FOXF2 Differentially Regulates Expression of Metabolic Genes In
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Screening and Identification of Key Biomarkers in Clear Cell Renal Cell Carcinoma Based on Bioinformatics Analysis
bioRxiv preprint doi: https://doi.org/10.1101/2020.12.21.423889; this version posted December 23, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Screening and identification of key biomarkers in clear cell renal cell carcinoma based on bioinformatics analysis Basavaraj Vastrad1, Chanabasayya Vastrad*2 , Iranna Kotturshetti 1. Department of Biochemistry, Basaveshwar College of Pharmacy, Gadag, Karnataka 582103, India. 2. Biostatistics and Bioinformatics, Chanabasava Nilaya, Bharthinagar, Dharwad 580001, Karanataka, India. 3. Department of Ayurveda, Rajiv Gandhi Education Society`s Ayurvedic Medical College, Ron, Karnataka 562209, India. * Chanabasayya Vastrad [email protected] Ph: +919480073398 Chanabasava Nilaya, Bharthinagar, Dharwad 580001 , Karanataka, India bioRxiv preprint doi: https://doi.org/10.1101/2020.12.21.423889; this version posted December 23, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Abstract Clear cell renal cell carcinoma (ccRCC) is one of the most common types of malignancy of the urinary system. The pathogenesis and effective diagnosis of ccRCC have become popular topics for research in the previous decade. In the current study, an integrated bioinformatics analysis was performed to identify core genes associated in ccRCC. An expression dataset (GSE105261) was downloaded from the Gene Expression Omnibus database, and included 26 ccRCC and 9 normal kideny samples. Assessment of the microarray dataset led to the recognition of differentially expressed genes (DEGs), which was subsequently used for pathway and gene ontology (GO) enrichment analysis. -

Mediator of DNA Damage Checkpoint 1 (MDC1) Is a Novel Estrogen Receptor Co-Regulator in Invasive 6 Lobular Carcinoma of the Breast 7 8 Evelyn K
bioRxiv preprint doi: https://doi.org/10.1101/2020.12.16.423142; this version posted December 16, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC 4.0 International license. 1 Running Title: MDC1 co-regulates ER in ILC 2 3 Research article 4 5 Mediator of DNA damage checkpoint 1 (MDC1) is a novel estrogen receptor co-regulator in invasive 6 lobular carcinoma of the breast 7 8 Evelyn K. Bordeaux1+, Joseph L. Sottnik1+, Sanjana Mehrotra1, Sarah E. Ferrara2, Andrew E. Goodspeed2,3, James 9 C. Costello2,3, Matthew J. Sikora1 10 11 +EKB and JLS contributed equally to this project. 12 13 Affiliations 14 1Dept. of Pathology, University of Colorado Anschutz Medical Campus 15 2Biostatistics and Bioinformatics Shared Resource, University of Colorado Comprehensive Cancer Center 16 3Dept. of Pharmacology, University of Colorado Anschutz Medical Campus 17 18 Corresponding author 19 Matthew J. Sikora, PhD.; Mail Stop 8104, Research Complex 1 South, Room 5117, 12801 E. 17th Ave.; Aurora, 20 CO 80045. Tel: (303)724-4301; Fax: (303)724-3712; email: [email protected]. Twitter: 21 @mjsikora 22 23 Authors' contributions 24 MJS conceived of the project. MJS, EKB, and JLS designed and performed experiments. JLS developed models 25 for the project. EKB, JLS, SM, and AEG contributed to data analysis and interpretation. SEF, AEG, and JCC 26 developed and performed informatics analyses. MJS wrote the draft manuscript; all authors read and revised the 27 manuscript and have read and approved of this version of the manuscript. -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

