Revealed by Chromosome Mapping of Rdna Loci
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

The Exclusively Neotropical Genus Scaphispatha Was Formerly Considered Monospecific. the Type Species
A REVISION OF SCAPHISPATHA (ARACEAE – CALADIEAE) INCLUDING A NEW SPECIES Eduardo Gomes Gonçalves1 ABSTRACT (A revision of Scaphispatha (Araceae – Caladieae) including a new species) The formerly considered monospecific genus Scaphispatha (Araceae – Caladieae) is here revised. Scaphispatha robusta E.G.Gonç, a second species for the genus is newly described from the Cerrado Biome and the transition Cerrado- Amazonia. It differs from S. gracilis Brongn. ex Schott by the much more robust petioles and leaves, primary lateral veins drying clearer than the lamina, lateral secondary veins conspicously more prominent than tertiary veins and for the female spadix with 11-15 rows of flowers visible in side view. A key to separate both species is provided, as well as ink illustrations and general remarks on the genus. Key-words: Scaphispatha, Cerrado, Caladieae, Araceae, geophyte. RESUMO (Revisão de Scaphispatha (Araceae – Caladieae), incluíndo a descrição de uma nova espécie para o gênero) O gênero Scaphispatha (Araceae – Caladieae), até então considerado monoespecífico, é aqui revisado. Scaphispatha robusta E.G.Gonç., uma segunda espécie para o gênero é descrita para o bioma Cerrado e a transição Cerrado-Amazonia. Difere de S. gracilis Brongn. ex Schott pelos pecíolos e folhas muito mais robustas, nervuras laterais mais claras que o limbo quando secas, nervuras laterais secundárias mais proemi- nentes que as terciárias e pela porção feminina do espádice com 11-15 espirais de flores visíveis em vista lateral. Uma chave para separar as espécies, assim como ilustrações em nanquim e aspectos gerais para o gênero são apresentados. Palavras-chave: Scaphispatha, Cerrado, Caladieae, Araceae, geófita. INTRODUCTION was recognized when plants from Pará state The exclusively neotropical genus (Northern Brazil) flowered in cultivation. -

History and Current Status of Systematic Research with Araceae
HISTORY AND CURRENT STATUS OF SYSTEMATIC RESEARCH WITH ARACEAE Thomas B. Croat Missouri Botanical Garden P. O. Box 299 St. Louis, MO 63166 U.S.A. Note: This paper, originally published in Aroideana Vol. 21, pp. 26–145 in 1998, is periodically updated onto the IAS web page with current additions. Any mistakes, proposed changes, or new publications that deal with the systematics of Araceae should be brought to my attention. Mail to me at the address listed above, or e-mail me at [email protected]. Last revised November 2004 INTRODUCTION The history of systematic work with Araceae has been previously covered by Nicolson (1987b), and was the subject of a chapter in the Genera of Araceae by Mayo, Bogner & Boyce (1997) and in Curtis's Botanical Magazine new series (Mayo et al., 1995). In addition to covering many of the principal players in the field of aroid research, Nicolson's paper dealt with the evolution of family concepts and gave a comparison of the then current modern systems of classification. The papers by Mayo, Bogner and Boyce were more comprehensive in scope than that of Nicolson, but still did not cover in great detail many of the participants in Araceae research. In contrast, this paper will cover all systematic and floristic work that deals with Araceae, which is known to me. It will not, in general, deal with agronomic papers on Araceae such as the rich literature on taro and its cultivation, nor will it deal with smaller papers of a technical nature or those dealing with pollination biology. -

