Snapshot: the Replisome Nina Y
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Introduction of Human Telomerase Reverse Transcriptase to Normal Human Fibroblasts Enhances DNA Repair Capacity
Vol. 10, 2551–2560, April 1, 2004 Clinical Cancer Research 2551 Introduction of Human Telomerase Reverse Transcriptase to Normal Human Fibroblasts Enhances DNA Repair Capacity Ki-Hyuk Shin,1 Mo K. Kang,1 Erica Dicterow,1 INTRODUCTION Ayako Kameta,1 Marcel A. Baluda,1 and Telomerase, which consists of the catalytic protein subunit, No-Hee Park1,2 human telomerase reverse transcriptase (hTERT), the RNA component of telomerase (hTR), and several associated pro- 1School of Dentistry and 2Jonsson Comprehensive Cancer Center, University of California, Los Angeles, California teins, has been primarily associated with maintaining the integ- rity of cellular DNA telomeres in normal cells (1, 2). Telomer- ase activity is correlated with the expression of hTERT, but not ABSTRACT with that of hTR (3, 4). Purpose: From numerous reports on proteins involved The involvement of DNA repair proteins in telomere main- in DNA repair and telomere maintenance that physically tenance has been well documented (5–8). In eukaryotic cells, associate with human telomerase reverse transcriptase nonhomologous end-joining requires a DNA ligase and the (hTERT), we inferred that hTERT/telomerase might play a DNA-activated protein kinase, which is recruited to the DNA role in DNA repair. We investigated this possibility in nor- ends by the DNA-binding protein Ku. Ku binds to hTERT mal human oral fibroblasts (NHOF) with and without ec- without the need for telomeric DNA or hTR (9), binds the topic expression of hTERT/telomerase. telomere repeat-binding proteins TRF1 (10) and TRF2 (11), and Experimental Design: To study the effect of hTERT/ is thought to regulate the access of telomerase to telomere DNA telomerase on DNA repair, we examined the mutation fre- ends (12, 13). -

Replisome Assembly at Oric, the Replication Origin of E. Coli, Reveals an Explanation for Initiation Sites Outside an Origin
Molecular Cell, Vol. 4, 541±553, October, 1999, Copyright 1999 by Cell Press Replisome Assembly at oriC, the Replication Origin of E. coli, Reveals an Explanation for Initiation Sites outside an Origin Linhua Fang,*§ Megan J. Davey,² and Mike O'Donnell²³ have not been addressed. For example, is the local un- *Microbiology Department winding sufficiently large for two helicases to assemble Joan and Sanford I. Weill Graduate School of Medical for bidirectional replication, or does one helicase need Sciences of Cornell University to enter first and expand the bubble via helicase action New York, New York 10021 to make room for the second helicase? The known rep- ² The Rockefeller University and licative helicases are hexameric and encircle ssDNA. Howard Hughes Medical Institute Which strand does the initial helicase(s) at the origin New York, New York 10021 encircle, and if there are two, how are they positioned relative to one another? Primases generally require at least transient interaction with helicase to function. Can Summary primase function with the helicase(s) directly after heli- case assembly at the origin, or must helicase-catalyzed This study outlines the events downstream of origin DNA unwinding occur prior to RNA primer synthesis? unwinding by DnaA, leading to assembly of two repli- Chromosomal replicases are comprised of a ring-shaped cation forks at the E. coli origin, oriC. We show that protein clamp that encircles DNA, a clamp-loading com- two hexamers of DnaB assemble onto the opposing plex that uses ATP to assemble the clamp around DNA, strands of the resulting bubble, expanding it further, and a DNA polymerase that binds the circular clamp, yet helicase action is not required. -

