The Flagellar Regulon of Legionella—A Review
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Metaproteogenomic Insights Beyond Bacterial Response to Naphthalene
ORIGINAL ARTICLE ISME Journal – Original article Metaproteogenomic insights beyond bacterial response to 5 naphthalene exposure and bio-stimulation María-Eugenia Guazzaroni, Florian-Alexander Herbst, Iván Lores, Javier Tamames, Ana Isabel Peláez, Nieves López-Cortés, María Alcaide, Mercedes V. del Pozo, José María Vieites, Martin von Bergen, José Luis R. Gallego, Rafael Bargiela, Arantxa López-López, Dietmar H. Pieper, Ramón Rosselló-Móra, Jesús Sánchez, Jana Seifert and Manuel Ferrer 10 Supporting Online Material includes Text (Supporting Materials and Methods) Tables S1 to S9 Figures S1 to S7 1 SUPPORTING TEXT Supporting Materials and Methods Soil characterisation Soil pH was measured in a suspension of soil and water (1:2.5) with a glass electrode, and 5 electrical conductivity was measured in the same extract (diluted 1:5). Primary soil characteristics were determined using standard techniques, such as dichromate oxidation (organic matter content), the Kjeldahl method (nitrogen content), the Olsen method (phosphorus content) and a Bernard calcimeter (carbonate content). The Bouyoucos Densimetry method was used to establish textural data. Exchangeable cations (Ca, Mg, K and 10 Na) extracted with 1 M NH 4Cl and exchangeable aluminium extracted with 1 M KCl were determined using atomic absorption/emission spectrophotometry with an AA200 PerkinElmer analyser. The effective cation exchange capacity (ECEC) was calculated as the sum of the values of the last two measurements (sum of the exchangeable cations and the exchangeable Al). Analyses were performed immediately after sampling. 15 Hydrocarbon analysis Extraction (5 g of sample N and Nbs) was performed with dichloromethane:acetone (1:1) using a Soxtherm extraction apparatus (Gerhardt GmbH & Co. -

NCTC) Bacterial Strain Equivalents to American Type Culture Collection (ATCC) Bacterial Strains
This list shows National Collection of Type Cultures (NCTC) bacterial strain equivalents to American Type Culture Collection (ATCC) bacterial strains. NCTC Number CurrentName ATCC Number NCTC 7212 Acetobacter pasteurianus ATCC 23761 NCTC 10138 Acholeplasma axanthum ATCC 25176 NCTC 10171 Acholeplasma equifetale ATCC 29724 NCTC 10128 Acholeplasma granularum ATCC 19168 NCTC 10172 Acholeplasma hippikon ATCC 29725 NCTC 10116 Acholeplasma laidlawii ATCC 23206 NCTC 10134 Acholeplasma modicum ATCC 29102 NCTC 10188 Acholeplasma morum ATCC 33211 NCTC 10150 Acholeplasma oculi ATCC 27350 NCTC 10198 Acholeplasma parvum ATCC 29892 NCTC 8582 Achromobacter denitrificans ATCC 15173 NCTC 10309 Achromobacter metalcaligenes ATCC 17910 NCTC 10807 Achromobacter xylosoxidans subsp. xylosoxidans ATCC 27061 NCTC 10808 Achromobacter xylosoxidans subsp. xylosoxidans ATCC 17062 NCTC 10809 Achromobacter xylosoxidans subsp. xylosoxidans ATCC 27063 NCTC 12156 Acinetobacter baumannii ATCC 19606 NCTC 10303 Acinetobacter baumannii ATCC 17904 NCTC 7844 Acinetobacter calcoaceticus ATCC 15308 NCTC 12983 Acinetobacter calcoaceticus ATCC 23055 NCTC 8102 acinetobacter dna group 13 ATCC 17903 NCTC 10304 Acinetobacter genospecies 13 ATCC 17905 NCTC 10306 Acinetobacter haemolyticus ATCC 17907 NCTC 10305 Acinetobacter haemolyticus subsp haemolyticus ATCC 17906 NCTC 10308 Acinetobacter johnsonii ATCC 17909 NCTC 10307 Acinetobacter junii ATCC 17908 NCTC 5866 Acinetobacter lwoffii ATCC 15309 NCTC 12870 Actinobacillus delphinicola ATCC 700179 NCTC 8529 Actinobacillus equuli ATCC 19392 -

Legionella Gresilensis Sp. Nov. and Legionella Beliardensis Sp. Nov., Isolated from Water in France
