Qt1zv1x22m.Pdf
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Small Cell Ovarian Carcinoma: Genomic Stability and Responsiveness to Therapeutics
Gamwell et al. Orphanet Journal of Rare Diseases 2013, 8:33 http://www.ojrd.com/content/8/1/33 RESEARCH Open Access Small cell ovarian carcinoma: genomic stability and responsiveness to therapeutics Lisa F Gamwell1,2, Karen Gambaro3, Maria Merziotis2, Colleen Crane2, Suzanna L Arcand4, Valerie Bourada1,2, Christopher Davis2, Jeremy A Squire6, David G Huntsman7,8, Patricia N Tonin3,4,5 and Barbara C Vanderhyden1,2* Abstract Background: The biology of small cell ovarian carcinoma of the hypercalcemic type (SCCOHT), which is a rare and aggressive form of ovarian cancer, is poorly understood. Tumourigenicity, in vitro growth characteristics, genetic and genomic anomalies, and sensitivity to standard and novel chemotherapeutic treatments were investigated in the unique SCCOHT cell line, BIN-67, to provide further insight in the biology of this rare type of ovarian cancer. Method: The tumourigenic potential of BIN-67 cells was determined and the tumours formed in a xenograft model was compared to human SCCOHT. DNA sequencing, spectral karyotyping and high density SNP array analysis was performed. The sensitivity of the BIN-67 cells to standard chemotherapeutic agents and to vesicular stomatitis virus (VSV) and the JX-594 vaccinia virus was tested. Results: BIN-67 cells were capable of forming spheroids in hanging drop cultures. When xenografted into immunodeficient mice, BIN-67 cells developed into tumours that reflected the hypercalcemia and histology of human SCCOHT, notably intense expression of WT-1 and vimentin, and lack of expression of inhibin. Somatic mutations in TP53 and the most common activating mutations in KRAS and BRAF were not found in BIN-67 cells by DNA sequencing. -