ULEARUM SAGITTATUM Peter Boyce the Subject of This Plate Is a Little-Known Aroid Originating from Tropical South America
273. ULEARUM SAGITTATUM Peter Boyce The subject of this plate is a little-known aroid originating from tropical South America. Ulearum sagittatum Engl. (Engler, 1905) was based on material collected by the German botanist E.H.G. Ule from the Departamento Loreto in the Amazonian region of Peru. Ulearum is one of four genera in the Zomicarpeae, a tribe of the subfamily Aroideae, the others being Zomicarpa Schott (Schott, 1856), Zomicarpella N.E. Br. (Brown, 1881) and Filarum Nicolson (Nicolson, 1967). The genera are rare in the wild and very seldom seen in cultivation. Ulearum appears to be most closely related to Filarurn; both have seeds without endosperm whereas Zomicarpa and Zomicarpella pro- duce seeds with copious endosperm. Ulearum differs from Filarum in lacking a conspicuous elongated anther connective and in having a creeping rhizomatous, not tuberous, stem. The silver-variegated foliage is the most attractive feature of Ulearum, since the inflorescences are small and inconspicuous. Many of the aroids which are prized for their colourful and attractive foliage, such as Caladium Vent. and Dieffenbachia Schott, have soft, thin-textured leaves and are prone to fungal attack and leafdamage in cultivation. Ulearum, with its thin but tough-textured leaves, seems more resistant and is not affected by such problems, so it would appear that the plant has considerable horticultural poten- tial. CULTIVATION.The plants of Ulearum at Kew were presented by Josef Bogner of Munich Botanical Garden who received them from an amateur plant enthusiast in Brazil. Ulearum is a rain-forest plant and in cultivation requires a high temperature and constant humidity. -

Curriculum Vitae
CURRICULUM VITAE THOMAS B. CROAT, P. A. Schulze Curator of Botany Missouri Botanical Garden P.O. Box 299, St. Louis, Missouri 63166 USA; 314-577-5163 Personal Data Born: 23 May 1938, St. Mary’s, Iowa, U.S.A. Education 1956 U.S. Army Guided Missiles School, Ft. Sill, Oklahoma, radar operations AN-MPQ-10 1957 U.S. Army Signal School - Europe - Ansbach, Germany, electronics, radar repair B.A. 1962 Simpson College, Indianola, Iowa. Major, Biology; Minor, Chemistry. Professional teacher's certificate, University fellowships, NDEA student loans; 4 of 8 semesters on Dean's List; Tri-Beta, honorary biological fraternity M.A. 1966 University of Kansas, Lawrence, Kansas; NDEA fellowship; Botany participation in Organization for Tropical Studies Program, summer 1965, "Anatomy of Tropical Monocotyledons (Tomlinson). Thesis: The genus Solidago in Kansas Ph.D. 1967 University of Kansas; NDEA fellowship; NSF Evolutionary Biology. Botany Award (for fieldwork in Great Plains of U.S. and Canada). Thesis: The genus Solidago of the north central Great Plains Appointments and Teaching Experience 1956-1958 Radar Technician, U.S. Army 1962-1963 Teacher, Charlotte Amalie High School, St. Thomas, Virgin Island 1963-1964 Teacher, Knoxville Public Schools, Knoxville, Iowa 1966-1967 Assistant Instructor, University of Kansas 1967-1970 Research Associate, Center for the Biology of Natural Systems, Washington University, St. Louis 1967-1971 Assistant Botanist, Missouri Botanical Garden 1968-1971 Visiting Fellow, Smithsonian Tropical Research Institute 1970-1973 Adjunct Assistant Professor, Washington University, St. Louis 1970-1971 Curator, Summit Herbarium and Library, Canal Zone 1971-1976 Curator of Phanerogams, Missouri Botanical Garden 1972-l974 Member of NSF Advisory Committee for Resources in Systematic Botany 1974- Adjunct Faculty Appointment, University of Missouri, St. -
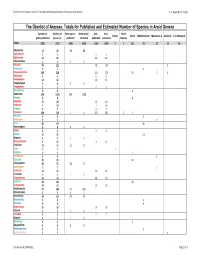
The Überlist of Araceae, Totals for Published and Estimated Number of Species in Aroid Genera P.C
The Überlist of Araceae, Totals for Published and Estimated Number of Species in Aroid Genera P.C. Boyce & T.B. Croat The Überlist of Araceae, Totals for Published and Estimated Number of Species in Aroid Genera Species per Species per Neotropical ‐ Neotropical ‐ Asia ‐ Asia ‐ North Boreal Africa Mediterranean Mascarene Is Australia Circumtropical genus published genus est. published estimated published estimated America Totals 3305 5422 1889 3143 1105 1968 2 7 152 78 20 23 29 Adelonema 13 20 13 20 Aglaodorum 11 11 Aglaonema 22 22 22 22 Alloschemone 2222 Alocasia 79 111 78 110 1 Ambrosina 11 1 Amorphophallus 196 218 153 175 35 7 1 Amydrium 55 55 Anadendrum 12 35 12 35 Anaphyllopsis 3434 Anaphyllum 22 22 Anchomanes 66 6 Anthurium 905 1500 905 1500 Anubias 88 8 Apoballis 12 20 12 20 Aridarum 716 716 Ariopsis 22 22 Arisaema 208 210 1 1 200 202 2 5 Arisarum 33 3 Arophyton 77 7 Arum 40 40 40 Asterostigma 8888 Bakoa 23 23 Biarum 21 21 21 Bognera 1111 Bucephalandra 315 315 Caladium 12 17 12 17 Calla 11 1 Callopsis 11 1 Carlephyton 33 3 Cercestis 10 10 10 Chlorospatha 28 71 28 71 Colletogyne 11 1 Colocasia 13 15 13 15 Croatiella 1111 Cryptocoryne 58 70 58 70 Culcasia 28 28 28 Cyrtosperma 13 13 13 13 Dieffenbachia 57 140 57 140 Dracontioides 2222 Dracontium 24 24 24 24 Dracunculus 22 2 Eminium 99 9 Epipremnum 15 18 15 18 Filarum 1111 Furtadoa 22 22 Gearum 1111 Gonatopus 55 5 Gorgonidium 8888 Gymnomesium 11 1 Gymnostachys 11 1 Current as of 10JAN2012 Page 1 of 3 The Überlist of Araceae, Totals for Published and Estimated Number of Species in Aroid Genera P.C. -