Dissertation Submitted to the Combined Faculties for the Natural Sciences and for Mathematics of the Ruperto-Carola University O
Dissertation submitted to the Combined Faculties for the Natural Sciences and for Mathematics of the Ruperto-Carola University of Heidelberg, Germany for the degree of Doctor of Natural Sciences presented by Master of Information Technology Anna-Lena Kranz born in: Bielefeld Oral-examination: 28.03.2011 Analyzing combinatorial regulation of transcription in mammalian cells Referees: PD Dr. Rainer K¨onig PD Dr. Stefan Wiemann Abstract Analyzing combinatorial regulation of transcription in mammalian cells During development and differentiation of an organism, accurate gene regulation is central for cells to maintain and balance their differentiation processes. Tran- scriptional interactions between cis-acting DNA-elements such as promoters and enhancers are the basis for precise and balanced transcriptional regulation. In this thesis, proximal and distal regulatory regions upstream of all transcription start sites were considered in silico to identify regulatory modules consisting of combina- tions of transcription factors (TFs) with binding sites at promoters and enhancers. Applying these modules to a broad variety of gene expression profiles demonstrated that the identified modules regulate gene expression during mouse embryonic de- velopment and human stem cell differentiation in a tissue- and temporal-specific manner. Whereas tissue-specific regulation is mainly controlled by combinations of TFs binding at promoters, the combination of TFs binding at promoters together with TFs binding at the respective enhancers determines the regulation of tem- poral progression during development. The identified regulatory modules showed considerably good predictive power to discriminate genes being differentially reg- ulated at a specific time interval. In addition, TF binding sites are immanently different for promoter and enhancer regions. -

Gene Regulation Is Governed by a Core Network in Hepatocellular Carcinoma Zuguang Gu, Chenyu Zhang* and Jin Wang*
Gu et al. BMC Systems Biology 2012, 6:32 http://www.biomedcentral.com/1752-0509/6/32 RESEARCH ARTICLE Open Access Gene regulation is governed by a core network in hepatocellular carcinoma Zuguang Gu, Chenyu Zhang* and Jin Wang* Abstract Background: Hepatocellular carcinoma (HCC) is one of the most lethal cancers worldwide, and the mechanisms that lead to the disease are still relatively unclear. However, with the development of high-throughput technologies it is possible to gain a systematic view of biological systems to enhance the understanding of the roles of genes associated with HCC. Thus, analysis of the mechanism of molecule interactions in the context of gene regulatory networks can reveal specific sub-networks that lead to the development of HCC. Results: In this study, we aimed to identify the most important gene regulations that are dysfunctional in HCC generation. Our method for constructing gene regulatory network is based on predicted target interactions, experimentally-supported interactions, and co-expression model. Regulators in the network included both transcription factors and microRNAs to provide a complete view of gene regulation. Analysis of gene regulatory network revealed that gene regulation in HCC is highly modular, in which different sets of regulators take charge of specific biological processes. We found that microRNAs mainly control biological functions related to mitochondria and oxidative reduction, while transcription factors control immune responses, extracellular activity and the cell cycle. On the higher level of gene regulation, there exists a core network that organizes regulations between different modules and maintains the robustness of the whole network. There is direct experimental evidence for most of the regulators in the core gene regulatory network relating to HCC. -
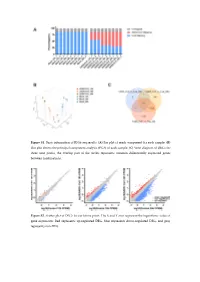
Figure S1. Basic Information of RNA-Seq Results. (A) Bar Plot of Reads Component for Each Sample
Figure S1. Basic information of RNA-seq results. (A) Bar plot of reads component for each sample. (B) Dot plot shows the principal component analysis (PCA) of each sample. (C) Venn diagram of DEGs for three time points, the overlap part of the circles represents common differentially expressed genes between combinations. Figure S2. Scatter plot of DEGs for each time point. The X and Y axes represent the logarithmic value of gene expression. Red represents up-regulated DEG, blue represents down-regulated DEG, and gray represents non-DEG. Table S1. Primers used for quantitative real-time PCR analysis of DEGs. Gene Primer Sequence Forward 5’-CTACGAGTGGATGGTCAAGAGC-3’ FOXO1 Reverse 5’-CCAGTTCCTTCATTCTGCACACG-3’ Forward 5’-GACGTCCGGCATCAGAGAAA-3’ IRS2 Reverse 5’-TCCACGGCTAATCGTCACAG-3’ Forward 5’-CACAACCAGGACCTCACACC-3’ IRS1 Reverse 5’-CTTGGCACGATAGAGAGCGT-3’ Forward 5’-AGGATACCACTCCCAACAGACCT-3’ IL6 Reverse 5’-CAAGTGCATCATCGTTGTTCATAC-3’ Forward 5’-TCACGTTGTACGCAGCTACC-3’ CCL5 Reverse 5’-CAGTCCTCTTACAGCCTTTGG-3’ Forward 5’-CTGTGCAGCCGCAGTGCCTACC-3’ BMP7 Reverse 5’-ATCCCTCCCCACCCCACCATCT-3’ Forward 5’-CTCTCCCCCTCGACTTCTGA-3’ BCL2 Reverse 5’-AGTCACGCGGAACACTTGAT-3’ Forward 5’-CTGTCGAACACAGTGGTACCTG-3’ FGF7 Reverse 5’-CCAACTGCCACTGTCCTGATTTC-3’ Forward 5’-GGGAGCCAAAAGGGTCATCA-3’ GAPDH Reverse 5’-CGTGGACTGTGGTCATGAGT-3’ Supplementary material: Differentially expressed genes log2(SADS-CoV_12h/ Qvalue (SADS-CoV _12h/ Gene Symbol Control_12h) Control_12h) PTGER4 -1.03693 6.79E-04 TMEM72 -3.08132 3.66E-04 IFIT2 -1.02918 2.11E-07 FRAT2 -1.09282 4.66E-05 -