International Journal of Systematic and Evolutionary Microbiology (2001), 51, 1949–1957 Printed in Great Britain Legionella gresilensis sp. nov. and Legionella beliardensis sp. nov., isolated from water in France 1 Centre National de Franc: ois Lo Presti,1‡ Serge Riffard,1 He! le' ne Meugnier,1 Re! fe! rence des Legionella 1 2 3 UPRES EA1655, Faculte! de Monique Reyrolle, Yves Lasne, † Patrick A. D. Grimont, Me! decine RTH Laennec, Francine Grimont,3 Robert F. Benson,4 Don J. Brenner,4 Rue Guillaume Paradin, 4 1 1 69372 Lyon cedex 08, Arnold G. Steigerwalt, Jerome Etienne and Jean Freney France 2 Laboratoire des Author for correspondence: Franc: ois Lo Presti. Tel: j33 0 169 79 79 60. Fax: j33 0 169 79 79 20. Radioisotopes et de e-mail: Francois.Lo-Presti!sanofi-synthelabo.com Biochimie mole! culaire, Ho# pital Edouard Herriot, ! ! 69437 Lyon cedex 03, Novel Legionella-like isolates, strains Montbeliard A1T and Greoux 11 D13T, France isolated from two different French water sources, were studied taxonomically 3 Unite! des Ente! robacte! ries, and phylogenetically. Morphological and biochemical characterization revealed Institut Pasteur, 75724 Paris cedex 15, France that they were Gram-negative, aerobic, non-spore-forming bacilli with a cut- glass appearance that grew only on L-cysteine-supplemented buffered charcoal 4 Respiratory Diseases Branch, Meningitis and yeast extract agar. Phenotypic characterization using fatty acid and ubiquinone Special Pathogens Branch, profiles and SDS-PAGE analysis confirmed that they were closely related, but National Center for distinct from, other species of the genus Legionella, since serotyping could not Infectious Diseases, Centers for Disease Control and relate them to any existing serogroup. -

Evolutionary Origin of Insect–Wolbachia Nutritional Mutualism
Evolutionary origin of insect–Wolbachia nutritional mutualism Naruo Nikoha,1, Takahiro Hosokawab,1, Minoru Moriyamab,1, Kenshiro Oshimac, Masahira Hattoric, and Takema Fukatsub,2 aDepartment of Liberal Arts, The Open University of Japan, Chiba 261-8586, Japan; bBioproduction Research Institute, National Institute of Advanced Industrial Science and Technology, Tsukuba 305-8566, Japan; and cCenter for Omics and Bioinformatics, Graduate School of Frontier Sciences, University of Tokyo, Kashiwa 277-8561, Japan Edited by Nancy A. Moran, University of Texas at Austin, Austin, TX, and approved June 3, 2014 (received for review May 20, 2014) Obligate insect–bacterium nutritional mutualism is among the insects, generally conferring negative fitness consequences to most sophisticated forms of symbiosis, wherein the host and the their hosts and often causing hosts’ reproductive aberrations to symbiont are integrated into a coherent biological entity and un- enhance their own transmission in a selfish manner (7, 8). Re- able to survive without the partnership. Originally, however, such cently, however, a Wolbachia strain associated with the bedbug obligate symbiotic bacteria must have been derived from free-living Cimex lectularius,designatedaswCle, was shown to be es- bacteria. How highly specialized obligate mutualisms have arisen sential for normal growth and reproduction of the blood- from less specialized associations is of interest. Here we address this sucking insect host via provisioning of B vitamins (9). Hence, it –Wolbachia evolutionary -

Legionella Shows a Diverse Secondary Metabolism Dependent on a Broad Spectrum Sfp-Type Phosphopantetheinyl Transferase