Leaf and Inflorescence Evidence for Near-Basal Araceae and an Unexpected Diversity of Other Monocots from the Late Early Cretaceous of Spain
Journal of Systematic Palaeontology ISSN: 1477-2019 (Print) 1478-0941 (Online) Journal homepage: http://www.tandfonline.com/loi/tjsp20 Leaf and inflorescence evidence for near-basal Araceae and an unexpected diversity of other monocots from the late Early Cretaceous of Spain Luis Miguel Sender, James A. Doyle, Garland R. Upchurch Jr, Uxue Villanueva- Amadoz & José B. Diez To cite this article: Luis Miguel Sender, James A. Doyle, Garland R. Upchurch Jr, Uxue Villanueva-Amadoz & José B. Diez (2018): Leaf and inflorescence evidence for near-basal Araceae and an unexpected diversity of other monocots from the late Early Cretaceous of Spain, Journal of Systematic Palaeontology, DOI: 10.1080/14772019.2018.1528999 To link to this article: https://doi.org/10.1080/14772019.2018.1528999 View supplementary material Published online: 09 Nov 2018. Submit your article to this journal View Crossmark data Full Terms & Conditions of access and use can be found at http://www.tandfonline.com/action/journalInformation?journalCode=tjsp20 Journal of Systematic Palaeontology, 2018 Vol. 0, No. 0, 1–34, http://doi.org/10.1080/14772019.2018.1528999 Leaf and inflorescence evidence for near-basal Araceae and an unexpected diversity of other monocots from the late Early Cretaceous of Spain aà b c d e Luis Miguel Sender , James A. Doyle , Garland R. Upchurch Jr , Uxue Villanueva-Amadoz and Jose B. Diez aDepartment of Biological Sciences, Faculty of Science and Engineering, Chuo University, 1-13-27 Kasuga, Bunkyo, Tokyo, Japan; bDepartment of Evolution and Ecology, -

