The Brown Adipocyte Protein CIDEA Promotes Lipid Droplet Fusion
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

GRIM19 Impedes Obesity by Regulating Inflammatory White Fat
cells Article GRIM19 Impedes Obesity by Regulating Inflammatory White Fat Browning and Promoting Th17/Treg Balance JooYeon Jhun 1,†, Jin Seok Woo 1,† , Seung Hoon Lee 2, Jeong-Hee Jeong 1, KyungAh Jung 3, Wonhee Hur 4, Seon-Yeong Lee 1, Jae Yoon Ryu 1, Young-Mee Moon 1, Yoon Ju Jung 5, Kyo Young Song 5, Kiyuk Chang 6, Seung Kew Yoon 4,7 , Sung-Hwan Park 1,8 and Mi-La Cho 1,8,* 1 The Rheumatism Research Center, Catholic Research Institute of Medical Science, The Catholic University of Korea, Seoul 137-040, Korea; [email protected] (J.J.); [email protected] (J.S.W.); [email protected] (J.-H.J.); [email protected] (S.-Y.L.); [email protected] (J.Y.R.); [email protected] (Y.-M.M.); [email protected] (S.-H.P.) 2 Division of Immunology, Department of Microbiology and Immunobiology, Harvard Medical School, Boston, MA 02115, USA; [email protected] 3 Research Center, Impact Biotech, Seoul 137-040, Korea; [email protected] 4 The Catholic University Liver Research Center & WHO Collaborating Center of Viral Hepatitis, College of Medicine, The Catholic University of Korea, Seoul 137-040, Korea; [email protected] (W.H.); [email protected] (S.K.Y.) 5 Division of Gastrointestinal Surgery, Department of General Surgery, Seoul St. Mary’s Hospital, The Catholic University of Korea, Seoul 137-040, Korea; [email protected] (Y.J.J.); [email protected] (K.Y.S.) 6 Cardiovascular Center and Cardiology Division, Seoul St. Mary’s Hospital, College of Medicine, The Catholic University of Korea, Seoul 137-040, Korea; [email protected] 7 Department of Internal Medicine, Seoul St. -

Characterization of the Human CIDEA Promoter in Fat Cells
International Journal of Obesity (2008) 32, 1380–1387 & 2008 Macmillan Publishers Limited All rights reserved 0307-0565/08 $32.00 www.nature.com/ijo ORIGINAL ARTICLE Characterization of the human CIDEA promoter in fat cells AT Pettersson1, J Laurencikiene1, EA Nordstro¨m1, BM Stenson1, V van Harmelen1, C Murphy2, I Dahlman1 and M Ryde´n1 1Department of Medicine, Huddinge, Lipid Laboratory, Novum, Karolinska Institutet, Stockholm, Sweden and 2Department of Laboratory Medicine, Karolinska Institutet, Stockholm, Sweden. Background: Cell death-inducing DFFA (DNA fragmentation factor-a)-like effector A (CIDEA) is a protein that regulates lipolysis in human adipocytes through cross-talk involving tumor necrosis factor-a (TNF-a). TNF-a downregulates CIDEA mRNA although it is unclear whether this is mediated through transcriptional or post-transcriptional mechanisms. CIDEA has important metabolic effects in human fat cells and genetic variations in the human CIDEA gene have been correlated to the development of obesity. However, little is known about the factors regulating CIDEA expression in human adipocytes. We set out to describe the transcriptional control of human CIDEA. Methods: A 1.1-kb genomic fragment upstream of the transcriptional start site (TSS) of human CIDEA was cloned and deletion fragments were generated. Transcriptional activity of the promoter was analyzed by luciferase reporter assays in in vitro- differentiated human adipocytes. The effect of TNF-a was assessed in human adipocytes and murine 3T3-L1 cells transfected with deletion fragments of the CIDEA promoter. Protein–DNA interactions were analyzed by electrophoretic mobility shift assays (EMSA). Results: Basal transcriptional activity was found in a 97-bp region upstream of the TSS. -
![Anti-CIDE a Antibody [V62P1E3*B10] (ARG10830)](https://docslib.b-cdn.net/cover/1546/anti-cide-a-antibody-v62p1e3-b10-arg10830-181546.webp)