Legionella shows a diverse secondary metabolism dependent on a broad spectrum Sfp-type phosphopantetheinyl transferase Nicholas J. Tobias1, Tilman Ahrendt1, Ursula Schell2, Melissa Miltenberger1, Hubert Hilbi2,3 and Helge B. Bode1,4 1 Fachbereich Biowissenschaften, Merck Stiftungsprofessur fu¨r Molekulare Biotechnologie, Goethe Universita¨t, Frankfurt am Main, Germany 2 Max von Pettenkofer Institute, Ludwig-Maximilians-Universita¨tMu¨nchen, Munich, Germany 3 Institute of Medical Microbiology, University of Zu¨rich, Zu¨rich, Switzerland 4 Buchmann Institute for Molecular Life Sciences, Goethe Universita¨t, Frankfurt am Main, Germany ABSTRACT Several members of the genus Legionella cause Legionnaires’ disease, a potentially debilitating form of pneumonia. Studies frequently focus on the abundant number of virulence factors present in this genus. However, what is often overlooked is the role of secondary metabolites from Legionella. Following whole genome sequencing, we assembled and annotated the Legionella parisiensis DSM 19216 genome. Together with 14 other members of the Legionella, we performed comparative genomics and analysed the secondary metabolite potential of each strain. We found that Legionella contains a huge variety of biosynthetic gene clusters (BGCs) that are potentially making a significant number of novel natural products with undefined function. Surprisingly, only a single Sfp-like phosphopantetheinyl transferase is found in all Legionella strains analyzed that might be responsible for the activation of all carrier proteins in primary (fatty acid biosynthesis) and secondary metabolism (polyketide and non-ribosomal peptide synthesis). Using conserved active site motifs, we predict Submitted 29 June 2016 some novel compounds that are probably involved in cell-cell communication, Accepted 25 October 2016 Published 24 November 2016 differing to known communication systems. -

Which Organisms Are Used for Anti-Biofouling Studies
Table S1. Semi-systematic review raw data answering: Which organisms are used for anti-biofouling studies? Antifoulant Method Organism(s) Model Bacteria Type of Biofilm Source (Y if mentioned) Detection Method composite membranes E. coli ATCC25922 Y LIVE/DEAD baclight [1] stain S. aureus ATCC255923 composite membranes E. coli ATCC25922 Y colony counting [2] S. aureus RSKK 1009 graphene oxide Saccharomycetes colony counting [3] methyl p-hydroxybenzoate L. monocytogenes [4] potassium sorbate P. putida Y. enterocolitica A. hydrophila composite membranes E. coli Y FESEM [5] (unspecified/unique sample type) S. aureus (unspecified/unique sample type) K. pneumonia ATCC13883 P. aeruginosa BAA-1744 composite membranes E. coli Y SEM [6] (unspecified/unique sample type) S. aureus (unspecified/unique sample type) graphene oxide E. coli ATCC25922 Y colony counting [7] S. aureus ATCC9144 P. aeruginosa ATCCPAO1 composite membranes E. coli Y measuring flux [8] (unspecified/unique sample type) graphene oxide E. coli Y colony counting [9] (unspecified/unique SEM sample type) LIVE/DEAD baclight S. aureus stain (unspecified/unique sample type) modified membrane P. aeruginosa P60 Y DAPI [10] Bacillus sp. G-84 LIVE/DEAD baclight stain bacteriophages E. coli (K12) Y measuring flux [11] ATCC11303-B4 quorum quenching P. aeruginosa KCTC LIVE/DEAD baclight [12] 2513 stain modified membrane E. coli colony counting [13] (unspecified/unique colony counting sample type) measuring flux S. aureus (unspecified/unique sample type) modified membrane E. coli BW26437 Y measuring flux [14] graphene oxide Klebsiella colony counting [15] (unspecified/unique sample type) P. aeruginosa (unspecified/unique sample type) graphene oxide P. aeruginosa measuring flux [16] (unspecified/unique sample type) composite membranes E. -

Microbial Diversity Under Extreme Euxinia: Mahoney Lake, Canada V
