G-Protein-Coupled Receptor Signaling and Polarized Actin Dynamics Drive
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

G Protein Alpha 13 (GNA13) (NM 006572) Human Tagged ORF Clone Lentiviral Particle Product Data
OriGene Technologies, Inc. 9620 Medical Center Drive, Ste 200 Rockville, MD 20850, US Phone: +1-888-267-4436 [email protected] EU: [email protected] CN: [email protected] Product datasheet for RC207762L3V G protein alpha 13 (GNA13) (NM_006572) Human Tagged ORF Clone Lentiviral Particle Product data: Product Type: Lentiviral Particles Product Name: G protein alpha 13 (GNA13) (NM_006572) Human Tagged ORF Clone Lentiviral Particle Symbol: GNA13 Synonyms: G13 Vector: pLenti-C-Myc-DDK-P2A-Puro (PS100092) ACCN: NM_006572 ORF Size: 1131 bp ORF Nucleotide The ORF insert of this clone is exactly the same as(RC207762). Sequence: OTI Disclaimer: The molecular sequence of this clone aligns with the gene accession number as a point of reference only. However, individual transcript sequences of the same gene can differ through naturally occurring variations (e.g. polymorphisms), each with its own valid existence. This clone is substantially in agreement with the reference, but a complete review of all prevailing variants is recommended prior to use. More info OTI Annotation: This clone was engineered to express the complete ORF with an expression tag. Expression varies depending on the nature of the gene. RefSeq: NM_006572.3 RefSeq Size: 4744 bp RefSeq ORF: 1134 bp Locus ID: 10672 UniProt ID: Q14344, A0A024R8M0 Domains: G-alpha Protein Families: Druggable Genome Protein Pathways: Long-term depression, Regulation of actin cytoskeleton, Vascular smooth muscle contraction MW: 44 kDa This product is to be used for laboratory only. Not for diagnostic or therapeutic use. View online » ©2021 OriGene Technologies, Inc., 9620 Medical Center Drive, Ste 200, Rockville, MD 20850, US 1 / 2 G protein alpha 13 (GNA13) (NM_006572) Human Tagged ORF Clone Lentiviral Particle – RC207762L3V Gene Summary: Guanine nucleotide-binding proteins (G proteins) are involved as modulators or transducers in various transmembrane signaling systems (PubMed:15240885, PubMed:16787920, PubMed:16705036, PubMed:27084452). -

Entosis Enables a Population Response to Starvation
www.impactjournals.com/oncotarget/ Oncotarget, 2017, Vol. 8, (No. 35), pp: 57934-57935 Editorial Entosis enables a population response to starvation Jens C. Hamann and Michael Overholtzer Commentary on: Hamann et al. Entosis is induced by glucose starvation. Cell Rep. 2017; 20:201-210. https://doi.org/10.1016/j. celrep.2017.06.037 Cell death isn’t just about apoptosis anymore. In cells in a population to scavenge extracellular nutrients and the last decade, numerous alternative mechanisms have accumulate the biomass necessary to support proliferation emerged, including regulated forms of necrosis, as well (Figure 1). Macropinocytosis has been shown to act as entosis, a unique mechanism that is executed as a cell similarly to allow cancer cells to scavenge extracellular murder rather than cell suicide [1]. Hamann et al. now proteins [5]. Entosis instead supplies bulk nutrients in identify glucose starvation as a key inducer of this non- the form of whole cells, where, on average, winner cells cell-autonomous form of cell death [2]. ingest two losers, providing a large nutrient supply that Cells that undergo entosis are killed as a result of is well suited to support the outgrowth of selected cells ingestion into their neighbors. They are not phagocytosed, in the population. Indeed, winners with ingested losers but rather form junctions with their neighbors and then have a nearly 10-fold proliferative advantage, and entosis invade into them, ultimately becoming killed through a is required for population re-growth during long-term mechanism involving autophagy proteins and lysosomal starvation. enzymes [1, 3]. This is a competitive process that results Our findings identify entosis as a population-scale in elimination of stiffer cells (“losers”) by softer ones starvation response with parallels to cell competition (“winners”), an effect linked to control over entotic cell occurring in developing tissues [6]. -

