Mapping Axon Initial Segment Structure and Function by Multiplexed Proximity Biotinylation
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Transcriptomic Analysis of Human Brain Microvascular Endothelial
www.nature.com/scientificreports OPEN Transcriptomic analysis of human brain microvascular endothelial cells exposed to laminin binding protein (adhesion lipoprotein) and Streptococcus pneumoniae Irene Jiménez‑Munguía1, Zuzana Tomečková1, Evelína Mochnáčová1, Katarína Bhide1, Petra Majerová2 & Mangesh Bhide1,2* Streptococcus pneumoniae invades the CNS and triggers a strong cellular response. To date, signaling events that occur in the human brain microvascular endothelial cells (hBMECs), in response to pneumococci or its surface adhesins are not mapped comprehensively. We evaluated the response of hBMECs to the adhesion lipoprotein (a laminin binding protein—Lbp) or live pneumococci. Lbp is a surface adhesin recently identifed as a potential ligand, which binds to the hBMECs. Transcriptomic analysis was performed by RNA‑seq of three independent biological replicates and validated with qRT‑PCR using 11 genes. In total 350 diferentially expressed genes (DEGs) were identifed after infection with S. pneumoniae, whereas 443 DEGs when challenged with Lbp. Total 231 DEGs were common in both treatments. Integrative functional analysis revealed participation of DEGs in cytokine, chemokine, TNF signaling pathways and phagosome formation. Moreover, Lbp induced cell senescence and breakdown, and remodeling of ECM. This is the frst report which maps complete picture of cell signaling events in the hBMECs triggered against S. pneumoniae and Lbp. The data obtained here could contribute in a better understanding of the invasion of pneumococci across BBB and underscores role of Lbp adhesin in evoking the gene expression in neurovascular unit. Streptococcus pneumoniae (also known as pneumococcus) is a life-threatening pathogen responsible for high morbidity and mortality rates worldwide1. It can cross the blood–brain barrier (BBB) and cause meningitis, commonly known as pneumococcal meningitis, a rare but life-threatening medical emergency. -

Supplementary Data
Supplemental Figure 1: In vitro viability data A C HDQ-P1 basal-like cells HDQ-P1 untreated HDQ-P1 40 0.04 uM MEKi 35 30 25 MCF7 20 15 CAL85-1 %BrdU 10 Incorporation 5 MDA-MB-231 0 MDA-MB-468 0 6E-05 3E-04 0.002 0.008 0.04 0.2 1 5 25 MEKi (ȝM) T47D luminal cells 50 T-47D untreated T-47D 40 0.2uM MEKi 30 20 PD0325901 (ȝM) %BrdU 10 Incorporation 0 0 6E-05 3E-04 0.002 0.008 0.04 0.2 1 5 25 MEKi (ȝM) B D HDQ-P1 Vehicle Treated 1PM MEKi 100 G1: 45.2% 1500 G1: 62.5% MCF7 1000 S: 4.2% S: 3.4% 75 BT474 800 G2/M: 34.9% 1000 G2/M: 27.4% MDA-MB-231 34.9 27.4 50 600 HDQ-P1 # Cells 4.17 #Cells 3.36 Cell viability (%) 400 CAL85-1 45.2 500 62.5 25 0.001 0.01 0.1 1 10 200 MEK inhibitor 2 (PM) 0 0 0 200 400 600 800 1000 0 200 400 600 800 1000 Propidium Iodide Propidium Iodide EGFR B A WEGFR Levels 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 --- MDA-MB-134VI Basal-like HDQ-P1 BT549 Basal-like Her2 DU4475 CAL51 + HCC1806 Supplemental Figure2:PTENandEGFRinBreastLines - MDA-MB-435s CAL85-1 + HCC70 CAL120 -- MX1 CAL148 CAL-120 + CAL85-1 HCC70 -- SW527 HCC1143 CAL-51 HCC1395 + MDA-MB-231 HCC1937 + HCC1143 + BT20 MDA-MB-231 + Hs578T MDA-MB-436 + MDA-MB-468 MDA-MB-468 - HCC1395 MFM223 + BT-549 - CAL-148 AU565 + HCC1937 BT474 + HCC38 EFM192A + MDA-MB-436 + HCC1954 HCC1419 + AU565 ------ HCC1569 BT474 Her2 orLuminal EFM-192A HCC1954 HCC1419 HCC2218 HCC1569 HCC202 MDA-MB-361 HCC2218 MDA-MB-453 + JIM-T --- SKBR3 KPL4 MDA-MB-361 UACC812 MDA-MB-453 UACC893 + SKBR3 ZR75-30 ---------- UACC-812 -