Saccharomyces Rrm3p, a 5 to 3 DNA Helicase That Promotes Replication
Downloaded from genesdev.cshlp.org on September 24, 2021 - Published by Cold Spring Harbor Laboratory Press Saccharomyces Rrm3p, a 5 to 3 DNA helicase that promotes replication fork progression through telomeric and subtelomeric DNA Andreas S. Ivessa,1 Jin-Qiu Zhou,1,2 Vince P. Schulz, Ellen K. Monson, and Virginia A. Zakian3 Department of Molecular Biology, Princeton University, Princeton, New Jersey 08544-1014, USA In wild-type Saccharomyces cerevisiae, replication forks slowed during their passage through telomeric C1–3A/TG1–3 tracts. This slowing was greatly exacerbated in the absence of RRM3, shown here to encode a 5 ,to 3 DNA helicase. Rrm3p-dependent fork progression was seen at a modified Chromosome VII-L telomere at the natural X-bearing Chromosome III-L telomere, and at Y-bearing telomeres. Loss of Rrm3p also resulted in replication fork pausing at specific sites in subtelomeric DNA, such as at inactive replication origins, and at internal tracts of C1–3A/TG1–3 DNA. The ATPase/helicase activity of Rrm3p was required for its role in telomeric and subtelomeric DNA replication. Because Rrm3p was telomere-associated in vivo, it likely has a direct role in telomere replication. [Key Words: Telomere; helicase; telomerase; replication; RRM3; yeast] Received February 7, 2002; revised version accepted April 10, 2002. Telomeres are the natural ends of eukaryotic chromo- Because conventional DNA polymerases cannot repli- somes. In most organisms, the very ends of chromo- cate the very ends of linear DNA molecules, special somes consist of simple repeated sequences. For ex- mechanisms are required to prevent the loss of terminal ample, Saccharomyces cerevisiae chromosomes end in DNA. -

The Architecture of a Eukaryotic Replisome
The Architecture of a Eukaryotic Replisome Jingchuan Sun1,2, Yi Shi3, Roxana E. Georgescu3,4, Zuanning Yuan1,2, Brian T. Chait3, Huilin Li*1,2, Michael E. O’Donnell*3,4 1 Biosciences Department, Brookhaven National Laboratory, Upton, New York, USA 2 Department of Biochemistry & Cell Biology, Stony Brook University, Stony Brook, New York, USA. 3 The Rockefeller University, 1230 York Avenue, New York, New York, USA. 4 Howard Hughes Medical Institute *Correspondence and requests for materials should be addressed to M.O.D. ([email protected]) or H.L. ([email protected]) ABSTRACT At the eukaryotic DNA replication fork, it is widely believed that the Cdc45-Mcm2-7-GINS (CMG) helicase leads the way in front to unwind DNA, and that DNA polymerases (Pol) trail behind the helicase. Here we use single particle electron microscopy to directly image a replisome. Contrary to expectations, the leading strand Pol ε is positioned ahead of CMG helicase, while Ctf4 and the lagging strand Pol α-primase (Pol α) are behind the helicase. This unexpected architecture indicates that the leading strand DNA travels a long distance before reaching Pol ε, it first threads through the Mcm2-7 ring, then makes a U-turn at the bottom to reach Pol ε at the top of CMG. Our work reveals an unexpected configuration of the eukaryotic replisome, suggests possible reasons for this architecture, and provides a basis for further structural and biochemical replisome studies. INTRODUCTION DNA is replicated by a multi-protein machinery referred to as a replisome 1,2. Replisomes contain a helicase to unwind DNA, DNA polymerases that synthesize the leading and lagging strands, and a primase that makes short primed sites to initiate DNA synthesis on both strands. -

USP7 Couples DNA Replication Termination to Mitotic Entry
bioRxiv preprint doi: https://doi.org/10.1101/305318; this version posted April 20, 2018. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. USP7 couples DNA replication termination to mitotic entry Antonio Galarreta1*, Emilio Lecona1*, Pablo Valledor1, Patricia Ubieto1,2, Vanesa Lafarga1, Julia Specks1 & Oscar Fernandez-Capetillo1,3 1Genomic Instability Group, Spanish National Cancer Research Centre (CNIO), Madrid 28029, Spain 2Current Address: DNA Replication Group, Spanish National Cancer Research Centre (CNIO), Madrid 28029, Spain 3Science for Life Laboratory, Division of Genome Biology, Department of Medical Biochemistry and Biophysics, Karolinska Institute, S-171 21 Stockholm, Sweden *Co-first authors Correspondence: E.L. ([email protected]) or O.F. ([email protected]) Lead Contact: Oscar Fernandez-Capetillo Spanish National Cancer Research Centre (CNIO) Melchor Fernandez Almagro, 3 Madrid 28029, Spain Tel.: +34.91.732.8000 Ext: 3480 Fax: +34.91.732.8028 Email: [email protected] KEYWORDS: USP7; CDK1; DNA REPLICATION; MITOSIS; S/M TRANSITION. bioRxiv preprint doi: https://doi.org/10.1101/305318; this version posted April 20, 2018. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. USP7 coordinates the S/M transition 2 SUMMARY To ensure a faithful segregation of chromosomes, DNA must be fully replicated before mitotic entry. However, how cells sense the completion of DNA replication and to what extent this is linked to the activation of the mitotic machinery remains poorly understood. We previously showed that USP7 is a replisome-associated deubiquitinase with an essential role in DNA replication. -