Discovery of Biased Orientation of Human DNA Motif Sequences
bioRxiv preprint doi: https://doi.org/10.1101/290825; this version posted January 27, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY 4.0 International license. 1 Discovery of biased orientation of human DNA motif sequences 2 affecting enhancer-promoter interactions and transcription of genes 3 4 Naoki Osato1* 5 6 1Department of Bioinformatic Engineering, Graduate School of Information Science 7 and Technology, Osaka University, Osaka 565-0871, Japan 8 *Corresponding author 9 E-mail address: [email protected], [email protected] 10 1 bioRxiv preprint doi: https://doi.org/10.1101/290825; this version posted January 27, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY 4.0 International license. 11 Abstract 12 Chromatin interactions have important roles for enhancer-promoter interactions 13 (EPI) and regulating the transcription of genes. CTCF and cohesin proteins are located 14 at the anchors of chromatin interactions, forming their loop structures. CTCF has 15 insulator function limiting the activity of enhancers into the loops. DNA binding 16 sequences of CTCF indicate their orientation bias at chromatin interaction anchors – 17 forward-reverse (FR) orientation is frequently observed. DNA binding sequences of 18 CTCF were found in open chromatin regions at about 40% - 80% of chromatin 19 interaction anchors in Hi-C and in situ Hi-C experimental data. -

Genome-Wide Profiling of Druggable Active Tumor Defense Mechanisms to Enhance Cancer Immunotherapy
bioRxiv preprint doi: https://doi.org/10.1101/843185; this version posted November 15, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Genome-wide profiling of druggable active tumor defense mechanisms to enhance cancer immunotherapy Rigel J. Kishton1,2,*,#, Shashank J. Patel1,2,†,*, Suman K. Vodnala1,2, Amy E. Decker3, Yogin Patel1,2, Madhusudhanan Sukumar1,2, Tori N. Yamamoto1,2,4, Zhiya Yu1,2, Michelle Ji1,2, Amanda N. Henning1,2, Devikala Gurusamy1,2, Douglas C. Palmer1,2, Winifred Lo1, Anna Pasetto1, Parisa Malekzadeh1, Drew C. Deniger1, Kris C. Wood3, Neville E. Sanjana5,6, Nicholas P. Restifo1,2, #, § 1Surgery Branch, Center for Cancer Research, National Cancer Institute, Bethesda, MD 20892, USA 2Center for Cell-Based Therapy, National Cancer Institute, Bethesda, MD 20892, USA 3Department of Pharmacology & Cancer Biology, Duke University School of Medicine, Durham, NC, USA 4Immunology Graduate Group, University of Pennsylvania, Philadelphia, PA 19104, USA 5New York Genome Center, New York, NY 10013 USA 6Department of Biology, New York University, New York, NY 10003, USA *These authors contributed equally to this work. †Present address: NextCure Inc., Beltsville, MD 20705, USA §Present address: Lyell Immunopharma, South San Francisco, CA 94080, USA #Corresponding authors. NPR: [email protected]. RJK: [email protected]. bioRxiv preprint doi: https://doi.org/10.1101/843185; this version posted November 15, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. -