Supplementary Figures
Mena regulates the LINC complex to control actin–nuclear lamina associations, trans-nuclear membrane signalling and cancer gene expression Frederic Li Mow Chee!, Bruno Beernaert!, Alexander Loftus!, Yatendra Kumar", Billie G. C. Griffith!, Jimi C. Wills!, Ann P. Wheeler#, J. Douglas Armstrong$, Maddy Parsons%, Irene M. Leigh,(, Charlotte M. Proby&, Alex von Kriegsheim!, Wendy A. Bickmore", Margaret C. Frame,* & Adam Byron,* Supplementary Information Supplementary Figure 1 Supplementary Figure 2 Supplementary Figure 3 Supplementary Table 1 Supplementary Table 2 Supplementary Table 3 Supplementary Table 4 !Cancer Research UK Edinburgh Centre, Institute of Genetics and Cancer, University of Edinburgh, Edinburgh EH< =XR, UK. "MRC Human Genetics Unit, Institute of Genetics and Cancer, University of Edinburgh, Edinburgh EH< =XU, UK. #Advanced Imaging Resource, Institute of Genetics and Cancer, University of Edinburgh, Edinburgh EH< =XU, UK. $Simons Initiative for the Developing Brain, School of Informatics, University of Edinburgh, Edinburgh EHH IYL, UK. %Randall Centre for Cell and Molecular Biophysics, King’s College London, London SEM MUL, UK. &Division of Molecular and Clinical Medicine, School of Medicine, University of Dundee, Dundee DD <HN, UK. 'Institute of Dentistry, Barts and the London School of Medicine and Dentistry, Queen Mary University of London, London EM =AT, UK. *email: [email protected] or [email protected] 1 a cSCC IAC correlation b cSCC IAC pathways c Core adhesome network ENAH −log10(q) MACF1 CSRP1 Met1 Met4 0 5 10 + + CORO2A Integrin signalling + CFL1 pathway PRNP ILK + HSPB1 PALLD PPFIA1 TES RDX Cytoskeletal regulation + VASP + + ARPC2 by Rho GTPase PPP2CA + Met1 + LASP1 MYH9 + VIM TUBA4A Huntington ITGA3 + disease ITGB4 VCL CAV1 ACTB ROCK1 KTN1 FLNA+ CALR DNA FBLIM1 CORO1B RAC1 + replication +ACTN1 ITGA6 + Met4 ITGAV Parkinson ITGB1 disease Actin cytoskel. -

Universidade Estadual De Campinas Instituto De Biologia
UNIVERSIDADE ESTADUAL DE CAMPINAS INSTITUTO DE BIOLOGIA VERÔNICA APARECIDA MONTEIRO SAIA CEREDA O PROTEOMA DO CORPO CALOSO DA ESQUIZOFRENIA THE PROTEOME OF THE CORPUS CALLOSUM IN SCHIZOPHRENIA CAMPINAS 2016 1 VERÔNICA APARECIDA MONTEIRO SAIA CEREDA O PROTEOMA DO CORPO CALOSO DA ESQUIZOFRENIA THE PROTEOME OF THE CORPUS CALLOSUM IN SCHIZOPHRENIA Dissertação apresentada ao Instituto de Biologia da Universidade Estadual de Campinas como parte dos requisitos exigidos para a obtenção do Título de Mestra em Biologia Funcional e Molecular na área de concentração de Bioquímica. Dissertation presented to the Institute of Biology of the University of Campinas in partial fulfillment of the requirements for the degree of Master in Functional and Molecular Biology, in the area of Biochemistry. ESTE ARQUIVO DIGITAL CORRESPONDE À VERSÃO FINAL DA DISSERTAÇÃO DEFENDIDA PELA ALUNA VERÔNICA APARECIDA MONTEIRO SAIA CEREDA E ORIENTADA PELO DANIEL MARTINS-DE-SOUZA. Orientador: Daniel Martins-de-Souza CAMPINAS 2016 2 Agência(s) de fomento e nº(s) de processo(s): CNPq, 151787/2F2014-0 Ficha catalográfica Universidade Estadual de Campinas Biblioteca do Instituto de Biologia Mara Janaina de Oliveira - CRB 8/6972 Saia-Cereda, Verônica Aparecida Monteiro, 1988- Sa21p O proteoma do corpo caloso da esquizofrenia / Verônica Aparecida Monteiro Saia Cereda. – Campinas, SP : [s.n.], 2016. Orientador: Daniel Martins de Souza. Dissertação (mestrado) – Universidade Estadual de Campinas, Instituto de Biologia. 