Unexpected and Variable Phenotypes in a Family with JAK3 Deficiency
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Promoter Janus Kinase 3 Proximal Characterization and Analysis Of
The Journal of Immunology Characterization and Analysis of the Proximal Janus Kinase 3 Promoter1 Martin Aringer,2*† Sigrun R. Hofmann,2* David M. Frucht,* Min Chen,* Michael Centola,* Akio Morinobu,* Roberta Visconti,* Daniel L. Kastner,* Josef S. Smolen,† and John J. O’Shea3* Janus kinase 3 (Jak3) is a nonreceptor tyrosine kinase essential for signaling via cytokine receptors that comprise the common ␥-chain (␥c), i.e., the receptors for IL-2, IL-4, IL-7, IL-9, IL-15, and IL-21. Jak3 is preferentially expressed in hemopoietic cells and is up-regulated upon cell differentiation and activation. Despite the importance of Jak3 in lymphoid development and immune function, the mechanisms that govern its expression have not been defined. To gain insight into this issue, we set out to characterize the Jak3 promoter. The 5-untranslated region of the Jak3 gene is interrupted by a 3515-bp intron. Upstream of this intron and the transcription initiation site, we identified an ϳ1-kb segment that exhibited lymphoid-specific promoter activity and was responsive to TCR signals. Truncation of this fragment revealed that core promoter activity resided in a 267-bp fragment that contains putative Sp-1, AP-1, Ets, Stat, and other binding sites. Mutation of the AP-1 sites significantly diminished, whereas mutation of the Ets sites abolished, the inducibility of the promoter construct. Chromatin immunoprecipitation assays showed that histone acetylation correlates with mRNA expression and that Ets-1/2 binds this region. Thus, transcription factors that bind these sites, especially Ets family members, are likely to be important regulators of Jak3 expression. -
HCC and Cancer Mutated Genes Summarized in the Literature Gene Symbol Gene Name References*
HCC and cancer mutated genes summarized in the literature Gene symbol Gene name References* A2M Alpha-2-macroglobulin (4) ABL1 c-abl oncogene 1, receptor tyrosine kinase (4,5,22) ACBD7 Acyl-Coenzyme A binding domain containing 7 (23) ACTL6A Actin-like 6A (4,5) ACTL6B Actin-like 6B (4) ACVR1B Activin A receptor, type IB (21,22) ACVR2A Activin A receptor, type IIA (4,21) ADAM10 ADAM metallopeptidase domain 10 (5) ADAMTS9 ADAM metallopeptidase with thrombospondin type 1 motif, 9 (4) ADCY2 Adenylate cyclase 2 (brain) (26) AJUBA Ajuba LIM protein (21) AKAP9 A kinase (PRKA) anchor protein (yotiao) 9 (4) Akt AKT serine/threonine kinase (28) AKT1 v-akt murine thymoma viral oncogene homolog 1 (5,21,22) AKT2 v-akt murine thymoma viral oncogene homolog 2 (4) ALB Albumin (4) ALK Anaplastic lymphoma receptor tyrosine kinase (22) AMPH Amphiphysin (24) ANK3 Ankyrin 3, node of Ranvier (ankyrin G) (4) ANKRD12 Ankyrin repeat domain 12 (4) ANO1 Anoctamin 1, calcium activated chloride channel (4) APC Adenomatous polyposis coli (4,5,21,22,25,28) APOB Apolipoprotein B [including Ag(x) antigen] (4) AR Androgen receptor (5,21-23) ARAP1 ArfGAP with RhoGAP domain, ankyrin repeat and PH domain 1 (4) ARHGAP35 Rho GTPase activating protein 35 (21) ARID1A AT rich interactive domain 1A (SWI-like) (4,5,21,22,24,25,27,28) ARID1B AT rich interactive domain 1B (SWI1-like) (4,5,22) ARID2 AT rich interactive domain 2 (ARID, RFX-like) (4,5,22,24,25,27,28) ARID4A AT rich interactive domain 4A (RBP1-like) (28) ARID5B AT rich interactive domain 5B (MRF1-like) (21) ASPM Asp (abnormal -

Supplementary Table 1. in Vitro Side Effect Profiling Study for LDN/OSU-0212320. Neurotransmitter Related Steroids