Anti-CIDE a Antibody [V62P1E3*B10] (ARG10830)
Product datasheet [email protected] ARG10830 Package: 100 μg anti-CIDE A antibody [V62P1E3*B10] Store at: -20°C Summary Product Description Mouse Monoclonal antibody recognizes CIDE A Tested Reactivity Hu Tested Application IHC-P, WB Host Mouse Clonality Monoclonal Clone V62P1E3*B10 Isotype IgG1, kappa Target Name CIDE A Antigen Species Human Immunogen Ovalbumin-conjugated synthetic peptide. (QAKGRFTCG) Conjugation Un-conjugated Alternate Names CIDE-A; Cell death-inducing DFFA-like effector A; Cell death activator CIDE-A Application Instructions Application table Application Dilution IHC-P Assay-dependent WB Assay-dependent Application Note Antigen Retrieval: Boil tissue section for 10 - 20 min at 800 - 950W microwave with 10 mM Citrate buffer or Sodium citrate buffer (pH 6.0). * The dilutions indicate recommended starting dilutions and the optimal dilutions or concentrations should be determined by the scientist. Calculated Mw 25 kDa Properties Form Liquid Purification Affinity purification with immunogen. Storage instruction For continuous use, store undiluted antibody at 2-8°C for up to a week. For long-term storage, aliquot and store at -20°C or below. Storage in frost free freezers is not recommended. Avoid repeated freeze/thaw cycles. Suggest spin the vial prior to opening. The antibody solution should be gently mixed before use. Note For laboratory research only, not for drug, diagnostic or other use. Bioinformation www.arigobio.com 1/3 Gene Symbol CIDEA Gene Full Name cell death-inducing DFFA-like effector a Background This gene encodes the homolog of the mouse protein Cidea that has been shown to activate apoptosis. This activation of apoptosis is inhibited by the DNA fragmentation factor DFF45 but not by caspase inhibitors. -

The Genetics of Bipolar Disorder
Molecular Psychiatry (2008) 13, 742–771 & 2008 Nature Publishing Group All rights reserved 1359-4184/08 $30.00 www.nature.com/mp FEATURE REVIEW The genetics of bipolar disorder: genome ‘hot regions,’ genes, new potential candidates and future directions A Serretti and L Mandelli Institute of Psychiatry, University of Bologna, Bologna, Italy Bipolar disorder (BP) is a complex disorder caused by a number of liability genes interacting with the environment. In recent years, a large number of linkage and association studies have been conducted producing an extremely large number of findings often not replicated or partially replicated. Further, results from linkage and association studies are not always easily comparable. Unfortunately, at present a comprehensive coverage of available evidence is still lacking. In the present paper, we summarized results obtained from both linkage and association studies in BP. Further, we indicated new potential interesting genes, located in genome ‘hot regions’ for BP and being expressed in the brain. We reviewed published studies on the subject till December 2007. We precisely localized regions where positive linkage has been found, by the NCBI Map viewer (http://www.ncbi.nlm.nih.gov/mapview/); further, we identified genes located in interesting areas and expressed in the brain, by the Entrez gene, Unigene databases (http://www.ncbi.nlm.nih.gov/entrez/) and Human Protein Reference Database (http://www.hprd.org); these genes could be of interest in future investigations. The review of association studies gave interesting results, as a number of genes seem to be definitively involved in BP, such as SLC6A4, TPH2, DRD4, SLC6A3, DAOA, DTNBP1, NRG1, DISC1 and BDNF. -

The Brown Adipocyte Protein CIDEA Promotes Lipid Droplet Fusion