PLXNB1 (Plexin
Atlas of Genetics and Cytogenetics in Oncology and Haematology OPEN ACCESS JOURNAL AT INIST-CNRS Gene Section Mini Review PLXNB1 (plexin B1) José Javier Gómez-Román, Montserrat Nicolas Martínez, Servando Lazuén Fernández, José Fernando Val-Bernal Department of Anatomical Pathology, Marques de Valdecilla University Hospital, Medical Faculty, University of Cantabria, Santander, Spain (JJGR, MN, SL, JFVB) Published in Atlas Database: March 2009 Online updated version: http://AtlasGeneticsOncology.org/Genes/PLXNB1ID43413ch3p21.html DOI: 10.4267/2042/44702 This work is licensed under a Creative Commons Attribution-Noncommercial-No Derivative Works 2.0 France Licence. © 2010 Atlas of Genetics and Cytogenetics in Oncology and Haematology Identity Pseudogene No. Other names: KIAA0407; MGC149167; OTTHUMP00000164806; PLEXIN-B1; PLXN5; SEP Protein HGNC (Hugo): PLXNB1 Location: 3p21.31 Description Local order: The Plexin B1 gene is located between 2135 Amino acids (AA). Plexins are receptors for axon FBXW12 and CCDC51 genes. molecular guidance molecules semaphorins. Plexin signalling is important in pathfinding and patterning of both neurons and developing blood vessels. Plexin-B1 is a surface cell receptor. When it binds to its ligand SEMA4D it activates several pathways by binding of cytoplasmic ligands, like RHOA activation and subsequent changes of the actin cytoskeleton, axon guidance, invasive growth and cell migration. It monomers and heterodimers with PLXNB2 after proteolytic processing. Binds RAC1 that has been activated by GTP binding. It binds PLXNA1 and by similarity ARHGEF11, Note ARHGEF12, ERBB2, MET, MST1R, RND1, NRP1 Size: 26,200 bases. and NRP2. Orientation: minus strand. This family features the C-terminal regions of various plexins. The cytoplasmic region, which has been called DNA/RNA a SEX domain in some members of this family is involved in downstream signalling pathways, by Description interaction with proteins such as Rac1, RhoD, Rnd1 and other plexins. -

CXCL13/CXCR5 Interaction Facilitates VCAM-1-Dependent Migration in Human Osteosarcoma
International Journal of Molecular Sciences Article CXCL13/CXCR5 Interaction Facilitates VCAM-1-Dependent Migration in Human Osteosarcoma 1, 2,3,4, 5 6 7 Ju-Fang Liu y, Chiang-Wen Lee y, Chih-Yang Lin , Chia-Chia Chao , Tsung-Ming Chang , Chien-Kuo Han 8, Yuan-Li Huang 8, Yi-Chin Fong 9,10,* and Chih-Hsin Tang 8,11,12,* 1 School of Oral Hygiene, College of Oral Medicine, Taipei Medical University, Taipei City 11031, Taiwan; [email protected] 2 Department of Orthopaedic Surgery, Chang Gung Memorial Hospital, Puzi City, Chiayi County 61363, Taiwan; [email protected] 3 Department of Nursing, Division of Basic Medical Sciences, and Chronic Diseases and Health Promotion Research Center, Chang Gung University of Science and Technology, Puzi City, Chiayi County 61363, Taiwan 4 Research Center for Industry of Human Ecology and Research Center for Chinese Herbal Medicine, Chang Gung University of Science and Technology, Guishan Dist., Taoyuan City 33303, Taiwan 5 School of Medicine, China Medical University, Taichung 40402, Taiwan; [email protected] 6 Department of Respiratory Therapy, Fu Jen Catholic University, New Taipei City 24205, Taiwan; [email protected] 7 School of Medicine, Institute of Physiology, National Yang-Ming University, Taipei City 11221, Taiwan; [email protected] 8 Department of Biotechnology, College of Health Science, Asia University, Taichung 40402, Taiwan; [email protected] (C.