Geobiology (2012), 10, 223–235 DOI: 10.1111/j.1472-4669.2012.00317.x Microbial diversity under extreme euxinia: Mahoney Lake, Canada V. KLEPAC-CERAJ,1,2 C. A. HAYES,3 W. P. GILHOOLY,4 T. W. LYONS,5 R. KOLTER2 AND A. PEARSON3 1Department of Molecular Genetics, Forsyth Institute, Cambridge, MA, USA 2Department of Microbiology and Molecular Genetics, Harvard Medical School, Boston, MA, USA 3Department of Earth and Planetary Sciences, Harvard University, Cambridge, MA, USA 4Department of Earth and Planetary Sciences, Washington University, Saint Louis, MO, USA 5Department of Earth Sciences, University of California, Riverside, CA, USA ABSTRACT Mahoney Lake, British Columbia, Canada, is a stratified, 15-m deep saline lake with a euxinic (anoxic, sulfidic) hypolimnion. A dense plate of phototrophic purple sulfur bacteria is found at the chemocline, but to date the rest of the Mahoney Lake microbial ecosystem has been underexamined. In particular, the microbial community that resides in the aphotic hypolimnion and ⁄ or in the lake sediments is unknown, and it is unclear whether the sulfate reducers that supply sulfide for phototrophy live only within, or also below, the plate. Here we profiled distribu- tions of 16S rRNA genes using gene clone libraries and PhyloChip microarrays. Both approaches suggest that microbial diversity is greatest in the hypolimnion (8 m) and sediments. Diversity is lowest in the photosynthetic plate (7 m). Shallower depths (5 m, 7 m) are rich in Actinobacteria, Alphaproteobacteria, and Gammaproteo- bacteria, while deeper depths (8 m, sediments) are rich in Crenarchaeota, Natronoanaerobium, and Verrucomi- crobia. The heterogeneous distribution of Deltaproteobacteria and Epsilonproteobacteria between 7 and 8 m is consistent with metabolisms involving sulfur intermediates in the chemocline, but complete sulfate reduction in the hypolimnion. -

The Risk to Human Health from Free-Living Amoebae Interaction with Legionella in Drinking and Recycled Water Systems
THE RISK TO HUMAN HEALTH FROM FREE-LIVING AMOEBAE INTERACTION WITH LEGIONELLA IN DRINKING AND RECYCLED WATER SYSTEMS Dissertation submitted by JACQUELINE MARIE THOMAS BACHELOR OF SCIENCE (HONOURS) AND BACHELOR OF ARTS, UNSW In partial fulfillment of the requirements for the award of DOCTOR OF PHILOSOPHY in ENVIRONMENTAL ENGINEERING SCHOOL OF CIVIL AND ENVIRONMENTAL ENGINEERING FACULTY OF ENGINEERING MAY 2012 SUPERVISORS Professor Nicholas Ashbolt Office of Research and Development United States Environmental Protection Agency Cincinnati, Ohio USA and School of Civil and Environmental Engineering Faculty of Engineering The University of New South Wales Sydney, Australia Professor Richard Stuetz School of Civil and Environmental Engineering Faculty of Engineering The University of New South Wales Sydney, Australia Doctor Torsten Thomas School of Biotechnology and Biomolecular Sciences Faculty of Science The University of New South Wales Sydney, Australia ORIGINALITY STATEMENT '1 hereby declare that this submission is my own work and to the best of my knowledge it contains no materials previously published or written by another person, or substantial proportions of material which have been accepted for the award of any other degree or diploma at UNSW or any other educational institution, except where due acknowledgement is made in the thesis. Any contribution made to the research by others, with whom 1 have worked at UNSW or elsewhere, is explicitly acknowledged in the thesis. I also declare that the intellectual content of this thesis is the product of my own work, except to the extent that assistance from others in the project's design and conception or in style, presentation and linguistic expression is acknowledged.' Signed ~ ............................ -