On the Taxonomic Importance of Relocating Poorly Collected Species
96 AROIDEANA, Vol. 35 Lost Aroids: On the taxonomic importance of relocating poorly collected species Peter C. Boyce Pusat Pengajian Sains Kajihayat [School of Biological Sciences] Universiti Sains Malaysia 11800 USM Pulau Pinang, Malaysia [email protected] Wong Sin Yeng Department of Plant Science & Environmental Ecology Faculty of Resource Science & Technology Universiti Malaysia Sarawak 94300 Kota Samarahan, Sarawak, Malaysia [email protected] ABSTRACT Aroids, perhaps by reason of their often originating from almost inaccessible tropi- Aridarum montanum Ridl. and Piptos- cal forests, are host to a remarkable number patha insignis N.E.Br. (Araceae: Schisma- of such ‘lost’ species. Remarkable, too, is toglottideae), aroids originating from Bor- that quite some number of long-lost species neo that are each known from a single has been re-found over the past 20 years. collection, are discussed and illustrated. Of particular note [with the period ‘‘lost’’ in The history of their discovery is reviewed, together with what is known or speculated years] are: Gearum brasiliense N.E.Br. of their ecology. The biological significance [150 years] (Mayo et al., 1994), Mangonia of the collection locality of A. montanum is tweediana Schott [142 years] (Bogner & highlighted. The species’ individual impor- Marchesi, 2000), Zomicarpella maculata tance to modern systematics is highlighted. N.E.Br. [116 years] (Bogner, 2007, 2009), and Ulearum sagittatum Engl. [90 years] (Boyce, 1995; Bogner, 1997). KEY WORDS However, many aroid species remain Araceae, Aridarum, Piptospatha, Bor- elusive. Two of these, from Borneo, are the neo, Malaysia, Sarawak, Santubong. subject of this short piece. INTRODUCTION Aridarum montanum Ridl. –Figs.2and3 ‘Lost’ plant species – species tantalizingly In 1909 Cecil Joslin Brooks, a metallur- only known from a single herbarium col- gical chemist and competent amateur lection, or frustratingly from just an old botanist in the employ of the gold-mining illustration, hold an abiding fascination for division of the Borneo Co. -

Molecular Systematics and Historical Biogeography of Araceae at a Worldwide Scale and in Southeast Asia
Dissertation zur Erlangung des Doktorgrades an der Fakultät für Biologie der Ludwig-Maximilians-Universität München Molecular systematics and historical biogeography of Araceae at a worldwide scale and in Southeast Asia Lars Nauheimer München, 2. Juli 2012 Contents Table of Contents i Preface iv Statutory Declaration (Erklärung und ehrenwörtliche Versicherung) . iv List of Publications . .v Declaration of contribution as co-author . .v Notes ........................................... vi Summary . viii Zusammenfassung . ix 1 Introduction 1 General Introduction . .2 Estimating Divergence Times . .2 Fossil calibration . .2 Historical Biogeography . .3 Ancestral area reconstruction . .3 Incorporation of fossil ranges . .4 The Araceae Family . .5 General Introduction . .5 Taxonomy . .5 Biogeography . .6 The Malay Archipelago . .7 The Genus Alocasia ...................................8 Aim of this study . .9 Color plate . 10 2 Araceae 11 Global history of the ancient monocot family Araceae inferred with models accounting for past continental positions and previous ranges based on fossils 12 Supplementary Table 1: List of accessions . 25 Supplementary Table 2: List of Araceae fossils . 34 Supplementary Table 3: Dispersal matrices for ancestral area reconstruction 40 Supplementary Table 4: Results of divergence dating . 42 Supplementary Table 5: Results of ancestral area reconstructions . 45 Supplementary Figure 1: Inferred DNA substitution rates . 57 Supplementary Figure 2: Chronogram and AAR without fossil inclusion . 58 Supplementary Figure 3: Posterior distribution of fossil constraints . 59 3 Alocasia 61 Giant taro and its relatives - A phylogeny of the large genus Alocasia (Araceae) sheds light on Miocene floristic exchange in the Malesian region . 62 Supplementary Table 1: List of accessions . 71 i CONTENTS Supplementary Table 2: Crown ages of major nodes . 74 Supplementary Table 3: Clade support, divergence time estimates and ancestral area reconstruction . -