Neuronal Pentraxin 2: a Synapse-Derived CSF Biomarker in Genetic Frontotemporal Dementia
J Neurol Neurosurg Psychiatry: first published as 10.1136/jnnp-2019-322493 on 9 April 2020. Downloaded from Neurodegeneration ORIGINAL RESEARCH Neuronal pentraxin 2: a synapse- derived CSF biomarker in genetic frontotemporal dementia Emma L van der Ende ,1 Meifang Xiao,2 Desheng Xu,2 Jackie M Poos ,1 Jessica L Panman,1,3 Lize C Jiskoot ,1,4 Lieke H Meeter,1 Elise GP Dopper,1 Janne M Papma,1 Carolin Heller ,5 Rhian Convery ,4 Katrina Moore,4 Martina Bocchetta ,4 Mollie Neason,4 Georgia Peakman,4 David M Cash,4 Charlotte E Teunissen,6 Caroline Graff,7,8 Matthis Synofzik,9,10 Fermin Moreno,11 Elizabeth Finger ,12 Raquel Sánchez- Valle,13 Rik Vandenberghe,14 Robert Laforce Jr,15 Mario Masellis,16 Maria Carmela Tartaglia,17 James B Rowe ,18 Christopher R Butler,19 Simon Ducharme,20 Alex Gerhard,21,22 Adrian Danek,23 Johannes Levin,23,24,25 Yolande AL Pijnenburg,26 Markus Otto ,27 Barbara Borroni ,28 Fabrizio Tagliavini,29 Alexandre de Mendonca,30 Isabel Santana,31 Daniela Galimberti,32,33 Harro Seelaar,1 Jonathan D Rohrer,4 Paul F Worley,2,34 John C van Swieten ,1 on behalf of the Genetic Frontotemporal Dementia Initiative (GENFI) ► Additional material is ABSTRACT INTRODUCTION published online only. To view Introduction Synapse dysfunction is emerging as an Frontotemporal dementia (FTD), a common form please visit the journal online (http:// dx. doi. org/ 10. 1136/ early pathological event in frontotemporal dementia of early-onset dementia, has an autosomal dominant jnnp- 2019- 322493). (FTD), however biomarkers are lacking. We aimed inheritance in 20%–30% of patients, most often to investigate the value of cerebrospinal fluid (CSF) due to mutations in granulin (GRN), chromosome For numbered affiliations see neuronal pentraxins (NPTXs), a family of proteins 9 open reading frame 72 (C9orf72) or microtubule- end of article. -
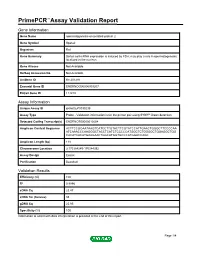
Primepcr™Assay Validation Report
PrimePCR™Assay Validation Report Gene Information Gene Name spermatogenesis-associated protein 2 Gene Symbol Spata2 Organism Rat Gene Summary Sertoli cell mRNA expression is induced by FSH; may play a role in spermatogenesis; localized to the nucleus Gene Aliases Not Available RefSeq Accession No. Not Available UniGene ID Rn.201291 Ensembl Gene ID ENSRNOG00000009207 Entrez Gene ID 114210 Assay Information Unique Assay ID qRnoCEP0030238 Assay Type Probe - Validation information is for the primer pair using SYBR® Green detection Detected Coding Transcript(s) ENSRNOT00000012604 Amplicon Context Sequence ACTTCCGGAATAAGTCATCCTTGTACTTCGTATCCATTGAACTGGGCTTCCCCAA ATCAAACCCAAGGGCTACCTCATCTCCCCCATGGCTCTGGGGCTGGAGGCTGG CACATCACATGAAGAACTGGCATGGTGCCCATGGACCAGC Amplicon Length (bp) 118 Chromosome Location 3:170384245-170384392 Assay Design Exonic Purification Desalted Validation Results Efficiency (%) 100 R2 0.9996 cDNA Cq 23.47 cDNA Tm (Celsius) 85 gDNA Cq 25.95 Specificity (%) 100 Information to assist with data interpretation is provided at the end of this report. Page 1/4 PrimePCR™Assay Validation Report Spata2, Rat Amplification Plot Amplification of cDNA generated from 25 ng of universal reference RNA Melt Peak Melt curve analysis of above amplification Standard Curve Standard curve generated using 20 million copies of template diluted 10-fold to 20 copies Page 2/4 PrimePCR™Assay Validation Report Products used to generate validation data Real-Time PCR Instrument CFX384 Real-Time PCR Detection System Reverse Transcription Reagent iScript™ Advanced cDNA Synthesis Kit for RT-qPCR Real-Time PCR Supermix SsoAdvanced™ SYBR® Green Supermix Experimental Sample qPCR Reference Total RNA Data Interpretation Unique Assay ID This is a unique identifier that can be used to identify the assay in the literature and online. Detected Coding Transcript(s) This is a list of the Ensembl transcript ID(s) that this assay will detect. -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

Genetics of Amyotrophic Lateral Sclerosis in the Han Chinese
Genetics of amyotrophic lateral sclerosis in the Han Chinese Ji He A thesis submitted for the degree of Master of Philosophy at The University of Queensland in 2015 The University of Queensland Diamantina Institute 1 Abstract Amyotrophic lateral sclerosis is the most frequently occurring neuromuscular degenerative disorders, and has an obscure aetiology. Whilst major progress has been made, the majority of the genetic variation involved in ALS is, as yet, undefined. In this thesis, multiple genetic studies have been conducted to advance our understanding of the genetic architecture of the disease. In the light of the paucity of comprehensive genetic studies performed in Chinese, the presented study focused on advancing our current understanding in genetics of ALS in the Han Chinese population. To identify genetic variants altering risk of ALS, a genome-wide association study (GWAS) was performed. The study included 1,324 Chinese ALS cases and 3,115 controls. After quality control, a number of analyses were performed in a cleaned dataset of 1,243 cases and 2,854 controls that included: a genome-wide association analysis to identify SNPs associated with ALS; a genomic restricted maximum likelihood (GREML) analysis to estimate the proportion of the phenotypic variance in ALS liability due to common SNPs; and a gene- based analysis to identify genes associated with ALS. There were no genome-wide significant SNPs or genes associated with ALS. However, it was estimated that 17% (SE: 0.05; P=6×10-5) of the phenotypic variance in ALS liability was due to common SNPs. The top associated SNP was within GNAS (rs4812037; p =7×10-7). -