The Biochemical Activities of the Saccharomyces Cerevisiae Pif1 Helicase Are Regulated by Its N-Terminal Domain
G C A T T A C G G C A T genes Article The Biochemical Activities of the Saccharomyces cerevisiae Pif1 Helicase Are Regulated by Its N-Terminal Domain David G. Nickens y, Christopher W. Sausen y and Matthew L. Bochman * Molecular and Cellular Biochemistry Department, Indiana University, Bloomington, IN 47405, USA; [email protected] (D.G.N.); [email protected] (C.W.S.) * Correspondence: [email protected] These authors contributed equally to this work. y Received: 31 March 2019; Accepted: 20 May 2019; Published: 28 May 2019 Abstract: Pif1 family helicases represent a highly conserved class of enzymes involved in multiple aspects of genome maintenance. Many Pif1 helicases are multi-domain proteins, but the functions of their non-helicase domains are poorly understood. Here, we characterized how the N-terminal domain (NTD) of the Saccharomyces cerevisiae Pif1 helicase affects its functions both in vivo and in vitro. Removal of the Pif1 NTD alleviated the toxicity associated with Pif1 overexpression in yeast. Biochemically, the N-terminally truncated Pif1 (Pif1DN) retained in vitro DNA binding, DNA unwinding, and telomerase regulation activities, but these activities differed markedly from those displayed by full-length recombinant Pif1. However, Pif1DN was still able to synergize with the Hrq1 helicase to inhibit telomerase activity in vitro, similar to full-length Pif1. These data impact our understanding of Pif1 helicase evolution and the roles of these enzymes in the maintenance of genome integrity. Keywords: DNA helicase; Saccharomyces cerevisiae; Pif1; telomerase; telomere 1. Introduction DNA helicases are enzymes that couple DNA binding and ATP hydrolysis to unwind double-stranded DNA (dsDNA) into its component single strands [1]. -

Structure of the Human Clamp Loader Reveals an Autoinhibited Conformation of a Substrate-Bound AAA+ Switch
Structure of the human clamp loader reveals an autoinhibited conformation of a substrate-bound AAA+ switch Christl Gaubitza,1, Xingchen Liua,b,1, Joseph Magrinoa,b, Nicholas P. Stonea, Jacob Landecka,b, Mark Hedglinc, and Brian A. Kelcha,2 aDepartment of Biochemistry and Molecular Pharmacology, University of Massachusetts Medical School, Worcester MA 01605; bGraduate School of Biomedical Sciences, University of Massachusetts Medical School, Worcester MA 01605; and cDepartment of Chemistry, The Pennsylvania State University, University Park, PA 16802 Edited by Michael E. O’Donnell, HHMI and Rockefeller University, New York, NY, and approved July 27, 2020 (received for review April 20, 2020) DNA replication requires the sliding clamp, a ring-shaped protein areflexia syndrome (15), Hutchinson–Gilford progeria syn- complex that encircles DNA, where it acts as an essential cofactor drome (16), and in the replication of some viruses (17–19). It for DNA polymerases and other proteins. The sliding clamp needs is unknown whether loading by RFC contributes to PARD to be opened and installed onto DNA by a clamp loader ATPase of disease. the AAA+ family. The human clamp loader replication factor C Clamp loaders are members of the AAA+ family of ATPases (RFC) and sliding clamp proliferating cell nuclear antigen (PCNA) (ATPases associated with various cellular activities), a large are both essential and play critical roles in several diseases. De- protein family that uses the chemical energy of adenosine 5′- spite decades of study, no structure of human RFC has been re- triphosphate (ATP) to generate mechanical force (20). Most solved. Here, we report the structure of human RFC bound to AAA+ proteins form hexameric motors that use an undulating PCNA by cryogenic electron microscopy to an overall resolution ∼ spiral staircase mechanism to processively translocate a substrate of 3.4 Å. -

