Mouse Cadm3 Conditional Knockout Project (CRISPR/Cas9)
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Supplementary Information
Supplementary Information This text file includes: Supplementary Methods Supplementary Figure 1-13, 15-30 Supplementary Table 1-8, 16, 20-21, 23, 25-37, 40-41 1 1. Samples, DNA extraction and genome sequencing 1.1 Ethical statements and sample storage The ethical statements of collecting and processing tissue samples for each species are listed as follows: Myotis myotis: All procedures were carried out in accordance with the ethical guidelines and permits (AREC-13-38-Teeling) delivered by the University College Dublin and the Préfet du Morbihan, awarded to Emma Teeling and Sébastien Puechmaille respectively. A single M. myotis individual was humanely sacrificed given that she had lethal injuries, and dissected. Rhinolophus ferrumequinum: All the procedures were conducted under the license (Natural England 2016-25216-SCI-SCI) issued to Gareth Jones. The individual bat died unexpectedly and suddenly during sampling and was dissected immediately. Pipistrellus kuhlii: The sampling procedure was carried out following all the applicable national guidelines for the care and use of animals. Sampling was done in accordance with all the relevant wildlife legislation and approved by the Ministry of Environment (Ministero della Tutela del Territorio e del Mare, Aut.Prot. N˚: 13040, 26/03/2014). Molossus molossus: All sampling methods were approved by the Ministerio de Ambiente de Panamá (SE/A-29-18) and by the Institutional Animal Care and Use Committee of the Smithsonian Tropical Research Institute (2017-0815-2020). Phyllostomus discolor: P. discolor bats originated from a breeding colony in the Department Biology II of the Ludwig-Maximilians-University in Munich. Approval to keep and breed the bats was issued by the Munich district veterinary office. -

Whole Exome Sequencing in Families at High Risk for Hodgkin Lymphoma: Identification of a Predisposing Mutation in the KDR Gene
Hodgkin Lymphoma SUPPLEMENTARY APPENDIX Whole exome sequencing in families at high risk for Hodgkin lymphoma: identification of a predisposing mutation in the KDR gene Melissa Rotunno, 1 Mary L. McMaster, 1 Joseph Boland, 2 Sara Bass, 2 Xijun Zhang, 2 Laurie Burdett, 2 Belynda Hicks, 2 Sarangan Ravichandran, 3 Brian T. Luke, 3 Meredith Yeager, 2 Laura Fontaine, 4 Paula L. Hyland, 1 Alisa M. Goldstein, 1 NCI DCEG Cancer Sequencing Working Group, NCI DCEG Cancer Genomics Research Laboratory, Stephen J. Chanock, 5 Neil E. Caporaso, 1 Margaret A. Tucker, 6 and Lynn R. Goldin 1 1Genetic Epidemiology Branch, Division of Cancer Epidemiology and Genetics, National Cancer Institute, NIH, Bethesda, MD; 2Cancer Genomics Research Laboratory, Division of Cancer Epidemiology and Genetics, National Cancer Institute, NIH, Bethesda, MD; 3Ad - vanced Biomedical Computing Center, Leidos Biomedical Research Inc.; Frederick National Laboratory for Cancer Research, Frederick, MD; 4Westat, Inc., Rockville MD; 5Division of Cancer Epidemiology and Genetics, National Cancer Institute, NIH, Bethesda, MD; and 6Human Genetics Program, Division of Cancer Epidemiology and Genetics, National Cancer Institute, NIH, Bethesda, MD, USA ©2016 Ferrata Storti Foundation. This is an open-access paper. doi:10.3324/haematol.2015.135475 Received: August 19, 2015. Accepted: January 7, 2016. Pre-published: June 13, 2016. Correspondence: [email protected] Supplemental Author Information: NCI DCEG Cancer Sequencing Working Group: Mark H. Greene, Allan Hildesheim, Nan Hu, Maria Theresa Landi, Jennifer Loud, Phuong Mai, Lisa Mirabello, Lindsay Morton, Dilys Parry, Anand Pathak, Douglas R. Stewart, Philip R. Taylor, Geoffrey S. Tobias, Xiaohong R. Yang, Guoqin Yu NCI DCEG Cancer Genomics Research Laboratory: Salma Chowdhury, Michael Cullen, Casey Dagnall, Herbert Higson, Amy A. -