Protein Kinase A-Mediated Septin7 Phosphorylation Disrupts Septin Filaments and Ciliogenesis
cells Article Protein Kinase A-Mediated Septin7 Phosphorylation Disrupts Septin Filaments and Ciliogenesis Han-Yu Wang 1,2, Chun-Hsiang Lin 1, Yi-Ru Shen 1, Ting-Yu Chen 2,3, Chia-Yih Wang 2,3,* and Pao-Lin Kuo 1,2,4,* 1 Department of Obstetrics and Gynecology, College of Medicine, National Cheng Kung University, Tainan 701, Taiwan; [email protected] (H.-Y.W.); [email protected] (C.-H.L.); [email protected] (Y.-R.S.) 2 Institute of Basic Medical Sciences, College of Medicine, National Cheng Kung University, Tainan 701, Taiwan; [email protected] 3 Department of Cell Biology and Anatomy, College of Medicine, National Cheng Kung University, Tainan 701, Taiwan 4 Department of Obstetrics and Gynecology, National Cheng-Kung University Hospital, Tainan 704, Taiwan * Correspondence: [email protected] (C.-Y.W.); [email protected] (P.-L.K.); Tel.: +886-6-2353535 (ext. 5338); (C.-Y.W.)+886-6-2353535 (ext. 5262) (P.-L.K.) Abstract: Septins are GTP-binding proteins that form heteromeric filaments for proper cell growth and migration. Among the septins, septin7 (SEPT7) is an important component of all septin filaments. Here we show that protein kinase A (PKA) phosphorylates SEPT7 at Thr197, thus disrupting septin filament dynamics and ciliogenesis. The Thr197 residue of SEPT7, a PKA phosphorylating site, was conserved among different species. Treatment with cAMP or overexpression of PKA catalytic subunit (PKACA2) induced SEPT7 phosphorylation, followed by disruption of septin filament formation. Constitutive phosphorylation of SEPT7 at Thr197 reduced SEPT7-SEPT7 interaction, but did not affect SEPT7-SEPT6-SEPT2 or SEPT4 interaction. -

1 Supporting Information for a Microrna Network Regulates
Supporting Information for A microRNA Network Regulates Expression and Biosynthesis of CFTR and CFTR-ΔF508 Shyam Ramachandrana,b, Philip H. Karpc, Peng Jiangc, Lynda S. Ostedgaardc, Amy E. Walza, John T. Fishere, Shaf Keshavjeeh, Kim A. Lennoxi, Ashley M. Jacobii, Scott D. Rosei, Mark A. Behlkei, Michael J. Welshb,c,d,g, Yi Xingb,c,f, Paul B. McCray Jr.a,b,c Author Affiliations: Department of Pediatricsa, Interdisciplinary Program in Geneticsb, Departments of Internal Medicinec, Molecular Physiology and Biophysicsd, Anatomy and Cell Biologye, Biomedical Engineeringf, Howard Hughes Medical Instituteg, Carver College of Medicine, University of Iowa, Iowa City, IA-52242 Division of Thoracic Surgeryh, Toronto General Hospital, University Health Network, University of Toronto, Toronto, Canada-M5G 2C4 Integrated DNA Technologiesi, Coralville, IA-52241 To whom correspondence should be addressed: Email: [email protected] (M.J.W.); yi- [email protected] (Y.X.); Email: [email protected] (P.B.M.) This PDF file includes: Materials and Methods References Fig. S1. miR-138 regulates SIN3A in a dose-dependent and site-specific manner. Fig. S2. miR-138 regulates endogenous SIN3A protein expression. Fig. S3. miR-138 regulates endogenous CFTR protein expression in Calu-3 cells. Fig. S4. miR-138 regulates endogenous CFTR protein expression in primary human airway epithelia. Fig. S5. miR-138 regulates CFTR expression in HeLa cells. Fig. S6. miR-138 regulates CFTR expression in HEK293T cells. Fig. S7. HeLa cells exhibit CFTR channel activity. Fig. S8. miR-138 improves CFTR processing. Fig. S9. miR-138 improves CFTR-ΔF508 processing. Fig. S10. SIN3A inhibition yields partial rescue of Cl- transport in CF epithelia. -