1. Esquizofrenia. 2. Espectrometria de massas. 3. Corpo caloso. -

Defining Functional Interactions During Biogenesis of Epithelial Junctions
ARTICLE Received 11 Dec 2015 | Accepted 13 Oct 2016 | Published 6 Dec 2016 | Updated 5 Jan 2017 DOI: 10.1038/ncomms13542 OPEN Defining functional interactions during biogenesis of epithelial junctions J.C. Erasmus1,*, S. Bruche1,*,w, L. Pizarro1,2,*, N. Maimari1,3,*, T. Poggioli1,w, C. Tomlinson4,J.Lees5, I. Zalivina1,w, A. Wheeler1,w, A. Alberts6, A. Russo2 & V.M.M. Braga1 In spite of extensive recent progress, a comprehensive understanding of how actin cytoskeleton remodelling supports stable junctions remains to be established. Here we design a platform that integrates actin functions with optimized phenotypic clustering and identify new cytoskeletal proteins, their functional hierarchy and pathways that modulate E-cadherin adhesion. Depletion of EEF1A, an actin bundling protein, increases E-cadherin levels at junctions without a corresponding reinforcement of cell–cell contacts. This unexpected result reflects a more dynamic and mobile junctional actin in EEF1A-depleted cells. A partner for EEF1A in cadherin contact maintenance is the formin DIAPH2, which interacts with EEF1A. In contrast, depletion of either the endocytic regulator TRIP10 or the Rho GTPase activator VAV2 reduces E-cadherin levels at junctions. TRIP10 binds to and requires VAV2 function for its junctional localization. Overall, we present new conceptual insights on junction stabilization, which integrate known and novel pathways with impact for epithelial morphogenesis, homeostasis and diseases. 1 National Heart and Lung Institute, Faculty of Medicine, Imperial College London, London SW7 2AZ, UK. 2 Computing Department, Imperial College London, London SW7 2AZ, UK. 3 Bioengineering Department, Faculty of Engineering, Imperial College London, London SW7 2AZ, UK. 4 Department of Surgery & Cancer, Faculty of Medicine, Imperial College London, London SW7 2AZ, UK. -

Absence of NEFL in Patient-Specific Neurons in Early-Onset Charcot-Marie-Tooth Neuropathy Markus T
ARTICLE OPEN ACCESS Absence of NEFL in patient-specific neurons in early-onset Charcot-Marie-Tooth neuropathy Markus T. Sainio, MSc, Emil Ylikallio, MD, PhD, Laura M¨aenp¨a¨a, MSc, Jenni Lahtela, PhD, Pirkko Mattila, PhD, Correspondence Mari Auranen, MD, PhD, Johanna Palmio, MD, PhD, and Henna Tyynismaa, PhD Dr. Tyynismaa [email protected] Neurol Genet 2018;4:e244. doi:10.1212/NXG.0000000000000244 Abstract Objective We used patient-specific neuronal cultures to characterize the molecular genetic mechanism of recessive nonsense mutations in neurofilament light (NEFL) underlying early-onset Charcot- Marie-Tooth (CMT) disease. Methods Motor neurons were differentiated from induced pluripotent stem cells of a patient with early- onset CMT carrying a novel