18167638.Pdf
REVIEW ARTICLE Amoebal pathogens as emerging causal agents of pneumonia Fred´ eric´ Lamoth1 & Gilbert Greub1,2 1Infectious Diseases Service, University of Lausanne, Lausanne, Switzerland; and 2Center for Research on Intracellular Bacteria, Institute of Microbiology, University Hospital Center and University of Lausanne, Lausanne, Switzerland Correspondence: Gilbert Greub, Center for Abstract Research on Intracellular Bacteria, Institute of Microbiology, University Hospital Center and Despite using modern microbiological diagnostic approaches, the aetiological University of Lausanne, Rue du Bugnon 46, agents of pneumonia remain unidentified in about 50% of cases. Some bacteria 1011 Lausanne, Switzerland. Tel.: 141 21 that grow poorly or not at all in axenic media used in routine clinical bacteriology 31449 79; fax: 141 21 31440 60; e-mail: laboratory but which can develop inside amoebae may be the agents of these lower [email protected] respiratory tract infections (RTIs) of unexplained aetiology. Such amoebae- resisting bacteria, which coevolved with amoebae to resist their microbicidal Received 24 September 2009; revised 30 machinery, may have developed virulence traits that help them survive within November 2009; accepted 2 December 2009. human macrophages, i.e. the first line of innate immune defence in the lung. We Final version published online 22 January 2010. review here the current evidence for the emerging pathogenic role of various DOI:10.1111/j.1574-6976.2009.00207.x amoebae-resisting microorganisms as agents of RTIs in humans. Specifically, we discuss the emerging pathogenic roles of Legionella-like amoebal pathogens, novel Editor: Colin Berry Chlamydiae (Parachlamydia acanthamoebae, Simkania negevensis), waterborne mycobacteria and Bradyrhizobiaceae (Bosea and Afipia spp.). Keywords free-living amoebae; amoebae-resisting bacteria; Legionella; Chlamydia-like bacteria; mycobacteria; pneumonia. -

Subcellular Location of Piscirickettsia Salmonis Heat Shock Protein 60 (Hsp60) Chaperone by Using Immunogold Labeling and Proteomic Analysis
microorganisms Article Subcellular Location of Piscirickettsia salmonis Heat Shock Protein 60 (Hsp60) Chaperone by Using Immunogold Labeling and Proteomic Analysis 1, 2,3, 4 5 Cristian Oliver y, Patricio Sánchez y , Karla Valenzuela , Mauricio Hernández , Juan Pablo Pontigo 3, Maria C. Rauch 3, Rafael A. Garduño 4,6 , Ruben Avendaño-Herrera 2,7,* and Alejandro J. Yáñez 2,8,* 1 Laboratorio de Inmunología y Estrés de Organismos Acuáticos, Instituto de Patología Animal, Facultad de Ciencias Veterinarias, Universidad Austral de Chile, Valdivia 5090000, Chile; [email protected] 2 Interdisciplinary Center for Aquaculture Research, (INCAR), Concepción 4070386, Chile; [email protected] 3 Instituto de Bioquímica y Microbiología, Facultad de Ciencias, Universidad Austral de Chile, Valdivia 5090000, Chile; [email protected] (J.P.P.); [email protected] (M.C.R.) 4 Microbiology and Immunology Department, Dalhousie University, Halifax, NS B3H 4R2, Canada; [email protected] (K.V.); [email protected] (R.A.G.) 5 Austral-OMICS, Faculty of Sciences, Universidad Austral de Chile, Valdivia 5090000, Chile; [email protected] 6 Canadian Food Inspection Agency, Dartmouth Laboratory, Dartmouth, NS B3B 1Y9, Canada 7 Universidad Andrés Bello, Laboratorio de Patología de Organismos Acuáticos y Biotecnología Acuícola, Facultad Ciencias de la Vida, Viña del Mar 2531015, Chile 8 Facultad de Ciencias, Universidad Austral de Chile, Valdivia 5090000, Chile * Correspondence: [email protected] (R.A.-H.); [email protected] (A.J.Y.) These authors contributed equally to this work. y Received: 12 November 2019; Accepted: 31 December 2019; Published: 15 January 2020 Abstract: Piscirickettsia salmonis is the causative bacterial agent of piscirickettsiosis, a systemic fish disease that significantly impacts the Chilean salmon industry. -