Regulation of Synapse Development by Activity Dependent Transcription in Inhibitory Neurons
Regulation of Synapse Development by Activity Dependent Transcription in Inhibitory Neurons The Harvard community has made this article openly available. Please share how this access benefits you. Your story matters Citation Mardinly, Alan Robert. 2013. Regulation of Synapse Development by Activity Dependent Transcription in Inhibitory Neurons. Doctoral dissertation, Harvard University. Citable link http://nrs.harvard.edu/urn-3:HUL.InstRepos:11151534 Terms of Use This article was downloaded from Harvard University’s DASH repository, and is made available under the terms and conditions applicable to Other Posted Material, as set forth at http:// nrs.harvard.edu/urn-3:HUL.InstRepos:dash.current.terms-of- use#LAA 2013 Alan Robert Mardinly All rights reserved Dissertation Advisor: Professor Michael E. Greenberg Alan Robert Mardinly Regulation of Synapse Development by Activity Dependent Transcription in Inhibitory Neurons Abstract Neuronal activity and subsequent calcium influx activates a signaling cascade that causes transcription factors in the nucleus to rapidly induce an early-response program of gene expression. This early-response program is composed of transcriptional regulators that in turn induce transcription of late-response genes, which are enriched for regulators of synaptic development and plasticity that act locally at the synapse. The main focus of this thesis is to describe the activity-induced transcriptional program in inhibitory neurons and identify its functional roles. We find strong similarity of the early-induced transcription factors induced by activity in excitatory and inhibitory neurons, but significant differences in the late-response genes. This finding suggests that commonly induced transcription factors may regulate distinct sets of target genes to perform cell-type specific functional roles. -

Genome-Wide Identification and Characterization of Functional Neuronal Activity–Dependent Enhancers
ART ic LE S Genome-wide identification and characterization of functional neuronal activity–dependent enhancers Athar N Malik1–3,7, Thomas Vierbuchen1,7, Martin Hemberg4, Alex A Rubin1, Emi Ling1,5, Cameron H Couch1, Hume Stroud1, Ivo Spiegel1, Kyle Kai-How Farh1,6, David A Harmin1 & Michael E Greenberg1 Experience-dependent gene transcription is required for nervous system development and function. However, the DNA regulatory elements that control this program of gene expression are not well defined. Here we characterize the enhancers that function across the genome to mediate activity-dependent transcription in mouse cortical neurons. We find that the subset of enhancers enriched for monomethylation of histone H3 Lys4 (H3K4me1) and binding of the transcriptional coactivator CREBBP (also called CBP) that shows increased acetylation of histone H3 Lys27 (H3K27ac) after membrane depolarization of cortical neurons functions to regulate activity-dependent transcription. A subset of these enhancers appears to require binding of FOS, which was previously thought to bind primarily to promoters. These findings suggest that FOS functions at enhancers to control activity- dependent gene programs that are critical for nervous system function and provide a resource of functional cis-regulatory elements that may give insight into the genetic variants that contribute to brain development and disease. In the nervous system, sensory experience promotes both the refine- There is still a striking gap in our understanding of how early ment and maturation of synaptic connections and the subsequent mod- response transcription factors such as FOS, C-JUN, NR4A1 and EGR1 ification of neural circuits during learning, memory and behavior1–6. -

Protein Interactions in the Cancer Proteome† Cite This: Mol
Molecular BioSystems View Article Online PAPER View Journal | View Issue Small-molecule binding sites to explore protein– protein interactions in the cancer proteome† Cite this: Mol. BioSyst., 2016, 12,3067 David Xu,ab Shadia I. Jalal,c George W. Sledge Jr.d and Samy O. Meroueh*aef The Cancer Genome Atlas (TCGA) offers an unprecedented opportunity to identify small-molecule binding sites on proteins with overexpressed mRNA levels that correlate with poor survival. Here, we analyze RNA-seq and clinical data for 10 tumor types to identify genes that are both overexpressed and correlate with patient survival. Protein products of these genes were scanned for binding sites that possess shape and physicochemical properties that can accommodate small-molecule probes or therapeutic agents (druggable). These binding sites were classified as enzyme active sites (ENZ), protein–protein interaction sites (PPI), or other sites whose function is unknown (OTH). Interestingly, the overwhelming majority of binding sites were classified as OTH. We find that ENZ, PPI, and OTH binding sites often occurred on the same structure suggesting that many of these OTH cavities can be used for allosteric modulation of Creative Commons Attribution 3.0 Unported Licence. enzyme activity or protein–protein interactions with small molecules. We discovered several ENZ (PYCR1, QPRT,andHSPA6)andPPI(CASC5, ZBTB32,andCSAD) binding sites on proteins that have been seldom explored in cancer. We also found proteins that have been extensively studied in cancer that have not been previously explored with small molecules that harbor ENZ (PKMYT1, STEAP3,andNNMT) and PPI (HNF4A, MEF2B,andCBX2) binding sites. All binding sites were classified by the signaling pathways to Received 29th March 2016, which the protein that harbors them belongs using KEGG. -