Supplementary Table 1. In vitro side effect profiling study for LDN/OSU-0212320. Percent Inhibition Receptor 10 µM Neurotransmitter Related Adenosine, Non-selective 7.29% Adrenergic, Alpha 1, Non-selective 24.98% Adrenergic, Alpha 2, Non-selective 27.18% Adrenergic, Beta, Non-selective -20.94% Dopamine Transporter 8.69% Dopamine, D1 (h) 8.48% Dopamine, D2s (h) 4.06% GABA A, Agonist Site -16.15% GABA A, BDZ, alpha 1 site 12.73% GABA-B 13.60% Glutamate, AMPA Site (Ionotropic) 12.06% Glutamate, Kainate Site (Ionotropic) -1.03% Glutamate, NMDA Agonist Site (Ionotropic) 0.12% Glutamate, NMDA, Glycine (Stry-insens Site) 9.84% (Ionotropic) Glycine, Strychnine-sensitive 0.99% Histamine, H1 -5.54% Histamine, H2 16.54% Histamine, H3 4.80% Melatonin, Non-selective -5.54% Muscarinic, M1 (hr) -1.88% Muscarinic, M2 (h) 0.82% Muscarinic, Non-selective, Central 29.04% Muscarinic, Non-selective, Peripheral 0.29% Nicotinic, Neuronal (-BnTx insensitive) 7.85% Norepinephrine Transporter 2.87% Opioid, Non-selective -0.09% Opioid, Orphanin, ORL1 (h) 11.55% Serotonin Transporter -3.02% Serotonin, Non-selective 26.33% Sigma, Non-Selective 10.19% Steroids Estrogen 11.16% 1 Percent Inhibition Receptor 10 µM Testosterone (cytosolic) (h) 12.50% Ion Channels Calcium Channel, Type L (Dihydropyridine Site) 43.18% Calcium Channel, Type N 4.15% Potassium Channel, ATP-Sensitive -4.05% Potassium Channel, Ca2+ Act., VI 17.80% Potassium Channel, I(Kr) (hERG) (h) -6.44% Sodium, Site 2 -0.39% Second Messengers Nitric Oxide, NOS (Neuronal-Binding) -17.09% Prostaglandins Leukotriene, -

Crystal Structure of the Jak3 Kinase Domain in Complex with a Staurosporine Analog
From www.bloodjournal.org by guest on September 11, 2016. For personal use only. TRANSPLANTATION Crystal structure of the Jak3 kinase domain in complex with a staurosporine analog Titus J. Boggon, Yiqun Li, Paul W. Manley, and Michael J. Eck Jak (Janus kinase) family nonreceptor target for development of lymphocyte- tion of the activation loop. Such a direct tyrosine kinases are central mediators of specific immunosuppressants. Here, we coupling has not been previously ob- cytokine signaling. The Jak kinases ex- present the crystal structure of the Jak3 served in tyrosine kinases and may be hibit distinct cytokine receptor associa- kinase domain in complex with stauro- unique to Jak kinases. The crystal struc- tion profiles and so transduce different sporine analog AFN941. The kinase do- ture provides a detailed view of the Jak3 signals. Jak3 expression is limited to the main is in the active conformation, with active site and will facilitate computa- immune system, where it plays a key role both activation loop tyrosine residues tional and structure-directed approaches in signal transduction from cytokine re- phosphorylated. The phosphate group on to development of Jak3-specific inhibi- ceptors containing the common gamma- pTyr981 in the activation loop is in part tors. (Blood. 2005;106:996-1002) chain, ␥c. Patients unable to signal via ␥c coordinated by an arginine residue in the present with severe combined immunode- regulatory C-helix, suggesting a direct ficiency (SCID). The finding that Jak3 mechanism by which the active position mutations result in SCID has made it a of the C-helix is induced by phosphoryla- © 2005 by The American Society of Hematology Introduction The Janus kinase (Jak) family of cytoplasmic tyrosine kinases are regulation of diverse cell processes, including proliferation, differ- essential for signal transduction from a wide variety of cell-surface entiation, migration, and apoptosis.1,2 receptors. -

JAK3 Deficiency, (SCID T-B+)
