Functional Evolution of C4 Pyruvate, Orthophosphate Dikinase
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

METACYC ID Description A0AR23 GO:0004842 (Ubiquitin-Protein Ligase
Electronic Supplementary Material (ESI) for Integrative Biology This journal is © The Royal Society of Chemistry 2012 Heat Stress Responsive Zostera marina Genes, Southern Population (α=0. -

In the Field of Clinical Examination, the Measurement of Creatine
J. Clin. Biochem. Nutr., 3, 17-25, 1987 Bacterial Glucokinase as an Enzymic Reagent of Good Stability for Measurement of Creatine Kinase Activity Hitoshi KONDO, * Takanari SHIRAISHI, Masao KAGEYAMA, Kazuhiko NAGATA, and Kosuke TOMITA Research and Development Center, UNITIKA Ltd., Uji 611, Japan (Received January 10, 1987) Summary An enzymic reagent, that has long-term stability even in the liquid state, was successfully employed for the measurement of serum creatine kinase (CK, EC 2.7.3.2) activity. The enzyme used was the thermostable glucokinase (GlcK, EC 2.7.1.2) obtained from the thermo- phile Bacillus stearothermophilus. The reagent was found to be stable in solution for about one month at 6•Ž and for about one week at 30•Ž. This substitution of glucokinase for the hexokinase of the most commonly used hexokinase-glucose-6-phosphate dehydrogenase (HK-G6PDH) method results in a remarkable improvement of the method. The CK activity measured by the GlcK-G6PDH method was linear up to about 2,000 U/ liter at 37•Ž. The GlcK-G6PDH method was found to give a satisfactory precision and reproducibility (coefficient of variation less than 2.17%). Over a wide range of CK activity, an excellent agreement was obtained between the GlcK-G6PDH and the HK-G6PDH methods. Furthermore several coexistents and anticoagulants were found to have little effect on the measured value of CK activity by the GlcK-G6PDH method. Key Words: creatine kinase activity, glucokinase, improved stability of reagent, creatine kinase determination, thermostable enzyme In the field of clinical examination, the measurement of creatine kinase (CK, ATP : creatine phosphotransferase, EC 2.7.3.2) activity in serum is one of the important examinations usually employed for diagnosis of cardiac diseases such as myocardial infarction or muscular diseases such as progressive muscular dystrophy. -

Phosphotransferase Activity of Liver Mitochondria (Oxidative Phosphorylation/Adenine Nucleotide Ni-Oxides/Substrate Specificity/In Vivo Phosphorylation) G
Proc. Nat. Acad. Sci. USA Vol. 71, No. 11, pp. 4630-4634, November 1974 Participation of N1-Oxide Derivatives of Adenine Nucleotides in the Phosphotransferase Activity of Liver Mitochondria (oxidative phosphorylation/adenine nucleotide Ni-oxides/substrate specificity/in vivo phosphorylation) G. JEBELEANU, N. G. TY, H. H. MANTSCH*, 0. BARZU, G. NIACt, AND I. ABRUDAN Department of Biochemistry, Medical and Pharmaceutical Institute, Cluj, * Institute of Chemistry, University of Cluj, Cluj, and t Department of Physical Chemistry, University of Craiova, Craiova, Romania Communicated by Henry Lardy, September 3, 1974 ABSTRACT The modified adenine nucleotides ATP- rivatives of adenine nucleotides once produced become "ac- NO, ADP-NO, and AMP-NO were tested as potential tive" components of the mitochondrial or cellular adenylate substrates and/or inhibitors of mitochondrial phospho- transferases. ADP-NO is not recognized by the translocase pool. membrane; system located in the inner mitochondrial AND METHODS however, it is rapidly phosphorylated to ATP-NO in the MATERIALS outer compartment of mitochondria, by way of the The following commercially available chemicals were used: nucleosidediplhosphate kinase (EC 2.7.4.6) reaction, pro-' de- vitded there is sufficient ATP in the mitochondria. AMP- crystalline bovine serum albumin, glucose-6-phosphate NO is not phosphorylated by liver mitochondria to the hydrogenase (Glc-6-P dehydrogenase; EC 1.1.1.49; BDH corresponding nucleoside diphosphate; it cannot serve as Chemicals, Ltd.), yeast hexokinase (EC 2.7.1.1) (Nutritional substrate for adenylate kinase (EC 2.7.4.3). ATP-NO and Biochemicals, Cleveland), (log muscle lactate dehydrogenase ADP-NO, however, are substrates of this enzyme. -

