institute consists of the Office of Measurement Services, the Office of Radiation Measurement and the following Center and divisions: Applied Mathematics — Electricity — Mechanics