JAK3 deficiency, (SCID T-B+) Author: Professor Luigi D. Notarangelo1,2 Creation Date: November 2001 Update: January 2005 1member of the European editorial committee of Orphanet encyclopedia 2Department of Pediatrics, University of Brescia, Spedali Civil, 25123 Brescia, Italy. [email protected] Abstract Keywords Diagnosis criteria/definition Differential diagnosis Prevalence Clinical description Treatment Etiology Diagnostic methods Genetic counseling Antenatal diagnosis Unresolved questions References Abstract JAK3 (Janus Kinase 3) deficiency is an autosomal recessive form of severe combined immune deficiency (SCID). It is characterized by lack of circulating T and NK (Natural Killer) cells and normal number of B lymphocytes. The disease is due to mutations in the JAK3 gene encoding an intracellular tyrosine kinase that is physically and functionally coupled with several cytokine receptors. Identification of gene anomalies has allowed physicians to make the diagnosis (even prenatal), and may prompt novel forms of treatment based on gene therapy. Although a relatively low number of JAK3-deficient subjects have been diagnosed, JAK3 deficiency represents an important cause of autosomal recessive SCID in the United States and its prevalence in Europe appears to be even higher. However it is considered as a rare disease (incidence is between 1/100,000 and 1/1,000,000 live births). JAK3-deficient patients present with the classical clinical features of SCID in the first few months of life, i.e. chronic diarrhea, failure to thrive, recurrent respiratory infection and/or generalized infections from opportunistic pathogens, or signs of graft-versus-host reaction (skin rash, abnormalities of liver function, pancytopenia) from transplacental acquired maternal T cells. -

Supplementary Table 2
Supplementary Table 2. Differentially Expressed Genes following Sham treatment relative to Untreated Controls Fold Change Accession Name Symbol 3 h 12 h NM_013121 CD28 antigen Cd28 12.82 BG665360 FMS-like tyrosine kinase 1 Flt1 9.63 NM_012701 Adrenergic receptor, beta 1 Adrb1 8.24 0.46 U20796 Nuclear receptor subfamily 1, group D, member 2 Nr1d2 7.22 NM_017116 Calpain 2 Capn2 6.41 BE097282 Guanine nucleotide binding protein, alpha 12 Gna12 6.21 NM_053328 Basic helix-loop-helix domain containing, class B2 Bhlhb2 5.79 NM_053831 Guanylate cyclase 2f Gucy2f 5.71 AW251703 Tumor necrosis factor receptor superfamily, member 12a Tnfrsf12a 5.57 NM_021691 Twist homolog 2 (Drosophila) Twist2 5.42 NM_133550 Fc receptor, IgE, low affinity II, alpha polypeptide Fcer2a 4.93 NM_031120 Signal sequence receptor, gamma Ssr3 4.84 NM_053544 Secreted frizzled-related protein 4 Sfrp4 4.73 NM_053910 Pleckstrin homology, Sec7 and coiled/coil domains 1 Pscd1 4.69 BE113233 Suppressor of cytokine signaling 2 Socs2 4.68 NM_053949 Potassium voltage-gated channel, subfamily H (eag- Kcnh2 4.60 related), member 2 NM_017305 Glutamate cysteine ligase, modifier subunit Gclm 4.59 NM_017309 Protein phospatase 3, regulatory subunit B, alpha Ppp3r1 4.54 isoform,type 1 NM_012765 5-hydroxytryptamine (serotonin) receptor 2C Htr2c 4.46 NM_017218 V-erb-b2 erythroblastic leukemia viral oncogene homolog Erbb3 4.42 3 (avian) AW918369 Zinc finger protein 191 Zfp191 4.38 NM_031034 Guanine nucleotide binding protein, alpha 12 Gna12 4.38 NM_017020 Interleukin 6 receptor Il6r 4.37 AJ002942 -
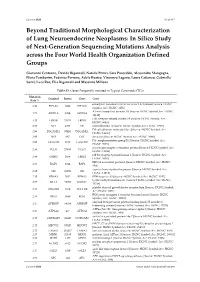
Beyond Traditional Morphological Characterization of Lung