Isozymes of Pyruvate Kinase in Liver and Hepatomas of the Rat1
[CANCER RESEARCH 34, 1439-1446, June 1974] Isozymes of Pyruvate Kinase in Liver and Hepatomas of the Rat1 Francis A. Farina,2 Jennie B. Shatton, Harold P. Morris, and Sidney Weinhouse The Fels Research Institute and the Department of Biochemistry, Temple University School oj Medicine, Philadelphia. Pennsylvania IV140 (F. A. F., J. B. S.. S. W.\, and the Department of Biochemistry. Howard University School of Medicine. Washington. D. C. 20001 [H. P. M .\ SUMMARY 23, 32, 41, 57). These alterations involve the replacement of those isozymes that are under dietary and hormonal control Pyruvate kinase (PK) (EC 2.7.1.40) isozymes were assayed by the host, and that have important metabolic functions in in normal rat liver and a series of transplantable rat the adult differentiated liver by other isozymes which are hepatomas ranging widely in growth rate and degree of normally either low in, or absent from, the adult tissue. As differentiation, with the use of gradient elution by chloride part of an ongoing study of this phenomenon, we have ion from columns of DEAE-cellulose. In agreement with examined the alteration of PK3 (EC 2.7.1.40) isozymes in other studies, three noninterconvertible forms were found in the Morris hepatomas (30. 31), a series of chemically rat tissues: isozyme I, the major form in adult rat liver: induced, transplantable rat hepatomas ranging widely in isozyme II, the sole form in heart and skeletal muscle: and growth rate and degree of differentiation. This enzyme isozyme III, the sole form in poorly differentiated hepato occupies a key position in the metabolism of cells and, as we mas, the major form in normal kidney and lung, and the pointed out previously (27, 28. -

Labeled in Thecourse of Glycolysis, Since Phosphoglycerate Kinase
THE STATE OF MAGNESIUM IN CELLS AS ESTIMATED FROM THE ADENYLATE KINASE EQUILIBRIUM* BY TRWIN A. RoSE THE INSTITUTE FOR CANCER RESEARCH, PHILADELPHIA Communicated by Thomas F. Anderson, August 30, 1968 Magnesium functions in many enzymatic reactions as a cofactor and in com- plex with nucleotides acting as substrates. Numerous examples of a possible regulatory role of Mg can be cited from studies with isolated enzymes,'- and it is known that Mg affects the structural integrity of macromolecules such as trans- fer RNA" and functional elements such as ribosomes.'0 The major problem in translating this information on isolated preparations to the functioning cell is the difficulty in determining the distribution of Mg and the nucleotides among the free and complexed forms that function in the region of the cell for which this information is desired. Nanningall based an attempt to calculate the free Mg2+ and Ca2+ ion concentrations of frog muscle on the total content of these metals and of the principal known ligands (adenosine 5'-triphosphate (ATP), creatine-P, and myosin) and the dissociation constants of the complexes. However, this method suffers from the necessity of evaluating the contribution of all ligands as well as from the assumption that all the known ligands are contributing their full complexing capacity. During studies concerned with the control of glycolysis in red cells and the control of the phosphoglycerate kinase step in particular, it became important to determine the fractions of the cell's ATP and adenosine 5'-diphosphate (ADP) that were present as Mg complexes. Just as the problem of determining the distribution of protonated and dissociated forms of an acid can be solved from a knowledge of pH and pKa of the acid, so it would be possible to determine the liganded and free forms of all rapidly established Mg complexes from a knowledge of Mg2+ ion concentration and the appropriate dissociation constants. -