Universidade Federal De Pernambuco Centro De Biociências Programa De Pós-Graduacão Em Genética Emanuelle Varão Vasconcelos
UNIVERSIDADE FEDERAL DE PERNAMBUCO CENTRO DE BIOCIÊNCIAS PROGRAMA DE PÓS-GRADUACÃO EM GENÉTICA EMANUELLE VARÃO VASCONCELOS DNA repetitivo na evolução cariotípica de espécies de Philodendron Schott e Thaumatophyllum Schott (Araceae) Recife 2018 EMANUELLE VARÃO VASCONCELOS DNA repetitivo na evolução cariotípica de espécies de Philodendron Schott e Thaumatophyllum Schott (Araceae) Tese de Doutorado apresentada ao Programa de Pós- Graduação em Genética da Universidade Federal de Pernambuco como parte dos requisitos exigidos para obtenção do título de Doutor em Genética. Orientadora: Profa Dra Ana Christina Brasileiro-Vidal Coorientadores: Profa Dra Ana Maria Benko-Iseppon Dr Santelmo Selmo de Vasconcelos Júnior Recife 2018 Dados Internacionais de Catalogação na Publicação (CIP) de acordo com ISBD Vasconcelos, Emanuelle Varão DNA repetitivo na evolução cariotípica de espécies Philodendron Schott e Thaumatophyllum Schott (Araceae) / Emanuelle Varão Vasconcelos. – 2018. 123 f. : il. Orientador: Profa. Dra. Ana Christina Brasileiro-Vidal. Coorientadora: Profa. Dra. Ana Maria Benko-Iseppon. Coorientador: Dr. Santelmo Selmo de Vasconcelos Júnior. Tese (doutorado) – Universidade Federal de Pernambuco. Centro de Biociências. Programa de Pós-graduação em Genética, Recife, 2018. Inclui referências. 1. Genética vegetal. 2. Biologia – Classificação. 3. Plantas – Variação. I. Brasileiro-Vidal, Ana Christina (Orientadora). II. Benko-Iseppon, Ana Maria (Coorientadora). III. Vasconcelos Júnior, Santelmo Selmo (Coorientador). III. Título. 581.35 CDD (22.ed.) UFPE/CB – 2018 - 405 Elaborado por Bruno Márcio Gouveia - CRB-4/1788 EMANUELLE VARÃO VASCONCELOS DNA repetitivo na evolução cariotípica de espécies de Philodendron Schott e Thaumatophyllum Schott (Araceae) Tese de Doutorado apresentada ao Programa de Pós- Graduação em Genética da Universidade Federal de Pernambuco como parte dos requisitos exigidos para obtenção do título de Doutor em Genética. -

Plastome Phylogeny Monocots SI Tables
Givnish et al. – American Journal of Botany – Appendix S2. Taxa included in the across- monocots study and sources of sequence data. Sources not included in the main bibliography are listed at the foot of this table. Order Famiy Species Authority Source Acorales Acoraceae Acorus americanus (Raf.) Raf. Leebens-Mack et al. 2005 Acorus calamus L. Goremykin et al. 2005 Alismatales Alismataceae Alisma triviale Pursh Ross et al. 2016 Astonia australiensis (Aston) S.W.L.Jacobs Ross et al. 2016 Baldellia ranunculoides (L.) Parl. Ross et al. 2016 Butomopsis latifolia (D.Don) Kunth Ross et al. 2016 Caldesia oligococca (F.Muell.) Buchanan Ross et al. 2016 Damasonium minus (R.Br.) Buchenau Ross et al. 2016 Echinodorus amazonicus Rataj Ross et al. 2016 (Rusby) Lehtonen & Helanthium bolivianum Myllys Ross et al. 2016 (Humb. & Bonpl. ex Hydrocleys nymphoides Willd.) Buchenau Ross et al. 2016 Limnocharis flava (L.) Buchenau Ross et al. 2016 Luronium natans Raf. Ross et al. 2016 (Rich. ex Kunth) Ranalisma humile Hutch. Ross et al. 2016 Sagittaria latifolia Willd. Ross et al. 2016 Wiesneria triandra (Dalzell) Micheli Ross et al. 2016 Aponogetonaceae Aponogeton distachyos L.f. Ross et al. 2016 Araceae Aglaonema costatum N.E.Br. Henriquez et al. 2014 Aglaonema modestum Schott ex Engl. Henriquez et al. 2014 Aglaonema nitidum (Jack) Kunth Henriquez et al. 2014 Alocasia fornicata (Roxb.) Schott Henriquez et al. 2014 (K.Koch & C.D.Bouché) K.Koch Alocasia navicularis & C.D.Bouché Henriquez et al. 2014 Amorphophallus titanum (Becc.) Becc. Henriquez et al. 2014 Anchomanes hookeri (Kunth) Schott Henriquez et al. 2014 Anthurium huixtlense Matuda Henriquez et al. -