M1224-100 Thermostable Rnase H
BioVision 03/17 For research use only Thermostable RNAse H CATALOG NO.: M1224-100 10X THERMOSTABLE RNAse H REACTION BUFFER: 500 mM Tris-HCl, 1000 mM NaCl, AMOUNT: 500 U (100 µl) 100 mM MgCl2, pH 7.5 PRODUCT SOURCE: Recombinant E. coli REACTION CONDITIONS: Use 1X Thermostable RNAse H Reaction Buffer and incubate CONCENTRATION: 100 U/µl at a chosen temperature between 65°C and 95°C (enzyme half-life is 2 hours at 70°C and 30 minutes at 95°C). FORM: Liquid. Enzyme supplied with 10X Reaction Buffer COMPONENTS: Product Name Size Part. No. Thermostable RNAse H (5 U/μl) 100 µl M1224-100-1 10X Thermostable RNAse Reaction Buffer 500 µl M1224-100-2 DESCRIPTION: Thermostable RNAse H (Ribonuclease H) is an endoribonuclease that specifically hydrolyzes the phosphodiester bonds of RNA strands in RNA-DNA hybrids. Unlike E. coli RNAse H which is inactivated at temperatures above 55°C, Thermostable RNase H can withstand much higher temperatures. These higher temperatures allow for higher hybridization stringency for RNA-DNA heteroduplexes resulting in more specific hydrolysis of RNA. Thermostable RNase H has optimal activity above 65°C and is active up to 95°C making it useful for a broad range of applications. APPLICATIONS: 1. High-stringency hybrid selection and mapping of mRNA structure 2. Removal of mRNA prior to synthesis of second strand cDNA 3. Removal of the poly(A) sequences from mRNA in the presence of oligo (dT) 4. Directed cleavage of RNA RELATED PRODUCTS: STORAGE CONDITIONS: Store at -20°C. Avoid repeated freeze-thaw cycles of all E. -

Arthur Kornberg Discovered (The First) DNA Polymerase Four
Arthur Kornberg discovered (the first) DNA polymerase Using an “in vitro” system for DNA polymerase activity: 1. Grow E. coli 2. Break open cells 3. Prepare soluble extract 4. Fractionate extract to resolve different proteins from each other; repeat; repeat 5. Search for DNA polymerase activity using an biochemical assay: incorporate radioactive building blocks into DNA chains Four requirements of DNA-templated (DNA-dependent) DNA polymerases • single-stranded template • deoxyribonucleotides with 5’ triphosphate (dNTPs) • magnesium ions • annealed primer with 3’ OH Synthesis ONLY occurs in the 5’-3’ direction Fig 4-1 E. coli DNA polymerase I 5’-3’ polymerase activity Primer has a 3’-OH Incoming dNTP has a 5’ triphosphate Pyrophosphate (PP) is lost when dNMP adds to the chain E. coli DNA polymerase I: 3 separable enzyme activities in 3 protein domains 5’-3’ polymerase + 3’-5’ exonuclease = Klenow fragment N C 5’-3’ exonuclease Fig 4-3 E. coli DNA polymerase I 3’-5’ exonuclease Opposite polarity compared to polymerase: polymerase activity must stop to allow 3’-5’ exonuclease activity No dNTP can be re-made in reversed 3’-5’ direction: dNMP released by hydrolysis of phosphodiester backboneFig 4-4 Proof-reading (editing) of misincorporated 3’ dNMP by the 3’-5’ exonuclease Fidelity is accuracy of template-cognate dNTP selection. It depends on the polymerase active site structure and the balance of competing polymerase and exonuclease activities. A mismatch disfavors extension and favors the exonuclease.Fig 4-5 Superimposed structure of the Klenow fragment of DNA pol I with two different DNAs “Fingers” “Thumb” “Palm” red/orange helix: 3’ in red is elongating blue/cyan helix: 3’ in blue is getting edited Fig 4-6 E. -