1 The brown adipocyte protein CIDEA promotes lipid droplet 2 fusion via a phosphatidic acid-binding amphipathic helix 3 David Barneda1, Joan Planas-Iglesias2, Maria L. Gaspar3, Dariush Mohammadyani4, 4 Sunil Prasannan2, Dirk Dormann5, Gil-Soo Han5, Stephen A. Jesch3, George M. 5 Carman6, Valerian Kagan4, Malcolm G. Parker1, Nicholas T. Ktistakis7, Judith Klein- 6 Seetharaman2, 4, Ann M. Dixon8, Susan A. Henry3, Mark Christian1,2*. 7 1 Institute of Reproductive and Developmental Biology, Imperial College London, London W12 ONN, 8 UK 9 2 Warwick Medical School, University of Warwick, Coventry, CV4 7AL, UK. 10 3 Department of Molecular Biology and Genetics, Cornell University, Ithaca, New York 14853, USA. 11 4 Department of Bioengineering, University of Pittsburgh, Pittsburgh, Pennsylvania 15219, USA. 12 5 Microscopy Facility, MRC Clinical Sciences Centre, Imperial College London, London W12 0NN, UK 13 6 Department of Food Science, Rutgers Center for Lipid Research, Rutgers University, New Brunswick, 14 New Jersey 08901, USA. 15 7 Signalling Programme, Babraham Institute, Cambridge CB22 3AT, UK. 16 8 Department of Chemistry, University of Warwick, Coventry, CV4 7AL, UK. 17 18 19 *Corresponding author. 20 E-mail: [email protected] 21 Phone number: 44 2476 96 8585 1 22 23 Summary 24 Maintenance of energy homeostasis depends on the highly regulated storage and 25 release of triacylglycerol primarily in adipose tissue and excessive storage is a feature of 26 common metabolic disorders. CIDEA is a lipid droplet (LD)-protein enriched in brown 27 adipocytes promoting the enlargement of LDs which are dynamic, ubiquitous organelles 28 specialized for storing neutral lipids. -

Proteomic Analysis of Monolayer-Integrated Proteins on Lipid Droplets Identifies Amphipathic Interfacial Α-Helical Membrane Anchors
Proteomic analysis of monolayer-integrated proteins on lipid droplets identifies amphipathic interfacial α-helical membrane anchors Camille I. Patakia, João Rodriguesb, Lichao Zhangc, Junyang Qiand, Bradley Efrond, Trevor Hastied, Joshua E. Eliasc, Michael Levittb, and Ron R. Kopitoe,1 aDepartment of Biochemistry, Stanford University, Stanford, CA 94305; bDepartment of Structural Biology, Stanford University, Stanford, CA 94305; cDepartment of Chemical and Systems Biology, Stanford University, Stanford, CA 94305; dDepartment of Statistics, Stanford University, Stanford, CA 94305; and eDepartment of Biology, Stanford University, Stanford, CA 94305 Edited by Jennifer Lippincott-Schwartz, Howard Hughes Medical Institute, Ashburn, VA, and approved July 23, 2018 (received for review May 9, 2018) Despite not spanning phospholipid bilayers, monotopic integral The first step in understanding the structural diversity of proteins (MIPs) play critical roles in organizing biochemical reac- monotopic HMDs is to identify a sufficiently large set of proteins tions on membrane surfaces. Defining the structural basis by with unequivocal monotopic topology. In this study, we exploited which these proteins are anchored to membranes has been the fact that lipid droplets (LDs) are enveloped by phospholipid hampered by the paucity of unambiguously identified MIPs and a monolayers that separate a highly hydrophobic lipid core from lack of computational tools that accurately distinguish monolayer- the aqueous cytoplasm (8). Because TMDs are flanked on both integrating motifs from bilayer-spanning transmembrane domains ends by soluble, hydrophilic domains, bilayer-spanning proteins (TMDs). We used quantitative proteomics and statistical modeling are strongly disfavored from integrating into the monolayer to identify 87 high-confidence candidate MIPs in lipid droplets, membranes of LDs. -

PRDM16 Binds MED1 and Controls Chromatin Architecture to Determine a Brown Fat Transcriptional Program