Interstitial Telomerelike Repeats in the Monocot Family Araceae
bs_bs_banner Botanical Journal of the Linnean Society, 2015, 177, 15–26. With 3 figures Interstitial telomere-like repeats in the monocot family Araceae ARETUZA SOUSA* and SUSANNE S. RENNER Department of Biology, University of Munich (LMU), 80638 Munich, Germany Received 6 May 2014; revised 26 September 2014; accepted for publication 4 October 2014 Combining molecular cytogenetics and phylogenetic modelling of chromosome number change can shed light on the types of evolutionary changes that may explain the haploid numbers observed today. Applied to the monocot family Araceae, with chromosome numbers of 2n = 8 to 2n = 160, this type of approach has suggested that descending dysploidy has played a larger role than polyploidy in the evolution of the current chromosome numbers. To test this, we carried out molecular cytogenetic analyses in 14 species from 11 genera, using probes for telomere repeats, 5S rDNA and 45S rDNA and a plastid phylogenetic tree covering the 118 genera of the family, many with multiple species. We obtained new chromosome counts for six species, modelled chromosome number evolution using all available counts for the family and carried out fluorescence in situ hybridization with three probes (5S rDNA, 45S rDNA and Arabidopsis-like telomeres) on 14 species with 2n =14to2n = 60. The ancestral state reconstruction provides support for a large role of descending dysploidy in Araceae, and interstitial telomere repeats (ITRs) were detected in Anthurium leuconerum, A. wendlingeri and Spathyphyllum tenerum, all with 2n = 30. The number of ITR signals in Anthurium (up to 12) is the highest so far reported in angiosperms, and the large repeats located in the pericentromeric regions of A. -

Genera Species Per Genus Accepted Species Per Genus Anticipated Neotropical
Boyce, P. C. Croat, T. B. (2011 onwards).The Überlist of Araceae, Totals for Published and Estimated Number of Species in Aroid Genera. http://www.aroid.org/genera/180211uberlist.pdf 1/2 Totals 3645 6489 2113 4129 1212 2042 1 8 150 80 23 3 26 29 Genera Species per genus Species per genus Neotropical ‐ Neotropical ‐ Asia ‐ Asia ‐ Circumboreal N. America Africa Mediterranean Madagascar Mascarene Is Australia Circumtropical accepted anticipated published anticipated published anticipated Adelonema 13 20 13 20 Aglaodorum 11 11 Aglaonema 22 25 22 25 Alloschemone 2222 Alocasia 78 121 77 120 1 Ambrosina 11 1 Amorphophallus 197 219 153 175 35 8 1 Amydrium 55 55 Anadendrum 12 35 12 35 Anaphyllopsis 3333 Anaphyllum 22 22 Anchomanes 66 6 Anthurium 950 2000 950 2000 Anubias 88 8 Apoballis 12 20 12 20 Aridarum 34 34 Ariopsis 22 22 Arisaema 185 209 1 1 176 200 3 5 Arisarum 33 3 Arophyton 77 7 Arum 40 40 40 Asterostigma 8888 Bakoa 11 11 Bakoaella 35 35 Biarum 23 23 23 Bognera 1111 Bucephalandra 27 50 27 50 Burttianthus 610 610 Caladium 12 17 12 17 Calla 11 1 Callopsis 11 1 Carlephyton 55 5 Cercestis 10 10 10 Chlorospatha 68 90 68 90 Colletogyne 11 1 Colobogynium 11 11 Colocasia 13 20 13 20 Croatiella 1111 Cryptocoryne 58 75 58 75 Culcasia 28 28 28 Cyrtosperma 13 13 13 13 Dieffenbachia 57 140 57 140 Dracontioides 2222 Dracontium 26 30 26 30 Dracunculus 22 2 Eminium 99 9 Englerarum 11 11 Epipremnum 15 18 15 18 Fenestratarum 25 25 Filarum 1111 Furtadoa 35 35 Galantharum 13 13 Gamogyne 710 710 Gearum 1111 Gonatopus 55 5 Gorgonidium 8888 Gosong 23 23 Gymnomesium 11 1 Gymnostachys 11 1 Hapaline 88 88 Helicodiceros 11 1 Hera 11 11 Heteroaridarum 34 34 Heteropsis 17 20 17 20 Holochlamys 11 11 Homalomena 98 500 98 500 Hottarum 13 13 Idimanthus 01 1 Incarum 1111 Jasarum 1111 Kiewia 33 33 Lagenandra 15 15 15 15 Landoltia 11 1 Lasia 11 11 <Lasia> concinna 11 11 Lasimorpha 11 1 Lazarum 18 18 18 Lemna 13 13 13 180211uberlist.xlsx 2/11/2018 Boyce, P.