-K.H.); [email protected] (Y.-L.H.) 9 Department of Sports Medicine, College of Health Care, China Medical University, Taichung 40402, Taiwan 10 Department of Orthopedic Surgery, China Medical University Beigang Hospital, Yunlin 65152, Taiwan 11 Department of Pharmacology, School of Medicine, China Medical University, Taichung 40402, Taiwan 12 Chinese Medicine Research Center, China Medical University, Taichung 40402, Taiwan * Correspondence: [email protected] (Y.-C.F.); [email protected] (C.-H.T.); Tel.: +886-4-2205-2121-7726 (C.-H.T.); Fax: +886-4-2233-3641 (C.-H.T.) These authors contributed equally to this work. -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

Identification and Characterization of RHOA-Interacting Proteins in Bovine Spermatozoa1
BIOLOGY OF REPRODUCTION 78, 184–192 (2008) Published online before print 10 October 2007. DOI 10.1095/biolreprod.107.062943 Identification and Characterization of RHOA-Interacting Proteins in Bovine Spermatozoa1 Sarah E. Fiedler, Malini Bajpai, and Daniel W. Carr2 Department of Medicine, Oregon Health & Sciences University and Veterans Affairs Medical Center, Portland, Oregon 97239 ABSTRACT Guanine nucleotide exchange factors (GEFs) catalyze the GDP for GTP exchange [2]. Activation is negatively regulated by In somatic cells, RHOA mediates actin dynamics through a both guanine nucleotide dissociation inhibitors (RHO GDIs) GNA13-mediated signaling cascade involving RHO kinase and GTPase-activating proteins (GAPs) [1, 2]. Endogenous (ROCK), LIM kinase (LIMK), and cofilin. RHOA can be RHO can be inactivated via C3 exoenzyme ADP-ribosylation, negatively regulated by protein kinase A (PRKA), and it and studies have demonstrated RHO involvement in actin-based interacts with members of the A-kinase anchoring (AKAP) cytoskeletal response to extracellular signals, including lyso- family via intermediary proteins. In spermatozoa, actin poly- merization precedes the acrosome reaction, which is necessary phosphatidic acid (LPA) [2–4]. LPA is known to signal through for normal fertility. The present study was undertaken to G-protein-coupled receptors (GPCRs) [4, 5]; specifically, LPA- determine whether the GNA13-mediated RHOA signaling activated GNA13 (formerly Ga13) promotes RHO activation pathway may be involved in acrosome reaction in bovine through GEFs [4, 6]. Activated RHO-GTP then signals RHO caudal sperm, and whether AKAPs may be involved in its kinase (ROCK), resulting in the phosphorylation and activation targeting and regulation. GNA13, RHOA, ROCK2, LIMK2, and of LIM-kinase (LIMK), which in turn phosphorylates and cofilin were all detected by Western blot in bovine caudal inactivates cofilin, an actin depolymerizer, the end result being sperm. -

In Amyotrophic Lateral Sclerosis