homozygous nonsense mutation in NEFL. Quantitative PCR, protein analytics, immunocytochemistry, electron microscopy, and single-cell transcriptomics were used to investigate patient and control neurons. Results We show that the recessive nonsense mutation causes a nearly total loss of NEFL messenger RNA (mRNA), leading to the complete absence of NEFL protein in patient’s cultured neurons. Yet the cultured neurons were able to differentiate and form neuronal networks and neuro- filaments. Single-neuron gene expression fingerprinting pinpointed NEFL as the most down- regulated gene in the patient neurons and provided data of intermediate filament transcript abundancy and dynamics in cultured neurons. Blocking of nonsense-mediated decay partially rescued the loss of NEFL mRNA. Conclusions The strict neuronal specificity of neurofilament has hindered the mechanistic studies of re- cessive NEFL nonsense mutations. Here, we show that such mutation leads to the absence of NEFL, causing childhood-onset neuropathy through a loss-of-function mechanism. -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

Supplementary Materials
1 Supplementary Materials: Supplemental Figure 1. Gene expression profiles of kidneys in the Fcgr2b-/- and Fcgr2b-/-. Stinggt/gt mice. (A) A heat map of microarray data show the genes that significantly changed up to 2 fold compared between Fcgr2b-/- and Fcgr2b-/-. Stinggt/gt mice (N=4 mice per group; p<0.05). Data show in log2 (sample/wild-type). 2 Supplemental Figure 2. Sting signaling is essential for immuno-phenotypes of the Fcgr2b-/-lupus mice. (A-C) Flow cytometry analysis of splenocytes isolated from wild-type, Fcgr2b-/- and Fcgr2b-/-. Stinggt/gt mice at the age of 6-7 months (N= 13-14 per group). Data shown in the percentage of (A) CD4+ ICOS+ cells, (B) B220+ I-Ab+ cells and (C) CD138+ cells. Data show as mean ± SEM (*p < 0.05, **p<0.01 and ***p<0.001). 3 Supplemental Figure 3. Phenotypes of Sting activated dendritic cells. (A) Representative of western blot analysis from immunoprecipitation with Sting of Fcgr2b-/- mice (N= 4). The band was shown in STING protein of activated BMDC with DMXAA at 0, 3 and 6 hr. and phosphorylation of STING at Ser357. (B) Mass spectra of phosphorylation of STING at Ser357 of activated BMDC from Fcgr2b-/- mice after stimulated with DMXAA for 3 hour and followed by immunoprecipitation with STING. (C) Sting-activated BMDC were co-cultured with LYN inhibitor PP2 and analyzed by flow cytometry, which showed the mean fluorescence intensity (MFI) of IAb expressing DC (N = 3 mice per group). 4 Supplemental Table 1. Lists of up and down of regulated proteins Accession No. -

Circular RNA Hsa Circ 0005114‑Mir‑142‑3P/Mir‑590‑5P‑ Adenomatous
ONCOLOGY LETTERS 21: 58, 2021 Circular RNA hsa_circ_0005114‑miR‑142‑3p/miR‑590‑5p‑ adenomatous polyposis coli protein axis as a potential target for treatment of glioma BO WEI1*, LE WANG2* and JINGWEI ZHAO1 1Department of Neurosurgery, China‑Japan Union Hospital of Jilin University, Changchun, Jilin 130033; 2Department of Ophthalmology, The First Hospital of Jilin University, Jilin University, Changchun, Jilin 130021, P.R. China Received September 12, 2019; Accepted October 22, 2020 DOI: 10.3892/ol.2020.12320 Abstract. Glioma is the most common type of brain tumor APC expression with a good overall survival