In Silico Learning of Tumor Evolution Through Mutational Time Series
In silico learning of tumor evolution through mutational time series Noam Auslandera, Yuri I. Wolfa, and Eugene V. Koonina,1 aNational Center for Biotechnology Information, National Library of Medicine, National Institutes of Health, Bethesda, MD 20894 Contributed by Eugene V. Koonin, March 15, 2019 (sent for review January 29, 2019; reviewed by Natalia L. Komarova and Andrei Zinovyev) Cancer arises through the accumulation of somatic mutations over potentially revealing the complex dynamics of the tumor evolu- time. Understanding the sequence of mutation occurrence during tion and enabling a variety of context-specific predictions. cancer progression can assist early and accurate diagnosis and To this end, we employed long short-term memory (LSTM) improve clinical decision-making. Here we employ long short-term networks (20), a type of recurrent neural network (RNN) (21) memory (LSTM) networks, a class of recurrent neural network, to capable of learning long-term dependencies in a time series se- learn the evolution of a tumor through an ordered sequence of quence. These networks have achieved major success in time mutations. We demonstrate the capacity of LSTMs to learn series prediction tasks and for learning evolution of recurrent complex dynamics of the mutational time series governing tumor systems (20, 22–24). We demonstrate here the utility of applying progression, allowing accurate prediction of the mutational LSTM to time-ordered mutational data in colon and lung ade- burden and the occurrence of mutations in the sequence. Using nocarcinomas. We define a pseudotemporal gene ranking, rep- the probabilities learned by the LSTM, we simulate mutational resenting each tumor sample as a binary text, which can be used data and show that the simulation results are statistically in- for training LSTM in a similar manner to that applied in natural distinguishable from the empirical data. -

Human Transcription Factors Responsive to Initial Reprogramming Predominantly Undergo Legitimate Reprogramming During Fbroblast Conversion to Ipscs Ricardo R
www.nature.com/scientificreports OPEN Human transcription factors responsive to initial reprogramming predominantly undergo legitimate reprogramming during fbroblast conversion to iPSCs Ricardo R. Cevallos1, Yvonne J. K. Edwards1,2, John M. Parant3, Bradley K. Yoder2 & Kejin Hu1* The four transcription factors OCT4, SOX2, KLF4, and MYC (OSKM) together can convert human fbroblasts to induced pluripotent stem cells (iPSCs). It is, however, perplexing that they can do so only for a rare population of the starting cells with a long latency. Transcription factors (TFs) defne identities of both the starting fbroblasts and the end product, iPSCs, and are also of paramount importance for the reprogramming process. It is critical to upregulate or activate the iPSC-enriched TFs while downregulate or silence the fbroblast-enriched TFs. This report explores the initial TF responses to OSKM as the molecular underpinnings for both the potency aspects and the limitation sides of the OSKM reprogramming. The authors frst defned the TF reprogramome, i.e., the full complement of TFs to be reprogrammed. Most TFs were resistant to OSKM reprogramming at the initial stages, an observation consistent with the inefciency and long latency of iPSC reprogramming. Surprisingly, the current analyses also revealed that most of the TFs (at least 83 genes) that did respond to OSKM induction underwent legitimate reprogramming. The initial legitimate transcriptional responses of TFs to OSKM reprogramming were also observed in the reprogramming fbroblasts from a diferent individual. Such early biased legitimate reprogramming of the responsive TFs aligns well with the robustness aspect of the otherwise inefcient and stochastic OSKM reprogramming. OCT4, SOX2, KLF4, and MYC (collectively OSKM) can convert human fbroblasts into induced pluripotent stem cells (iPSCs), which are the man-made version of embryonic stem cells (ESCs)1–3. -