ORIGINAL RESEARCH published: 25 March 2021 doi: 10.3389/fncel.2021.600872 Dual Role of Lysophosphatidic Acid Receptor 2 (LPA2) in Amyotrophic Lateral Sclerosis Maria Puigdomenech-Poch 1,2, Anna Martínez-Muriana 1, Pol Andrés-Benito 2,3, Isidre Ferrer 2,3, Jerold Chun 4 and Rubèn López-Vales 1,2* 1Departament de Biologia Cellular, Fisiologia i Immunologia, Institut de Neurociències, Centro de Investigación Biomédica en Red sobre Enfermedades Neurodegenerativas (CIBERNED), Universitat Autònoma de Barcelona, Bellaterra, Spain, 2Centro de Investigación Biomédica en Red de Enfermedades Neurodegenerativas (CIBERNED), Madrid, Spain, 3Departament de Patologia i Terapèutica Experimental, Hospital Universitari de Bellvitge, IDIBELL, L’Hospitalet de Llobregat, Universitat de Barcelona, Barcelona, Spain, 4Sanford Burnham Prebys Medical Discovery Institute, La Jolla, CA, United States Lysophosphatidic acid (LPA) is a pleiotropic extracellular lipid mediator with many Edited by: Ertugrul Kilic, physiological functions that signal through six known G protein-coupled receptors Istanbul Medipol University, Turkey (LPA1–6). In the central nervous system (CNS), LPA mediates a wide range of effects Reviewed by: including neural progenitor cell physiology, neuronal cell death, axonal retraction, and Savina Apolloni, inflammation. Since inflammation is a hallmark of most neurological conditions, we University of Rome Tor Vergata, Italy Mustafa Caglar Beker, hypothesized that LPA could be involved in the physiopathology of amyotrophic lateral Istanbul Medipol University, Turkey sclerosis (ALS). We found that LPA2 RNA was upregulated in post-mortem spinal cord *Correspondence: samples of ALS patients and in the sciatic nerve and skeletal muscle of SOD1G93A Rubèn López-Vales [email protected] mouse, the most widely used ALS mouse model. -

Targeting Lysophosphatidic Acid in Cancer: the Issues in Moving from Bench to Bedside
View metadata, citation and similar papers at core.ac.uk brought to you by CORE provided by IUPUIScholarWorks cancers Review Targeting Lysophosphatidic Acid in Cancer: The Issues in Moving from Bench to Bedside Yan Xu Department of Obstetrics and Gynecology, Indiana University School of Medicine, 950 W. Walnut Street R2-E380, Indianapolis, IN 46202, USA; [email protected]; Tel.: +1-317-274-3972 Received: 28 August 2019; Accepted: 8 October 2019; Published: 10 October 2019 Abstract: Since the clear demonstration of lysophosphatidic acid (LPA)’s pathological roles in cancer in the mid-1990s, more than 1000 papers relating LPA to various types of cancer were published. Through these studies, LPA was established as a target for cancer. Although LPA-related inhibitors entered clinical trials for fibrosis, the concept of targeting LPA is yet to be moved to clinical cancer treatment. The major challenges that we are facing in moving LPA application from bench to bedside include the intrinsic and complicated metabolic, functional, and signaling properties of LPA, as well as technical issues, which are discussed in this review. Potential strategies and perspectives to improve the translational progress are suggested. Despite these challenges, we are optimistic that LPA blockage, particularly in combination with other agents, is on the horizon to be incorporated into clinical applications. Keywords: Autotaxin (ATX); ovarian cancer (OC); cancer stem cell (CSC); electrospray ionization tandem mass spectrometry (ESI-MS/MS); G-protein coupled receptor (GPCR); lipid phosphate phosphatase enzymes (LPPs); lysophosphatidic acid (LPA); phospholipase A2 enzymes (PLA2s); nuclear receptor peroxisome proliferator-activated receptor (PPAR); sphingosine-1 phosphate (S1P) 1. -

ARHGEF12 Regulates Erythropoiesis and Is Involved in Erythroid Regeneration After Ferrata Storti Foundation Chemotherapy in Acute Lymphoblastic Leukemia Patients