Cancers 2020 S1 of S15 Beyond Traditional Morphological Characterization of Lung Neuroendocrine Neoplasms: In Silico Study of Next-Generation Sequencing Mutations Analysis across the Four World Health Organization Defined Groups Giovanni Centonze, Davide Biganzoli, Natalie Prinzi, Sara Pusceddu, Alessandro Mangogna, Elena Tamborini, Federica Perrone, Adele Busico, Vincenzo Lagano, Laura Cattaneo, Gabriella Sozzi, Luca Roz, Elia Biganzoli and Massimo Milione Table S1. Genes Frequently mutated in Typical Carcinoids (TCs). Mutation Original Entrez Gene Gene Rate % eukaryotic translation initiation factor 1A X-linked [Source: HGNC 4.84 EIF1AX 1964 EIF1AX Symbol; Acc: HGNC: 3250] AT-rich interaction domain 1A [Source: HGNC Symbol;Acc: HGNC: 4.71 ARID1A 8289 ARID1A 11110] LDL receptor related protein 1B [Source: HGNC Symbol; Acc: 4.35 LRP1B 53353 LRP1B HGNC: 6693] 3.53 NF1 4763 NF1 neurofibromin 1 [Source: HGNC Symbol;Acc: HGNC: 7765] DS cell adhesion molecule like 1 [Source: HGNC Symbol; Acc: 2.90 DSCAML1 57453 DSCAML1 HGNC: 14656] 2.90 DST 667 DST dystonin [Source: HGNC Symbol;Acc: HGNC: 1090] FA complementation group D2 [Source: HGNC Symbol; Acc: 2.90 FANCD2 2177 FANCD2 HGNC: 3585] piccolo presynaptic cytomatrix protein [Source: HGNC Symbol; Acc: 2.90 PCLO 27445 PCLO HGNC: 13406] erb-b2 receptor tyrosine kinase 2 [Source: HGNC Symbol; Acc: 2.44 ERBB2 2064 ERBB2 HGNC: 3430] BRCA1 associated protein 1 [Source: HGNC Symbol; Acc: HGNC: 2.35 BAP1 8314 BAP1 950] capicua transcriptional repressor [Source: HGNC Symbol; Acc: 2.35 CIC 23152 CIC HGNC: -

Advances in the Inhibitors of Janus Kinase
l ch cina em di is e tr M y Jiang et al., Med chem 2014, 4:8 Medicinal chemistry DOI: 10.4172/2161-0444.1000192 ISSN: 2161-0444 Review Article Open Access Advances in the Inhibitors of Janus Kinase Jun-Jie J Jiang1, Xiao-Ying Wang1, Yv Zhang2, Yi Jin1* and Jun Lin1* 1Key Laboratory of Medicinal Chemistry for Natural Resource (Yunnan University), Ministry Education, School of Chemical Science and Technology, Yunnan University, Kunming, 650091, P. R. China 2State Key Laboratory of Drug Research, Shanghai Institute of Materia Medica, Shanghai Institutes for Biological Sciences, Chinese Academy of Sciences, 201203, China Abstract The janus kinases (JAKs) comprise an important class of non-receptor protein tyrosine kinases. Cytokine and receptor binding can cause the activation of JAKs, and then activate the "signal transcripts and transcriptional activator”, making JAK enter the nucleus to induce target gene expression. JAKs regulate inflammatory diseases, bone marrow hyperplasia and a variety of malignant tumours. Using JAKs inhibitors as the therapy for organ transplantation and immune disease has a very important significance. In recent years, many of the JAKs inhibitors have been developed one after another. Pan JAKs inhibitor and selective JAK2 or JAK3 inhibitor research progress are reviewed in this article. Keywords: Janus kinase; Inhibitors; Anti-tumour; Anti- activator of transcription (STAT) proteins by phosphorylation sites inflammatory near the SH2 domain [6]. Introduction JAK1 signal conduction mainly relies on a variety of cytokines, making them phosphorylated and increasing their ability to activate In recent years, for the lesion site, studies at the molecular level of downstream signalling molecules. -

Src Directly Tyrosine-Phosphorylates STAT5 on Its Activation Site and Is Involved in Erythropoietin-Induced Signaling Pathway