Downloaded from genesdev.cshlp.org on September 24, 2021 - Published by Cold Spring Harbor Laboratory Press PRDM16 binds MED1 and controls chromatin architecture to determine a brown fat transcriptional program Matthew J. Harms,1,2,4 Hee-Woong Lim,1,3,4 Yugong Ho,3 Suzanne N. Shapira,1,2 Jeff Ishibashi,1,2 Sona Rajakumari,1,2 David J. Steger,1 Mitchell A. Lazar,1 Kyoung-Jae Won,1,3 and Patrick Seale1,2 1Institute for Diabetes, Obesity, and Metabolism, 2Department of Cell and Developmental Biology, 3Genetics Department, Smilow Center for Translational Research, Perelman School of Medicine, University of Pennsylvania, Philadelphia, Pennsylvania 19104, USA PR (PRD1–BF1–RIZ1 homologous) domain-containing 16 (PRDM16) drives a brown fat differentiation program, but the mechanisms by which PRDM16 activates brown fat-selective genes have been unclear. Through chromatin immunoprecipitation (ChIP) followed by deep sequencing (ChIP-seq) analyses in brown adipose tissue (BAT), we reveal that PRDM16 binding is highly enriched at a broad set of brown fat-selective genes. Importantly, we found that PRDM16 physically binds to MED1, a component of the Mediator complex, and recruits it to superenhancers at brown fat-selective genes. PRDM16 deficiency in BAT reduces MED1 binding at PRDM16 target sites and causes a fundamental change in chromatin architecture at key brown fat-selective genes. Together, these data indicate that PRDM16 controls chromatin architecture and superenhancer activity in BAT. [Keywords: PRDM16; PRDM3; MED1; Mediator; brown adipose tissue; UCP1] Supplemental material is available for this article. Received September 17, 2014; revised version accepted December 10, 2014. Obesity is a leading cause of preventable death in the United required for beige fat differentiation and the maintenance of States due to its link to many diseases, including type 2 brown adipose tissue (BAT) fate in adult mice (Cohen diabetes, cardiovascular disease, stroke, and certain cancers et al. -
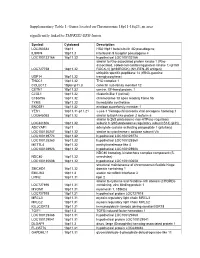
Supplementary Table 1: Genes Located on Chromosome 18P11-18Q23, an Area Significantly Linked to TMPRSS2-ERG Fusion
Supplementary Table 1: Genes located on Chromosome 18p11-18q23, an area significantly linked to TMPRSS2-ERG fusion Symbol Cytoband Description LOC260334 18p11 HSA18p11 beta-tubulin 4Q pseudogene IL9RP4 18p11.3 interleukin 9 receptor pseudogene 4 LOC100132166 18p11.32 hypothetical LOC100132166 similar to Rho-associated protein kinase 1 (Rho- associated, coiled-coil-containing protein kinase 1) (p160 LOC727758 18p11.32 ROCK-1) (p160ROCK) (NY-REN-35 antigen) ubiquitin specific peptidase 14 (tRNA-guanine USP14 18p11.32 transglycosylase) THOC1 18p11.32 THO complex 1 COLEC12 18pter-p11.3 collectin sub-family member 12 CETN1 18p11.32 centrin, EF-hand protein, 1 CLUL1 18p11.32 clusterin-like 1 (retinal) C18orf56 18p11.32 chromosome 18 open reading frame 56 TYMS 18p11.32 thymidylate synthetase ENOSF1 18p11.32 enolase superfamily member 1 YES1 18p11.31-p11.21 v-yes-1 Yamaguchi sarcoma viral oncogene homolog 1 LOC645053 18p11.32 similar to BolA-like protein 2 isoform a similar to 26S proteasome non-ATPase regulatory LOC441806 18p11.32 subunit 8 (26S proteasome regulatory subunit S14) (p31) ADCYAP1 18p11 adenylate cyclase activating polypeptide 1 (pituitary) LOC100130247 18p11.32 similar to cytochrome c oxidase subunit VIc LOC100129774 18p11.32 hypothetical LOC100129774 LOC100128360 18p11.32 hypothetical LOC100128360 METTL4 18p11.32 methyltransferase like 4 LOC100128926 18p11.32 hypothetical LOC100128926 NDC80 homolog, kinetochore complex component (S. NDC80 18p11.32 cerevisiae) LOC100130608 18p11.32 hypothetical LOC100130608 structural maintenance -

Distribution and Quantitative Analysis of Cidea and Cidec in Broiler Chickens: Accounting for Differential Fat Deposition Between Strains