Red Cell Biology & its Disorders ARTICLE ARHGEF12 regulates erythropoiesis and is involved in erythroid regeneration after Ferrata Storti Foundation chemotherapy in acute lymphoblastic leukemia patients Yangyang Xie,1* Li Gao,2* Chunhui Xu,3 Liming Chu,3,7 Lei Gao,4 Ruichi Wu,1 Yu Liu,1 Ting Liu,1 Xiao-jian Sun,5 Ruibao Ren,5 Jingyan Tang,1 Yi Zheng,6 Yong Zhou7 and Shuhong Shen1 1Key Lab of Pediatrics Hematology/Oncology, Ministry of Health, Department of Haematologica 2020 Hematology/Oncology, Shanghai Children's Medical Center, Shanghai Jiao Tong University, Volume 105(4):925-936 Shanghai, China; 2Department of Hematology and Oncology, Children's Hospital of Soochow University, Suzhou, China; 3Shanghai Institute of Nutrition and Health, University of Chinese Academy of Sciences, Chinese Academy of Sciences, Shanghai, China; 4CAS Key Laboratory of Genome Sciences and Information, Beijing Institute of Genomics, Beijing, China; 5State Key Laboratory for Medical Genomics, Shanghai Institute of Hematology, Ruijin Hospital, Shanghai, China; 6Division of Experimental Hematology and Cancer Biology, Cincinnati Children's Hospital Research Foundation, Cincinnati, OH, USA and 7CAS Key Laboratory of Tissue Microenvironment and Tumor, Shanghai Institute of Nutrition and Health, University of Chinese Academy of Sciences, Chinese Academy of Sciences, Shanghai, China *YX and LG contributed equally to this work. ABSTRACT ematopoiesis is a finely regulated process in vertebrates under both homeostatic and stress conditions. By whole exome sequencing, we Correspondence: Hstudied the genomics of acute lymphoblastic leukemia (ALL) YI ZHENG patients who needed multiple red blood cell (RBC) transfusions after inten- [email protected] sive chemotherapy treatment. ARHGEF12, encoding a RhoA guanine YONG ZHOU nucleotide exchange factor, was found to be associated with chemothera- [email protected] py-induced anemia by genome-wide association study analyses. -

Identification of Novel Potential Genes Involved in Cancer by Integrated
International Journal of Molecular Sciences Article Identification of Novel Potential Genes Involved in Cancer by Integrated Comparative Analyses Francesco Monticolo 1, Emanuela Palomba 2 and Maria Luisa Chiusano 1,2,* 1 Department of Agricultural Sciences, Università Degli Studi di Napoli Federico II, 80055 Naples, Italy; [email protected] 2 Department of RIMAR, Stazione Zoologica “Anton Dohrn”, 80122 Naples, Italy; [email protected] * Correspondence: [email protected] Received: 26 October 2020; Accepted: 11 December 2020; Published: 15 December 2020 Abstract: The main hallmarks of cancer diseases are the evasion of programmed cell death, uncontrolled cell division, and the ability to invade adjacent tissues. The explosion of omics technologies offers challenging opportunities to identify molecular agents and processes that may play relevant roles in cancer. They can support comparative investigations, in one or multiple experiments, exploiting evidence from one or multiple species. Here, we analyzed gene expression data from induction of programmed cell death and stress response in Homo sapiens and compared the results with Saccharomyces cerevisiae gene expression during the response to cell death. The aim was to identify conserved candidate genes associated with Homo sapiens cell death, favored by crosslinks based on orthology relationships between the two species. Weidentified differentially-expressed genes, pathways that are significantly dysregulated across treatments, and characterized genes among those involved in induced cell death. We investigated on co-expression patterns and identified novel genes that were not expected to be associated with death pathways, that have a conserved pattern of expression between the two species. Finally, we analyzed the resulting list by HumanNet and identified new genes predicted to be involved in cancer. -

G-Protein-Coupled Receptor Heteromer Dynamics