Oncogene (2001) 20, 6643 ± 6650 ã 2001 Nature Publishing Group All rights reserved 0950 ± 9232/01 $15.00 www.nature.com/onc Src directly tyrosine-phosphorylates STAT5 on its activation site and is involved in erythropoietin-induced signaling pathway Yuichi Okutani1, Akira Kitanaka1, Terukazu Tanaka*,2, Hiroshi Kamano6, Hiroaki Ohnishi3, Yoshitsugu Kubota4, Toshihiko Ishida1 and Jiro Takahara5 1First Department of Internal Medicine, Kagawa Medical University, Kagawa 761-0793, Japan; 2Environmental Health Sciences, Kagawa Medical University, Kagawa 761-0793, Japan; 3Laboratory Medicine, Kagawa Medical University, Kagawa 761-0793, Japan; 4Transfusion Medicine, Kagawa Medical University, Kagawa 761-0793, Japan; 5Kagawa Medical University, Kagawa 761- 0793, Japan; 6Health Science Center, Kagawa University, Kagawa 760-8521, Japan Signal transducers and activators of transcription Growth and dierentiation of hematopoietic cells are (STAT) proteins are transcription factors activated by regulated by the interaction of cell surface receptors phosphorylation on tyrosine residues after cytokine with their speci®c cytokines or growth factors. While stimulation. In erythropoietin receptor (EPOR)-mediated some of these receptors transmit signals via their signaling, STAT5 is tyrosine-phosphorylated by EPO intrinsic protein tyrosine kinase (PTK) activity, others stimulation. Although Janus Kinase 2 (JAK2) is reported use cytoplasmic non-receptor PTK for mediating to play a crucial role in EPO-induced activation of signals (van der Geer et al., 1994). The binding of STAT5, it is unclear whether JAK2 alone can tyrosine- erythropoietin (EPO) to an erythropoietin receptor phosphorylate STAT5 after EPO stimulation. Several (EPOR) which does not contain any intrinsic PTK studies indicate that STAT activation is caused by activity induces rapid and transient tyrosine phosphor- members of other families of protein tyrosine kinases ylation of intracellular proteins (Constantinescu et al., such as the Src family. -

Interpretation of Cytokine Signaling Through the Transcription Factors STAT5A and STAT5B
Downloaded from genesdev.cshlp.org on September 25, 2021 - Published by Cold Spring Harbor Laboratory Press REVIEW Interpretation of cytokine signaling through the transcription factors STAT5A and STAT5B Lothar Hennighausen1 and Gertraud W. Robinson Laboratory of Genetics and Physiology, National Institute of Diabetes and Digestive and Kidney Diseases, National Institutes of Health, Bethesda, Maryland 20892, USA Transcription factors from the family of Signal Trans- the “wrong” STATs and thus acquire inappropriate cues. ducers and Activators of Transcription (STAT) are acti- We propose that mice with mutations in various com- vated by numerous cytokines. Two members of this fam- ponents of the JAK–STAT signaling pathway are living ily, STAT5A and STAT5B (collectively called STAT5), laboratories, which will provide insight into the versa- have gained prominence in that they are activated by a tility of signaling hardware and the adaptability of the wide variety of cytokines such as interleukins, erythro- software. poietin, growth hormone, and prolactin. Furthermore, constitutive STAT5 activation is observed in the major- ity of leukemias and many solid tumors. Inactivation Historical perspective studies in mice as well as human mutations have pro- In 1994, Bernd Groner and colleagues (Wakao et al. vided insight into many of STAT5’s functions. Disrup- 1994), then at the Friedrich Miescher Institute in Basel, tion of cytokine signaling through STAT5 results in a cloned a cDNA from lactating ovine mammary tissue variety of cell-specific effects, ranging from a defective that encoded a transcription factor promoting prolactin- immune system and impaired erythropoiesis, the com- induced transcription of milk protein genes in mammary plete absence of mammary development during preg- epithelium. -

Direct Targets of Pstat5 Signalling in Erythropoiesis