Pyruvate Kinase: Function, Regulation and Role in Cancer
Pyruvate kinase: Function, regulation and role in cancer The MIT Faculty has made this article openly available. Please share how this access benefits you. Your story matters. Citation Israelsen, William J., and Matthew G. Vander Heiden. “Pyruvate Kinase: Function, Regulation and Role in Cancer.” Seminars in Cell & Developmental Biology 43 (2015): 43–51. As Published http://dx.doi.org/10.1016/j.semcdb.2015.08.004 Publisher Elsevier Version Author's final manuscript Citable link http://hdl.handle.net/1721.1/105833 Terms of Use Creative Commons Attribution-NonCommercial-NoDerivs License Detailed Terms http://creativecommons.org/licenses/by-nc-nd/4.0/ HHS Public Access Author manuscript Author Manuscript Author ManuscriptSemin Cell Author Manuscript Dev Biol. Author Author Manuscript manuscript; available in PMC 2016 August 13. Published in final edited form as: Semin Cell Dev Biol. 2015 July ; 43: 43–51. doi:10.1016/j.semcdb.2015.08.004. Pyruvate kinase: function, regulation and role in cancer William J. Israelsena,1,* and Matthew G. Vander Heidena,b,* aKoch Institute for Integrative Cancer Research, Massachusetts Institute of Technology, Cambridge, MA 02139, USA bDepartment of Medical Oncology, Dana-Farber Cancer Institute, Boston, MA 02115, USA Abstract Pyruvate kinase is an enzyme that catalyzes the conversion of phosphoenolpyruvate and ADP to pyruvate and ATP in glycolysis and plays a role in regulating cell metabolism. There are four mammalian pyruvate kinase isoforms with unique tissue expression patterns and regulatory properties. The M2 isoform of pyruvate kinase (PKM2) supports anabolic metabolism and is expressed both in cancer and normal tissue. The enzymatic activity of PKM2 is allosterically regulated by both intracellular signaling pathways and metabolites; PKM2 thus integrates signaling and metabolic inputs to modulate glucose metabolism according to the needs of the cell. -
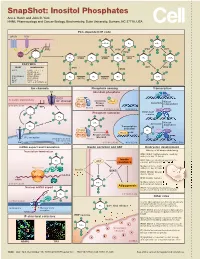
Snapshot: Inositol Phosphates Ace J
SnapShot: Inositol Phosphates Ace J. Hatch and John D. York HHMI, Pharmacology and Cancer Biology, Biochemistry, Duke University, Durham, NC 27710, USA PLC-dependent IP code GPCR RTK O O O O O O 5-PP-IP4 IP4 5-IP7 O O O O O O PIP2 O IP6K O IP6K O VIP1 O O O 2 O ITPK1 O 13 O PLC 2 O O O O O O O O 4 6 13 O 5 IP3 IPMK IP4 IPMK IP5 IPK1 IP6 1,5-IP8 4 6 O O 5 O O O O O O O O ENZYMES O O O O O O YEAST MAMMALIAN IP3K VIP1 IP6K IPMK PLC1 PLCβ, γ, δ, ε, ζ, η O - IP3KA, B, C - ITPK1 (IP56K) O O O O O O O O IPK2(ARG82) IPMK (IPK2) IP4 IP3 IP4 1-IP7 IPK1 IPK1 (IP5K) INPP5 ITPK1 KCS1 IP6K1, 2, 3 O O O O O O VIP1 VIP1, 2 (PPIP5K1, 2) O O Ion channels Phosphate sensing Transcription Cl- Abundant phosphate MCM1 ARG80 CIC3 P PLASMA MEMBRANE - Pho80 Cl channel Pho4 Kinase Kinase Assembly Pho85 independent CYTOPLASM activity 2 O PIP2 Pho81 13 CYTOPLASM NUCLEUS IPK2 ARG81 4 6 Phosphate starvation MCM1-ArgR O 5 O complex O O IP4 O O O O O O O 1-IP7 Kinase Activation dependent IP3 O O Transcription O O O activated Pho80 IP4 O X Pho4 O O Pho85 Kinase activity IP receptor blocked O 3 ENDOPLASMIC Pho81 RETICULUM Ca2+ CYTOPLASM NUCLEUS NUCLEUS mRNA export and translation Insulin secretion and AKT Embryonic development Translation termination Effects of IP kinase deficiency O IPMK (IPK2): Multiple defects, death by embryonic day 10 (mice) O O Insulin IPK1: Cillia are shortened and immotile IP6 AKT resistance causing patterning defects (zebrash) O O Multiple defects, death by Ribosome O embryonic day 8.5 (mice) GleI eRF1 Insulin GSK3β Dbp5 ITPK1 (IP56K): Neural tube -