British Poultry Science ISSN: 0007-1668 (Print) 1466-1799 (Online) Journal homepage: http://www.tandfonline.com/loi/cbps20 Distribution and quantitative analysis of CIDEa and CIDEc in broiler chickens: accounting for differential fat deposition between strains JZ. Huang, LM. Huang, QJ Zeng, EF. Huang, HP. Liang, Q. Wei, XH. Xie & JM. Ruan To cite this article: JZ. Huang, LM. Huang, QJ Zeng, EF. Huang, HP. Liang, Q. Wei, XH. Xie & JM. Ruan (2017): Distribution and quantitative analysis of CIDEa and CIDEc in broiler chickens: accounting for differential fat deposition between strains, British Poultry Science, DOI: 10.1080/00071668.2017.1415426 To link to this article: https://doi.org/10.1080/00071668.2017.1415426 Accepted author version posted online: 08 Dec 2017. Published online: 20 Dec 2017. Submit your article to this journal Article views: 7 View related articles View Crossmark data Full Terms & Conditions of access and use can be found at http://www.tandfonline.com/action/journalInformation?journalCode=cbps20 Download by: [RMIT University Library] Date: 23 December 2017, At: 01:09 BRITISH POULTRY SCIENCE, 2017 https://doi.org/10.1080/00071668.2017.1415426 Distribution and quantitative analysis of CIDEa and CIDEc in broiler chickens: accounting for differential fat deposition between strains JZ. Huanga, LM. Huangb, QJ Zenga, EF. Huangc, HP. Lianga, Q. Weia, XH. Xiea and JM. Ruana aJiangxi Provincial Key Laboratory for Animal Health, Institute of Animal Population Health, College of Animal Science and Technology, Jiangxi Agricultural University, Nanchang, P. R. China; bCollege of Life Sciences and Oceanography, Shenzhen University, Shenzhen, P. R. China; cDepartment of Animal Science, Jiangxi Biotech Vocational College, Nanchang, P. -

Differential Gene Expression by RNA-Seq Analysis of the Primo Vessel in the Rabbit Lymph
J Acupunct Meridian Stud 2019;12(1):11e19 Available online at www.sciencedirect.com Journal of Acupuncture and Meridian Studies journal homepage: www.jams-kpi.com Research Article Differential Gene Expression by RNA-Seq Analysis of the Primo Vessel in the Rabbit Lymph Jun-Young Shin, Sang-Heon Choi, Da-Woon Choi, Ye-Jin An, Jae-Hyuk Seo, Jong-Gu Choi, Min-Suk Rho, Sang-Suk Lee* Department of Oriental Biomedical Engineering, Sangji University, Wonju, Republic of Korea Available online 28 October 2018 Received: May 31, 2018 Abstract Revised: Jul 26, 2018 For the connectome of primo vascular system, some long-type primo vessels dyed with Accepted: Oct 23, 2018 Alcian blue injected into inguinal nodes, abdominal node, and axially nodes were visual- ized, which passed over around the vena cava of the rabbit. The Alcian blue dye revealed KEYWORDS primo vessels and colored blue in the rabbit lymph vessels. The length of long-type primo e m gene expression level; vessels was 18 cm on average, of which diameters were about 20 30 m, and the lymph e m lymph node; vessels had diameters of 100 150 m. Three different tissues of pure primo vessel, mixed primo connectome; primo þ lymph vessel, and only lymph vessel were made to undergo RNA-Seq analysis by RNA-Seq analysis next-generation sequencing. We also analyzed differentially expressed genes (DEGs) from the RNA-Seq data, in which 30 genes of the primo vessels, primo þ lymph vessels, and lymph vessels were selected for primo marker candidates. From the plot of DEG analysis, 10 genes had remarkably different expression pattern on the Group 1 (primo vessel) vs Group 3 (lymph vessel). -

Sympathetic Inputs Regulate Adaptive Thermogenesis in Brown Adipose