RESEARCH ARTICLE Direct targets of pSTAT5 signalling in erythropoiesis Kevin R. Gillinder1, Hugh Tuckey1,2, Charles C. Bell1, Graham W. Magor1, Stephen Huang1,2, Melissa D. Ilsley1,2, Andrew C. Perkins1,2,3* 1 Cancer Genomics Group, Mater Research Institute - University of Queensland, Translational Research Institute, Woolloongabba, Brisbane, Queensland, Australia, 2 Faculty of Medicine and Biomedical Sciences, University of Queensland, St. Lucia, Brisbane, Queensland, Australia, 3 Princess Alexandra Hospital, Brisbane, Queensland, Australia a1111111111 * [email protected] a1111111111 a1111111111 a1111111111 Abstract a1111111111 Erythropoietin (EPO) acts through the dimeric erythropoietin receptor to stimulate prolifera- tion, survival, differentiation and enucleation of erythroid progenitor cells. We undertook two complimentary approaches to find EPO-dependent pSTAT5 target genes in murine ery- throid cells: RNA-seq of newly transcribed (4sU-labelled) RNA, and ChIP-seq for pSTAT5 OPEN ACCESS 30 minutes after EPO stimulation. We found 302 pSTAT5-occupied sites: ~15% of these Citation: Gillinder KR, Tuckey H, Bell CC, Magor GW, Huang S, Ilsley MD, et al. (2017) Direct reside in promoters while the rest reside within intronic enhancers or intergenic regions, targets of pSTAT5 signalling in erythropoiesis. some >100kb from the nearest TSS. The majority of pSTAT5 peaks contain a central palin- PLoS ONE 12(7): e0180922. https://doi.org/ dromic GAS element, TTCYXRGAA. There was significant enrichment for GATA motifs and 10.1371/journal.pone.0180922 CACCC-box motifs within the neighbourhood of pSTAT5-bound peaks, and GATA1 and/or Editor: Kevin D Bunting, Emory University, UNITED KLF1 co-occupancy at many sites. Using 4sU-RNA-seq we determined the EPO-induced STATES transcriptome and validated differentially expressed genes using dynamic CAGE data and Received: May 19, 2017 qRT-PCR. -

Hirbe AC, Kaushal M, Sharma MK
Original Article Clinical Genomic Profiling Identifies TYK2 Mutation and Overexpression in Patients With Neurofibromatosis Type 1-Associated Malignant Peripheral Nerve Sheath Tumors Angela C. Hirbe, MD, PhD1; Madhurima Kaushal, MS2; Mukesh Kumar Sharma, PhD2; Sonika Dahiya, MD2; Melike Pekmezci, MD3; Arie Perry, MD3,4; and David H. Gutmann, MD, PhD5 BACKGROUND: Malignant peripheral nerve sheath tumors (MPNSTs) are aggressive sarcomas that arise at an estimated frequency of 8% to 13% in individuals with neurofibromatosis type 1 (NF1). Compared with their sporadic counterparts, NF1-associated MPNSTs (NF1-MPNSTs) develop in young adults, frequently recur (approximately 50% of cases), and carry a dismal prognosis. As such, most individuals affected with NF1-MPNSTs die within 5 years of diagnosis, despite surgical resection combined with radiotherapy and che- motherapy. METHODS: Clinical genomic profiling was performed using 1000 ng of DNA from 7 cases of NF1-MPNST, and bioinformatic analyses were conducted to identify genes with actionable mutations. RESULTS: A total of 3 women and 4 men with NF1-MPNST were identified (median age, 38 years). Nonsynonymous mutations were discovered in 4 genes (neurofibromatosis type 1 [NF1], ROS proto-oncogene 1 [ROS1], tumor protein p53 [TP53], and tyrosine kinase 2 [TYK2]), which in addition were mutated in other MPNST cases in this sample set. Consistent with their occurrence in individuals with NF1, all tumors had at least 1 mutation in the NF1 gene. Whereas TP53 gene mutations are frequently observed in other cancers, ROS1 mutations are common in melanoma (15%-35%), anoth- er neural crest-derived malignancy. In contrast, TYK2 mutations are uncommon in other malignancies (<7%).