The Characterization of Human Adenylate Kinases 7 and 8
The characterization of human adenylate kinases 7 and 8 demonstrates differences in kinetic parameters and structural organization among the family of adenylate kinase isoenzymes Christakis Panayiotou, Nicola Solaroli, Yunjian Xu, Magnus Johansson, Anna Karlsson To cite this version: Christakis Panayiotou, Nicola Solaroli, Yunjian Xu, Magnus Johansson, Anna Karlsson. The char- acterization of human adenylate kinases 7 and 8 demonstrates differences in kinetic parameters and structural organization among the family of adenylate kinase isoenzymes. Biochemical Journal, Port- land Press, 2011, 433 (3), pp.527-534. 10.1042/BJ20101443. hal-00558097 HAL Id: hal-00558097 https://hal.archives-ouvertes.fr/hal-00558097 Submitted on 21 Jan 2011 HAL is a multi-disciplinary open access L’archive ouverte pluridisciplinaire HAL, est archive for the deposit and dissemination of sci- destinée au dépôt et à la diffusion de documents entific research documents, whether they are pub- scientifiques de niveau recherche, publiés ou non, lished or not. The documents may come from émanant des établissements d’enseignement et de teaching and research institutions in France or recherche français ou étrangers, des laboratoires abroad, or from public or private research centers. publics ou privés. Biochemical Journal Immediate Publication. Published on 16 Nov 2010 as manuscript BJ20101443 The characterization of human adenylate kinases 7 and 8 demonstrates differences in kinetic parameters and structural organization among the family of adenylate kinase isoenzymes -

Effect of Enzyme/Substrate Ratio on the Antioxidant Properties of Hydrolysed African Yam Bean
African Journal of Biotechnology Vol. 11(50), pp. 11086-11091, 21 June, 2012 Available online at http://www.academicjournals.org/AJB DOI: 10.5897/AJB11.2271 ISSN 1684–5315 ©2012 Academic Journals Full Length Research Paper Effect of enzyme/substrate ratio on the antioxidant properties of hydrolysed African yam bean Fasasi Olufunmilayo*, Oyebode Esther and Fagbamila Oluwatoyin Department of Food Science and Technology, P. M. B. 704, Federal University of Technology, Akure, Nigeria. Accepted 8 June, 2012 The use of natural antioxidant as compared with synthetic antioxidant in food processing is a growing trend as consumers prefer natural to synthetic antioxidant mainly on emotional ground. This study investigates the antioxidant activity of hydrolysed African yam bean (Sphenostylis sternocarpa) which is regarded as one of the neglected underutilized species (NUS) of crop in Africa and Nigeria especially to improve food security and boost the economic importance of the crop. The antioxidant properties of African yam bean hydrolysates (AYH) produced at different enzyme to substrate (E/S) ratios of 1: 100 and 3: 100 (W/V) using pepsin (pH 2.0, 37°C) were studied. 2, 2-Diphenyl-1-picryl-hydrazyl (DPPH) radical scavenging activity of the hydrolysates was significantly influenced by the E\S ratio as DPPH radical scavenging activity ranged from 56.1 to 75.8% in AYH (1: 100) and 33.3 to 58.8% in AYH (3:100) with 1000 µg/ml having the highest DPPH radical scavenging activity. AYH (1:100) and AYH (3:100) had higher reducing activities of 0.42 and 0.23, respectively at a concentration of 1000 µg/ml. -

Regulation of Energy Substrate Metabolism in Endurance Exercise
International Journal of Environmental Research and Public Health Review Regulation of Energy Substrate Metabolism in Endurance Exercise Abdullah F. Alghannam 1,* , Mazen M. Ghaith 2 and Maha H. Alhussain 3 1 Lifestyle and Health Research Center, Health Sciences Research Center, Princess Nourah bInt. Abdulrahman University, Riyadh 84428, Saudi Arabia 2 Faculty of Applied Medical Sciences, Laboratory Medicine Department, Umm Al-Qura University, Al Abdeyah, Makkah 7607, Saudi Arabia; [email protected] 3 Department of Food Science and Nutrition, College of Food and Agriculture Sciences, King Saud University, Riyadh 11451, Saudi Arabia; [email protected] * Correspondence: [email protected] Abstract: The human body requires energy to function. Adenosine triphosphate (ATP) is the cellular currency for energy-requiring processes including mechanical work (i.e., exercise). ATP used by the cells is ultimately derived from the catabolism of energy substrate molecules—carbohydrates, fat, and protein. In prolonged moderate to high-intensity exercise, there is a delicate interplay between carbohydrate and fat metabolism, and this bioenergetic process is tightly regulated by numerous physiological, nutritional, and environmental factors such as exercise intensity and du- ration, body mass and feeding state. Carbohydrate metabolism is of critical importance during prolonged endurance-type exercise, reflecting the physiological need to regulate glucose homeostasis, assuring optimal glycogen storage, proper muscle fuelling, and delaying the onset of fatigue. Fat metabolism represents a sustainable source of energy to meet energy demands and preserve the ‘limited’ carbohydrate stores. Coordinated neural, hormonal and circulatory events occur during prolonged endurance-type exercise, facilitating the delivery of fatty acids from adipose tissue to the Citation: Alghannam, A.F.; Ghaith, working muscle for oxidation. -