Transcription-Replication Collisions—A Series of Unfortunate Events
biomolecules Review Transcription-Replication Collisions—A Series of Unfortunate Events Commodore St Germain 1,2,*, Hongchang Zhao 2 and Jacqueline H. Barlow 2,* 1 School of Mathematics and Science, Solano Community College, 4000 Suisun Valley Road, Fairfield, CA 94534, USA 2 Department of Microbiology and Molecular Genetics, University of California Davis, One Shields Avenue, Davis, CA 95616, USA; [email protected] * Correspondence: [email protected] (C.S.G.); [email protected] (J.H.B.) Abstract: Transcription-replication interactions occur when DNA replication encounters genomic regions undergoing transcription. Both replication and transcription are essential for life and use the same DNA template making conflicts unavoidable. R-loops, DNA supercoiling, DNA secondary structure, and chromatin-binding proteins are all potential obstacles for processive replication or transcription and pose an even more potent threat to genome integrity when these processes co-occur. It is critical to maintaining high fidelity and processivity of transcription and replication while navigating through a complex chromatin environment, highlighting the importance of defining cellular pathways regulating transcription-replication interaction formation, evasion, and resolution. Here we discuss how transcription influences replication fork stability, and the safeguards that have evolved to navigate transcription-replication interactions and maintain genome integrity in mammalian cells. Keywords: DNA replication; transcription; R-loops; replication stress Citation: St Germain, C.; Zhao, H.; Barlow, J.H. Transcription-Replication Collisions—A Series of Unfortunate Events. Biomolecules 2021, 11, 1249. 1. Introduction https://doi.org/10.3390/ From bacteria to humans, transcription has been identified as a source of genome biom11081249 instability, initially observed as spontaneous recombination referred to as transcription- associated recombination (TAR) [1]. -

Teacher Notes Science Nspired
DNA Replication TEACHER NOTES SCIENCE NSPIRED Science Objectives In this lesson, students will • Explore how the structure of DNA supports its semi-conservative replication • Identify the name and function of several key enzymes in DNA replication • Recognize the function of Okazaki fragments in DNA replication Vocabulary TI-Nspire™ Technology Skills: • Double helix • Leading strand • Download a TI-Nspire • Lagging strand • Polymerase I • Polymerase III • Helicase document • Base pairs • Okazaki Fragments • Open a document • Primase • Move between pages • Open Directions Box About the Lesson • Explore ‘Hot Spots’ • Using three simulations, students will interact with DNA replication to explore semi-conservative replication and identify Tech Tips: specific enzymes and their roles in replication. Assessments are Make sure that students know embedded in the activity to engage discussion and gauge how to move between pages by learning. pressing /¡ (left arrow) and • As a result, students will: /¢ (right arrow). • Learn the basic functions of the following DNA replication enzymes: helicase, primase, ligase, polymerase I and III. Lesson Materials: • Learn how the double helix reproduces in a semi- Student Activity • DNA_Replication_Student.do conservative fashion c • Learn the role and function of Okazaki fragments in • DNA_Replication_Student.pd replication of the lagging strand. f TI-Nspire document TI-Nspire™ Navigator™ • DNA_Replication.tns Not required. If using Navigator: • Send out the DNA_Replication.tns file. • Monitor student progress -
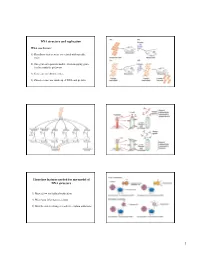
DNA Structure and Replication Three Key Features Needed for Any Model
DNA structure and replication What was known? 1) Hereditary factors were associated with specific traits 2) One-gene-one-protein model - from mapping genes for biosynthetic pathways 3) Genes are on chromosomes 4) Chromosomes are made up of DNA and protein Three key features needed for any model of DNA structure 1) Must allow for faithful replication 2) Must have information content 3) Must be able to change in order to explain mutations 1 Rosalind Franklin James Watson Francis Crick X-ray diffraction Maurice Wilkins Photograph of B-DNA Linus Pauling Nature 171, 737-738 (1953) © Macmillan Publishers Ltd. Molecular structure of Nucleic Acids WATSON, J. D. & CRICK, F. H. C. Features of the double helix A Structure for Deoxyribose Nucleic Acid 1) Two parallel strands We wish to suggest a structure for the salt of deoxyribose nucleic acid (D.N.A.). This structure has novel features which are of considerable biological interest………… 2) Bases held together by H-bonds 3) Phosphodiester backbone 4) The base is attached to the position 1 on the sugar 5) Base pair stack - provides Figure 1. This figure is purely diagrammatic. The two ribbons symbolize the two phophate-sugar chains, and the horizonal rods stability the pairs of bases holding the chains together. The vertical line marks the fibre axis. 6) Contains a major and …………….It has not escaped our notice that the specific pairing we minor groove have postulated immediately suggests a possible copying mechanism for the genetic material. Three key features needed for any model of DNA structure