rate. UALCAN and is associated with a high mortality rate. Despite recent analysis using TCGA data of glioblastoma multiforme and the advances in treatment options, the overall prognosis in patients GSE25632 and GSE103229 microarray datasets showed that with glioma remains poor. Studies have suggested that circular hsa‑miR‑142‑3p/hsa‑miR‑590‑5p was upregulated and APC (circ)RNAs serve important roles in the development and was downregulated. Thus, hsa‑miR‑142‑3p/hsa‑miR‑590‑5p‑ progression of glioma and may have potential as therapeutic APC‑related circ/ceRNA axes may be important in glioma, targets. However, the expression profiles of circRNAs and their and hsa_circ_0005114 interacted with both of these miRNAs. functions in glioma have rarely been studied. The present study Functional analysis showed that hsa_circ_0005114 was aimed to screen differentially expressed circRNAs (DECs) involved in insulin secretion, while APC was associated with between glioma and normal brain tissues using sequencing the Wnt signaling pathway. In conclusion, hsa_circ_0005114‑ data collected from the Gene Expression Omnibus database miR‑142‑3p/miR‑590‑5p‑APC ceRNA axes may be potential (GSE86202 and GSE92322 datasets) and explain their mecha‑ targets for the treatment of glioma. -
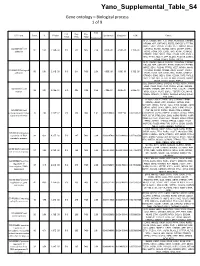
Supplemental Tables4.Pdf
Yano_Supplemental_Table_S4 Gene ontology – Biological process 1 of 9 Fold List Pop Pop GO Term Count % PValue Bonferroni Benjamini FDR Genes Total Hits Total Enrichment DLC1, CADM1, NELL2, CLSTN1, PCDHGA8, CTNNB1, NRCAM, APP, CNTNAP2, FERT2, RAPGEF1, PTPRM, MPDZ, SDK1, PCDH9, PTPRS, VEZT, NRXN1, MYH9, GO:0007155~cell CTNNA2, NCAM1, NCAM2, DDR1, LSAMP, CNTN1, 50 5.61 2.14E-08 510 311 7436 2.34 4.50E-05 4.50E-05 3.70E-05 adhesion ROR2, VCAN, DST, LIMS1, TNC, ASTN1, CTNND2, CTNND1, CDH2, NEO1, CDH4, CD24A, FAT3, PVRL3, TRO, TTYH1, MLLT4, LPP, NLGN1, PCDH19, LAMA1, ITGA9, CDH13, CDON, PSPC1 DLC1, CADM1, NELL2, CLSTN1, PCDHGA8, CTNNB1, NRCAM, APP, CNTNAP2, FERT2, RAPGEF1, PTPRM, MPDZ, SDK1, PCDH9, PTPRS, VEZT, NRXN1, MYH9, GO:0022610~biological CTNNA2, NCAM1, NCAM2, DDR1, LSAMP, CNTN1, 50 5.61 2.14E-08 510 311 7436 2.34 4.50E-05 4.50E-05 3.70E-05 adhesion ROR2, VCAN, DST, LIMS1, TNC, ASTN1, CTNND2, CTNND1, CDH2, NEO1, CDH4, CD24A, FAT3, PVRL3, TRO, TTYH1, MLLT4, LPP, NLGN1, PCDH19, LAMA1, ITGA9, CDH13, CDON, PSPC1 DCC, ENAH, PLXNA2, CAPZA2, ATP5B, ASTN1, PAX6, ZEB2, CDH2, CDH4, GLI3, CD24A, EPHB1, NRCAM, GO:0006928~cell CTTNBP2, EDNRB, APP, PTK2, ETV1, CLASP2, STRBP, 36 4.04 3.46E-07 510 205 7436 2.56 7.28E-04 3.64E-04 5.98E-04 motion NRG1, DCLK1, PLAT, SGPL1, TGFBR1, EVL, MYH9, YWHAE, NCKAP1, CTNNA2, SEMA6A, EPHA4, NDEL1, FYN, LRP6 PLXNA2, ADCY5, PAX6, GLI3, CTNNB1, LPHN2, EDNRB, LPHN3, APP, CSNK2A1, GPR45, NRG1, RAPGEF1, WWOX, SGPL1, TLE4, SPEN, NCAM1, DDR1, GRB10, GRM3, GNAQ, HIPK1, GNB1, HIPK2, PYGO1, GO:0007166~cell RNF138, ROR2, CNTN1, -

Phenomic Profiling Through Live-Cell Imaging in a Panel of Reporter Cell