www.nature.com/scientificreports OPEN Sympathetic inputs regulate adaptive thermogenesis in brown adipose tissue through cAMP-Salt Received: 9 November 2017 Accepted: 10 July 2018 inducible kinase axis Published: xx xx xxxx Esther Paulo1, Dongmei Wu1,5, Yangmeng Wang1,6, Yun Zhang1, Yixuan Wu1, Danielle L. Swaney2,3,4, Margaret Soucheray2,3,4, David Jimenez-Morales 2,3,4, Ajay Chawla1, Nevan J. Krogan2,3,4 & Biao Wang 1 Various physiological stimuli, such as cold environment, diet, and hormones, trigger brown adipose tissue (BAT) to produce heat through sympathetic nervous system (SNS)- and β-adrenergic receptors (βARs). The βAR stimulation increases intracellular cAMP levels through heterotrimeric G proteins and adenylate cyclases, but the processes by which cAMP modulates brown adipocyte function are not fully understood. Here we described that specifc ablation of cAMP production in brown adipocytes led to reduced lipolysis, mitochondrial biogenesis, uncoupling protein 1 (Ucp1) expression, and consequently defective adaptive thermogenesis. Elevated cAMP signaling by sympathetic activation inhibited Salt- inducible kinase 2 (Sik2) through protein kinase A (PKA)-mediated phosphorylation in brown adipose tissue. Inhibition of SIKs enhanced Ucp1 expression in diferentiated brown adipocytes and Sik2 knockout mice exhibited enhanced adaptive thermogenesis at thermoneutrality in an Ucp1-dependent manner. Taken together, our data indicate that suppressing Sik2 by PKA-mediated phosphorylation is a requisite for SNS-induced Ucp1 expression and adaptive thermogenesis in BAT, and targeting Sik2 may present a novel therapeutic strategy to ramp up BAT thermogenic activity in humans. Energy balance requires equivalent energy intake and energy expenditure, and when energy intake exceeds energy expenditure, animals store excess energy as fat in adipose and other metabolic tissues. -

The Brown Adipocyte Protein CIDEA Promotes
RESEARCH ARTICLE The brown adipocyte protein CIDEA promotes lipid droplet fusion via a phosphatidic acid-binding amphipathic helix David Barneda1, Joan Planas-Iglesias2, Maria L Gaspar3, Dariush Mohammadyani4, Sunil Prasannan5, Dirk Dormann6, Gil-Soo Han7, Stephen A Jesch3, George M Carman7, Valerian Kagan4, Malcolm G Parker1, Nicholas T Ktistakis8, Judith Klein-Seetharaman2,4, Ann M Dixon5, Susan A Henry3, Mark Christian1,2* 1Institute of Reproductive and Developmental Biology, Imperial College London, London, United Kingdom; 2Warwick Medical School, University of Warwick, Coventry, United Kingdom; 3Department of Molecular Biology and Genetics, Cornell University, Ithaca, United States; 4Department of Bioengineering, University of Pittsburgh, Pittsburgh, United States; 5Department of Chemistry, University of Warwick, Coventry, United Kingdom; 6Microscopy Facility, MRC Clinical Sciences Centre, Imperial College London, London, United Kingdom; 7Department of Food Science, Rutgers Center for Lipid Research, Rutgers University, New Brunswick, United States; 8Signalling Programme, Babraham Institute, Cambridge, United Kingdom Abstract Maintenance of energy homeostasis depends on the highly regulated storage and release of triacylglycerol primarily in adipose tissue, and excessive storage is a feature of common metabolic disorders. CIDEA is a lipid droplet (LD)-protein enriched in brown adipocytes promoting the enlargement of LDs, which are dynamic, ubiquitous organelles specialized for storing neutral *For correspondence: lipids. We demonstrate an essential role in this process for an amphipathic helix in CIDEA, which [email protected] facilitates embedding in the LD phospholipid monolayer and binds phosphatidic acid (PA). LD pairs Competing interests: The are docked by CIDEA trans-complexes through contributions of the N-terminal domain and a C- authors declare that no terminal dimerization region.