Commentary 4215 G-protein-coupled receptor heteromer dynamics Jean-Pierre Vilardaga1,2,*, Luigi F. Agnati3, Kjell Fuxe4 and Francisco Ciruela5 1Laboratory for GPCR Biology, Department of Pharmacology and Chemical Biology, University of Pittsburgh, School of Medicine, Pittsburgh, PA 15261, USA 2Endocrine Unit, Department of Medicine, Massachusetts General Hospital and Harvard Medical School, Boston, MA 02114, USA 3IRCCS, San Camillo, Lido Venezia, Italy 4Department of Neuroscience, Karolinska Institute, Stockholm SE-17177, Sweden 5Pharmacology Unit, Department of Pathology and Experimental Therapy, Faculty of Medicine, University of Barcelona, 08907 Barcelona, Spain *Author for correspondence ([email protected]) Journal of Cell Science 123, 000-000 © 2010. Published by The Company of Biologists Ltd doi:10.1242/jcs.063354 Summary G-protein-coupled receptors (GPCRs) represent the largest family of cell surface receptors, and have evolved to detect and transmit a large palette of extracellular chemical and sensory signals into cells. Activated receptors catalyze the activation of heterotrimeric G proteins, which modulate the propagation of second messenger molecules and the activity of ion channels. Classically thought to signal as monomers, different GPCRs often pair up with each other as homo- and heterodimers, which have been shown to modulate signaling to G proteins. Here, we discuss recent advances in GPCR heteromer systems involving the kinetics of the early steps in GPCR signal transduction, the dynamic property of receptor–receptor interactions, and how the formation of receptor heteromers modulate the kinetics of G-protein signaling. Key words: G-protein-coupled receptors, Heterodimers, Signaling Introduction the signaling and trafficking mechanisms of GPCRs is thus central G-protein-coupled receptors (GPCRs) constitute the main family for the development of new and safer therapies for many of cell surface receptors for a large variety of chemical stimuli physiological and psychological disorders. -
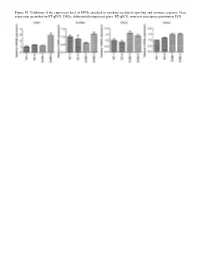
Figure S1. Validation of the Expression Level of Degs Enriched in Cytokine‑Mediated Signaling and Immune Response
Figure S1. Validation of the expression level of DEGs enriched in cytokine‑mediated signaling and immune response. Gene expression quantified by RT‑qPCR. DEGs, differentially expressed genes; RT‑qPCR, reverse‑transcription quantitative PCR. Figure S2. Validation of STAU1‑regulated AS events. IGV‑Sashimi plot revealed (A‑C) three A5SS AS events in three different genes. Reads distribution of each AS event was plotted in the left panel with the transcripts of each gene shown below. The sche‑ matic diagrams depict the structures of two AS events, AS1 (purple line) and AS2 (green line). The exon sequences are denoted by black boxes, the intron sequences by a horizontal line (right panel). RNA‑seq quantification and RT‑qPCR validation of ASEs are presented in the panels on the right. STAU1, double‑stranded RNA‑binding protein Staufen homolog 1; AS, alternative splicing; A5SS, alternative 5'splice site; RNA‑seq, RNA sequencing; RT‑qPCR, reverse‑transcription quantitative PCR. Error bars represent mean ± SEM. *P<0.05. Table SI. Primers used in gene validation experiments. IFIT2‑F CAGCCTACGGCAACTAAA IFIT2‑R GAGCCTTCTCAAAGCACA IFIT3‑F ACACCAAACAATGGCTAC IFIT3‑R TGGACAAACCCTCTAAAC OASL‑F AATGGTGACCGTGATGGG OASL‑R ACCTGAGGATGGAGCAGAG IFI27‑F TTCACTGCGGCGGGAATC IFI27‑R TGGCTGCTATGGAGGACGAG S1PR4‑F TGCTGAAGACGGTGCTGATG S1PR4‑R TGCGGAAGGAGTAGATGATGG CCL5‑F ACGACTGCTGGGTTGGAG CCL5‑R ACCCTGCTGCTTTGCCTA CCL2‑F CTAACCCAGAAACATCCAAT CCL2‑R GCTATGAGCAGCAGGCAC CD44‑F TGGAGGACAGAAAGCCAAGT CD44‑R TTCGCAATGAAACAATCAGTAG PLEKHG2‑M/As‑F CCAAAAGTAAGCCTGTCC PLEKHG2‑M‑R