Frequent Loss-Of-Heterozygosity in CRISPR-Cas9–Edited Early Human Embryos
Frequent loss-of-heterozygosity in CRISPR-Cas9–edited COLLOQUIUM PAPER early human embryos Gregorio Alanis-Lobatoa, Jasmin Zohrenb, Afshan McCarthya, Norah M. E. Fogartya,c, Nada Kubikovad,e, Emily Hardmana, Maria Grecof, Dagan Wellsd,g, James M. A. Turnerb, and Kathy K. Niakana,h,1 aHuman Embryo and Stem Cell Laboratory, The Francis Crick Institute, NW1 1AT London, United Kingdom; bSex Chromosome Biology Laboratory, The Francis Crick Institute, NW1 1AT London, United Kingdom; cCentre for Stem Cells and Regenerative Medicine, Guy’s Campus, King’s College London, SE1 9RT London, United Kingdom; dNuffield Department of Women’s and Reproductive Health, John Radcliffe Hospital, University of Oxford, OX3 9DU Oxford, United Kingdom; eJesus College, University of Oxford, OX1 3DW Oxford, United Kingdom; fAncient Genomics Laboratory, The Francis Crick Institute, NW1 1AT London, United Kingdom; gJuno Genetics, OX4 4GE Oxford, United Kingdom; and hThe Centre for Trophoblast Research, Department of Physiology, Development and Neuroscience, University of Cambridge, CB2 3EG Cambridge, United Kingdom Edited by Barbara J. Meyer, University of California, Berkeley, CA, and approved October 31, 2020 (received for review June 5, 2020) CRISPR-Cas9 genome editing is a promising technique for clinical homozygous WT embryos in both cases was not associated with applications, such as the correction of disease-associated alleles in use of the provided repair template for gene correction. Instead, somatic cells. The use of this approach has also been discussed in the authors suggest that in edited embryos the WT maternal the context of heritable editing of the human germ line. However, allele served as a template for the high-fidelity homology di- studies assessing gene correction in early human embryos report rected repair (HDR) pathway to repair the double-strand lesion low efficiency of mutation repair, high rates of mosaicism, and the caused by the Cas9 protein in the paternal allele (8). -

Supplementary Materials: Molecular Signature of Subtypes of Non- Small Cell Lung Cancer by Large-Scale Transcriptional Profiling
Cancers 2020 S1 of S18 Supplementary Materials: Molecular Signature of Subtypes of Non- Small Cell Lung Cancer by Large-Scale Transcriptional Profiling: Identification of Key Modules and Genes by Weighted Gene Co- Expression Network Analysis (WGCNA) Magdalena Niemira, Francois Collin, Anna Szalkowska, Agnieszka Bielska, Karolina Chwialkowska, Joanna Reszec, Jacek Niklinski, Miroslaw Kwasniewski and Adam Kretowski Cancers 2020 S2 of S18 A B Figure S1. The top-ranked enriched canonical pathway identified in (A) SCC and (B) ADC using IPA: Eicosanoid signalling pathway. Cancers 2020 S3 of S18 A Cancers 2020 S4 of S18 Figure S2. The second-ranked enriched canonical pathway identified in (A) SCC and (B) ADC using IPA: Agranulocyte adhesion and diapedesis. Cancers 2020 S5 of S18 Figure S3. The top-ranked enriched canonical pathway identified only in lung ADC: MIF regulation of innate immunity. A B Figure S4. Cluster dendograms of the gene clusters of (A) LUAD and (B) LUSC subset from TCGA database. Cancers 2020 S6 of S18 A B C D Figure S5. Protein-protein interaction (PPI) network of genes in the red (A), lightcyan (B), darkorange (C), yellow (D) modules in ADC. The networks were constructed using Cytoscape v. 3.7.2. software. Cancers 2020 S7 of S18 A B Figure S6. Protein-protein interaction (PPI) network of genes in the blue (A) and (B) modules in SCC. The networks were constructed using Cytoscape v. 3.7.2. software. Cancers 2020 S8 of S18 Table S1. Upstream regulator analysis of DEGs in lung SCC predicted by IPA. Upstream Prediction Target