Mouse Spata2 Conditional Knockout Project (CRISPR/Cas9)
https://www.alphaknockout.com Mouse Spata2 Conditional Knockout Project (CRISPR/Cas9) Objective: To create a Spata2 conditional knockout Mouse model (C57BL/6J) by CRISPR/Cas-mediated genome engineering. Strategy summary: The Spata2 gene (NCBI Reference Sequence: NM_170756 ; Ensembl: ENSMUSG00000047030 ) is located on Mouse chromosome 2. 3 exons are identified, with the ATG start codon in exon 2 and the TAG stop codon in exon 3 (Transcript: ENSMUST00000057627). Exon 2~3 will be selected as conditional knockout region (cKO region). Deletion of this region should result in the loss of function of the Mouse Spata2 gene. To engineer the targeting vector, homologous arms and cKO region will be generated by PCR using BAC clone RP24-144E24 as template. Cas9, gRNA and targeting vector will be co-injected into fertilized eggs for cKO Mouse production. The pups will be genotyped by PCR followed by sequencing analysis. Note: Homozygous knockout leads to small testes, oligospermia, asthenozoospermia, reduced male fertility and decreased male germ cell numbers. It also affects necroptosis and increases inflammatory responses. Exon 2~3 covers 100.0% of the coding region. Start codon is in exon 2, and stop codon is in exon 3. The size of intron 1 for 5'-loxP site insertion: 7165 bp. The size of effective cKO region: ~2396 bp. The cKO region does not have any other known gene. Page 1 of 8 https://www.alphaknockout.com Overview of the Targeting Strategy gRNA region Wildtype allele T A 5' gRNA region G 3' 1 2 3 Targeting vector T A G Targeted allele T A G Constitutive KO allele (After Cre recombination) Legends Exon of mouse Spata2 Homology arm cKO region loxP site Page 2 of 8 https://www.alphaknockout.com Overview of the Dot Plot Window size: 10 bp Forward Reverse Complement Sequence 12 Note: The sequence of homologous arms and cKO region is aligned with itself to determine if there are tandem repeats. -

Supplementary Table S4. FGA Co-Expressed Gene List in LUAD
Supplementary Table S4. FGA co-expressed gene list in LUAD tumors Symbol R Locus Description FGG 0.919 4q28 fibrinogen gamma chain FGL1 0.635 8p22 fibrinogen-like 1 SLC7A2 0.536 8p22 solute carrier family 7 (cationic amino acid transporter, y+ system), member 2 DUSP4 0.521 8p12-p11 dual specificity phosphatase 4 HAL 0.51 12q22-q24.1histidine ammonia-lyase PDE4D 0.499 5q12 phosphodiesterase 4D, cAMP-specific FURIN 0.497 15q26.1 furin (paired basic amino acid cleaving enzyme) CPS1 0.49 2q35 carbamoyl-phosphate synthase 1, mitochondrial TESC 0.478 12q24.22 tescalcin INHA 0.465 2q35 inhibin, alpha S100P 0.461 4p16 S100 calcium binding protein P VPS37A 0.447 8p22 vacuolar protein sorting 37 homolog A (S. cerevisiae) SLC16A14 0.447 2q36.3 solute carrier family 16, member 14 PPARGC1A 0.443 4p15.1 peroxisome proliferator-activated receptor gamma, coactivator 1 alpha SIK1 0.435 21q22.3 salt-inducible kinase 1 IRS2 0.434 13q34 insulin receptor substrate 2 RND1 0.433 12q12 Rho family GTPase 1 HGD 0.433 3q13.33 homogentisate 1,2-dioxygenase PTP4A1 0.432 6q12 protein tyrosine phosphatase type IVA, member 1 C8orf4 0.428 8p11.2 chromosome 8 open reading frame 4 DDC 0.427 7p12.2 dopa decarboxylase (aromatic L-amino acid decarboxylase) TACC2 0.427 10q26 transforming, acidic coiled-coil containing protein 2 MUC13 0.422 3q21.2 mucin 13, cell surface associated C5 0.412 9q33-q34 complement component 5 NR4A2 0.412 2q22-q23 nuclear receptor subfamily 4, group A, member 2 EYS 0.411 6q12 eyes shut homolog (Drosophila) GPX2 0.406 14q24.1 glutathione peroxidase