Prevalence of Pyruvate Kinase Deficiency
P3513 Prevalence of Red Cell Pyruvate Kinase Deficiency: A Systematic Literature Review Mike Storm1, Matthew H Secrest2*, Courtney Carrington2, Deb Casso2, Keely Gilroy1, Leanne Pladson1, Audra N Boscoe1 1Agios Pharmaceuticals Inc., Cambridge, MA, USA; 2IQVIA Epidemiology & Drug Safety, Seattle, WA and Cambridge, MA, USA; *Affiliation at the time research was conducted BACKGROUND METHODS (continued) RESULTS (continued) RESULTS (continued) • Pyruvate Kinase (PK) deficiency is a rare congenital hemolytic anemia Exclusion Criteria The remaining 34 studies were grouped based on methods and study Among these 4 studies, an important distinction was made between studies characterized by diminished activity of the PK enzyme in red blood population (Table 1). reporting diagnosed prevalence (n=3) and overall disease prevalence cells (RBC).1 • Non-human studies; (diagnosed and undiagnosed PK deficiency; n=1). Table 1. Distribution of extracted studies by type of study (n=34) • Low PK enzyme activity can lead to lifelong chronic hemolysis with • Publications that were not the primary report of the data • Two studies estimated diagnosed PK deficiency prevalence as 3.2 per associated symptoms and complications such as anemia, jaundice, (e.g., literature reviews); Type of study Number million4 and 8.5 per million5 by identifying diagnosed PK deficiency cases of studies gallstones, splenectomy and associated thrombosis, iron overload, and • Studies of PK deficiency prevalence/incidence conducted within a source from source populations of known size. liver cirrhosis.2 Population-based prevalence 2 population of patients with symptoms of PK deficiency such as anemia • We estimated the prevalence of diagnosed PK deficiency in a general • PK deficiency is caused by compound heterozygosity or homozygosity for or jaundice; Molecular PKLR screening in a general population 5 population to be 6.5 per million6 using data from another high-quality study 3 one or more of the >300 known mutations to the PKLR gene. -

Phosphoenolpyruvate-Dependent Phosphotransferase System in Lactobacillus Casei BRUCE M
JOURNAL OF BACTERIOLOGY, June 1983, p. 1204-1214 Vol. 154, No. 3 0021-9193/83/061204-11$02.00/0 Copyright C 1983, American Society for Microbiology Regulation and Characterization of the Galactose- Phosphoenolpyruvate-Dependent Phosphotransferase System in Lactobacillus casei BRUCE M. CHASSY* AND JOHN THOMPSON Microbiology Section, Laboratory of Microbiology and Immunology, National Institute of Dental Research, Bethesda, Maryland 20205 Received 8 November 1982/Accepted 5 March 1983 Cells ofLactobacillus casei grown in media containing galactose or a metaboliz- able ,-galactoside (lactose, lactulose, or arabinosyl-P-D-galactoside) were in- duced for a galactose-phosphoenolpyruvate-dependent phosphotransferase sys- tem (gal-PTS). This high-affinity system (Km for galactose, 11 ,uM) was inducible in eight strains examined, which were representative of all five subspecies of L. casei. The gal-PTS was also induced in strains defective in glucose- and lactose- phosphoenolpyruvate-dependent phosphotransferase systems during growth on galactose. Galactose 6-phosphate appeared to be the intracellular inducer of the gal-PTS. The gal-PTS was quite specific for D-galactose, and neither glucose, lactose, nor a variety of structural analogs of galactose caused significant inhibition of phosphotransferase system-mediated galactose transport in intact cells. The phosphoenolpyruvate-dependent phosphorylation of galactose in vitro required specific membrane and cytoplasmic components (including enzyme Illgal), which were induced only by growth of the cells on galactose or ,B- galactosides. Extracts prepared from such cells also contained an ATP-dependent galactokinase which converted galactose to galactose 1-phosphate. Our results demonstrate the separate identities of the gal-PTS and the lactose-phosphoenol- pyruvate-dependent phosphotransferase system in L.