www.nature.com/scientificreports OPEN Tales of 1,008 small molecules: phenomic profling through live‑cell imaging in a panel of reporter cell lines Michael J. Cox1,5, Stefen Jaensch1,5*, Jelle Van de Waeter1, Laure Cougnaud2, Daan Seynaeve2, Soulaiman Benalla1, Seong Joo Koo1, Ilse Van Den Wyngaert1, Jean‑Marc Neefs1, Dmitry Malkov3, Mart Bittremieux1, Margino Steemans1, Pieter J. Peeters1, Jörg Kurt Wegner1, Hugo Ceulemans1, Emmanuel Gustin1, Yolanda T. Chong1,4 & Hinrich W. H. Göhlmann1 Phenomic profles are high‑dimensional sets of readouts that can comprehensively capture the biological impact of chemical and genetic perturbations in cellular assay systems. Phenomic profling of compound libraries can be used for compound target identifcation or mechanism of action (MoA) prediction and other applications in drug discovery. To devise an economical set of phenomic profling assays, we assembled a library of 1,008 approved drugs and well‑characterized tool compounds manually annotated to 218 unique MoAs, and we profled each compound at four concentrations in live‑cell, high‑content imaging screens against a panel of 15 reporter cell lines, which expressed a diverse set of fuorescent organelle and pathway markers in three distinct cell lineages. For 41 of 83 testable MoAs, phenomic profles accurately ranked the reference compounds (AUC‑ROC ≥ 0.9). MoAs could be better resolved by screening compounds at multiple concentrations than by including replicates at a single concentration. Screening additional cell lineages and fuorescent markers increased the number of distinguishable MoAs but this efect quickly plateaued. There remains a substantial number of MoAs that were hard to distinguish from others under the current study’s conditions. -

Genome-Wide DNA Methylation Analysis of KRAS Mutant Cell Lines Ben Yi Tew1,5, Joel K
www.nature.com/scientificreports OPEN Genome-wide DNA methylation analysis of KRAS mutant cell lines Ben Yi Tew1,5, Joel K. Durand2,5, Kirsten L. Bryant2, Tikvah K. Hayes2, Sen Peng3, Nhan L. Tran4, Gerald C. Gooden1, David N. Buckley1, Channing J. Der2, Albert S. Baldwin2 ✉ & Bodour Salhia1 ✉ Oncogenic RAS mutations are associated with DNA methylation changes that alter gene expression to drive cancer. Recent studies suggest that DNA methylation changes may be stochastic in nature, while other groups propose distinct signaling pathways responsible for aberrant methylation. Better understanding of DNA methylation events associated with oncogenic KRAS expression could enhance therapeutic approaches. Here we analyzed the basal CpG methylation of 11 KRAS-mutant and dependent pancreatic cancer cell lines and observed strikingly similar methylation patterns. KRAS knockdown resulted in unique methylation changes with limited overlap between each cell line. In KRAS-mutant Pa16C pancreatic cancer cells, while KRAS knockdown resulted in over 8,000 diferentially methylated (DM) CpGs, treatment with the ERK1/2-selective inhibitor SCH772984 showed less than 40 DM CpGs, suggesting that ERK is not a broadly active driver of KRAS-associated DNA methylation. KRAS G12V overexpression in an isogenic lung model reveals >50,600 DM CpGs compared to non-transformed controls. In lung and pancreatic cells, gene ontology analyses of DM promoters show an enrichment for genes involved in diferentiation and development. Taken all together, KRAS-mediated DNA methylation are stochastic and independent of canonical downstream efector signaling. These epigenetically altered genes associated with KRAS expression could represent potential therapeutic targets in KRAS-driven cancer. Activating KRAS mutations can be found in nearly 25 percent of all cancers1.