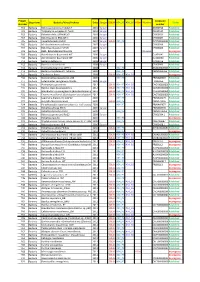
Project Number Organisms Bacteria/Virus/Archaea Date
Project_ Accession Organisms Bacteria/Virus/Archaea Date Sanger SOLiD 454_PE 454_SG PGM Illumina Status Number number P01 Bacteria Rickettsia conorii str.Malish 7 2001 Sanger AE006914 Published P02 Bacteria Tropheryma whipplei str.Twist 2003 Sanger AE014184 Published P03 Bacteria Rickettsia felis URRWXCal2 2005 Sanger CP000053 Published P04 Bacteria Rickettsia bellii RML369-C 2006 Sanger CP000087 Published P05 Bacteria Coxiella burnetii CB109 2007 Sanger SOLiD 454_PE AKYP00000000 Published P06 Bacteria Minibacterium massiliensis 2007 Sanger CP000269 Published P07 Bacteria Rickettsia massiliae MTU5 2007 Sanger CP000683 Published P08 Bacteria BaBL=Bête à Bernard Lascola 2007 Illumina In progress P09 Bacteria Acinetobacter baumannii AYE 2006 Sanger CU459141 Published P10 Bacteria Acinetobacter baumannii SDF 2006 Sanger CU468230 Published P11 Bacteria Borrelia duttonii Ly 2008 Sanger CP000976 Published P12 Bacteria Borrelia recurrentis A1 2008 Sanger CP000993 Published P13 Bacteria Francisella tularensis URFT1 2008 454_PE ABAZ00000000Published P14 Bacteria Borrelia crocidurae str. Achema 2009 454_PE PRJNA162335 Published P15 Bacteria Citrobacter koseri 2009 SOLiD 454_PE 454_SG In progress P16 Bacteria Diplorickettsia massiliensis 20B 2009 454_PE PRJNA86907 Published P17 Bacteria Enterobacter aerogenes EA1509E 2009 Sanger FO203355 Published P18 Bacteria Actinomyces grossensis 2012 SOLiD 454_PE 454_SG CAGY00000000Published P19 Bacteria Bacillus massiliosenegalensis 2012 SOLiD 454_PE 454_SG CAHJ00000000 Published P20 Bacteria Brevibacterium senegalensis -
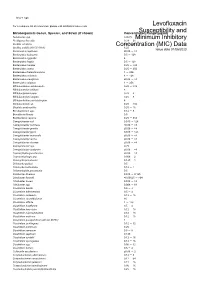
Susceptibility and Resistance Data
toku-e logo For a complete list of references, please visit antibiotics.toku-e.com Levofloxacin Microorganism Genus, Species, and Strain (if shown) Concentration Range (μg/ml)Susceptibility and Aeromonas spp. 0.0625 Minimum Inhibitory Alcaligenes faecalis 0.39 - 25 Bacillus circulans Concentration0.25 - 8 (MIC) Data Bacillus subtilis (ATCC 6051) 6.25 Issue date 01/06/2020 Bacteroides capillosus ≤0.06 - >8 Bacteroides distasonis 0.5 - 128 Bacteroides eggerthii 4 Bacteroides fragilis 0.5 - 128 Bacteroides merdae 0.25 - >32 Bacteroides ovatus 0.25 - 256 Bacteroides thetaiotaomicron 1 - 256 Bacteroides uniformis 4 - 128 Bacteroides ureolyticus ≤0.06 - >8 Bacteroides vulgatus 1 - 256 Bifidobacterium adolescentis 0.25 - >32 Bifidobacterium bifidum 8 Bifidobacterium breve 0.25 - 8 Bifidobacterium longum 0.25 - 8 Bifidobacterium pseudolongum 8 Bifidobacterium sp. 0.25 - >32 Bilophila wadsworthia 0.25 - 16 Brevibacterium spp. 0.12 - 8 Brucella melitensis 0.5 Burkholderia cepacia 0.25 - 512 Campylobacter coli 0.015 - 128 Campylobacter concisus ≤0.06 - >8 Campylobacter gracilis ≤0.06 - >8 Campylobacter jejuni 0.015 - 128 Campylobacter mucosalis ≤0.06 - >8 Campylobacter rectus ≤0.06 - >8 Campylobacter showae ≤0.06 - >8 Campylobacter spp. 0.25 Campylobacter sputorum ≤0.06 - >8 Capnocytophaga ochracea ≤0.06 - >8 Capnocytophaga spp. 0.006 - 2 Chlamydia pneumonia 0.125 - 1 Chlamydia psittaci 0.5 Chlamydia trachomatis 0.12 - 1 Chlamydophila pneumonia 0.5 Citrobacter diversus 0.015 - 0.125 Citrobacter freundii ≤0.00625 - >64 Citrobacter koseri 0.015 -