Mouse Cadm3 Knockout Project (CRISPR/Cas9)
https://www.alphaknockout.com Mouse Cadm3 Knockout Project (CRISPR/Cas9) Objective: To create a Cadm3 knockout Mouse model (C57BL/6N) by CRISPR/Cas-mediated genome engineering. Strategy summary: The Cadm3 gene (NCBI Reference Sequence: NM_053199 ; Ensembl: ENSMUSG00000005338 ) is located on Mouse chromosome 1. 9 exons are identified, with the ATG start codon in exon 1 and the TAG stop codon in exon 9 (Transcript: ENSMUST00000111220). Exon 2~6 will be selected as target site. Cas9 and gRNA will be co-injected into fertilized eggs for KO Mouse production. The pups will be genotyped by PCR followed by sequencing analysis. Note: Mice homozygous for a null allele exhibit delayed myelination. Other mice with ubiquitous conditional deletion of the gene do not display neurological abnormalities. Exon 2 starts from about 6.99% of the coding region. Exon 2~6 covers 58.42% of the coding region. The size of effective KO region: ~4894 bp. The KO region does not have any other known gene. Page 1 of 8 https://www.alphaknockout.com Overview of the Targeting Strategy Wildtype allele 5' gRNA region gRNA region 3' 1 2 3 4 5 6 9 Legends Exon of mouse Cadm3 Knockout region Page 2 of 8 https://www.alphaknockout.com Overview of the Dot Plot (up) Window size: 15 bp Forward Reverse Complement Sequence 12 Note: The 2000 bp section upstream of Exon 2 is aligned with itself to determine if there are tandem repeats. Tandem repeats are found in the dot plot matrix. The gRNA site is selected outside of these tandem repeats. Overview of the Dot Plot (down) Window size: 15 bp Forward Reverse Complement Sequence 12 Note: The 445 bp section downstream of Exon 6 is aligned with itself to determine if there are tandem repeats. -

Genetic and Functional Approaches to Understanding Autoimmune and Inflammatory Pathologies
University of Vermont ScholarWorks @ UVM Graduate College Dissertations and Theses Dissertations and Theses 2020 Genetic And Functional Approaches To Understanding Autoimmune And Inflammatory Pathologies Abbas Raza University of Vermont Follow this and additional works at: https://scholarworks.uvm.edu/graddis Part of the Genetics and Genomics Commons, Immunology and Infectious Disease Commons, and the Pathology Commons Recommended Citation Raza, Abbas, "Genetic And Functional Approaches To Understanding Autoimmune And Inflammatory Pathologies" (2020). Graduate College Dissertations and Theses. 1175. https://scholarworks.uvm.edu/graddis/1175 This Dissertation is brought to you for free and open access by the Dissertations and Theses at ScholarWorks @ UVM. It has been accepted for inclusion in Graduate College Dissertations and Theses by an authorized administrator of ScholarWorks @ UVM. For more information, please contact [email protected]. GENETIC AND FUNCTIONAL APPROACHES TO UNDERSTANDING AUTOIMMUNE AND INFLAMMATORY PATHOLOGIES A Dissertation Presented by Abbas Raza to The Faculty of the Graduate College of The University of Vermont In Partial Fulfillment of the Requirements for the Degree of Doctor of Philosophy Specializing in Cellular, Molecular, and Biomedical Sciences January, 2020 Defense Date: August 30, 2019 Dissertation Examination Committee: Cory Teuscher, Ph.D., Advisor Jonathan Boyson, Ph.D., Chairperson Matthew Poynter, Ph.D. Ralph Budd, M.D. Dawei Li, Ph.D. Dimitry Krementsov, Ph.D. Cynthia J. Forehand, Ph.D., Dean of the Graduate College ABSTRACT Our understanding of genetic predisposition to inflammatory and autoimmune diseases has been enhanced by large scale quantitative trait loci (QTL) linkage mapping and genome-wide association studies (GWAS). However, the resolution and interpretation of QTL linkage mapping or GWAS findings are limited. -

Peripheral Nerve Single-Cell Analysis Identifies Mesenchymal Ligands That Promote Axonal Growth
Research Article: New Research Development Peripheral Nerve Single-Cell Analysis Identifies Mesenchymal Ligands that Promote Axonal Growth Jeremy S. Toma,1 Konstantina Karamboulas,1,ª Matthew J. Carr,1,2,ª Adelaida Kolaj,1,3 Scott A. Yuzwa,1 Neemat Mahmud,1,3 Mekayla A. Storer,1 David R. Kaplan,1,2,4 and Freda D. Miller1,2,3,4 https://doi.org/10.1523/ENEURO.0066-20.2020 1Program in Neurosciences and Mental Health, Hospital for Sick Children, 555 University Avenue, Toronto, Ontario M5G 1X8, Canada, 2Institute of Medical Sciences University of Toronto, Toronto, Ontario M5G 1A8, Canada, 3Department of Physiology, University of Toronto, Toronto, Ontario M5G 1A8, Canada, and 4Department of Molecular Genetics, University of Toronto, Toronto, Ontario M5G 1A8, Canada Abstract Peripheral nerves provide a supportive growth environment for developing and regenerating axons and are es- sential for maintenance and repair of many non-neural tissues. This capacity has largely been ascribed to paracrine factors secreted by nerve-resident Schwann cells. Here, we used single-cell transcriptional profiling to identify ligands made by different injured rodent nerve cell types and have combined this with cell-surface mass spectrometry to computationally model potential paracrine interactions with peripheral neurons. These analyses show that peripheral nerves make many ligands predicted to act on peripheral and CNS neurons, in- cluding known and previously uncharacterized ligands. While Schwann cells are an important ligand source within injured nerves, more than half of the predicted ligands are made by nerve-resident mesenchymal cells, including the endoneurial cells most closely associated with peripheral axons. At least three of these mesen- chymal ligands, ANGPT1, CCL11, and VEGFC, promote growth when locally applied on sympathetic axons. -

F IDENTIFICATION of AMYOTROPHIC LATERAL
IDENTIFICATION OF AMYOTROPHIC LATERAL SCLEROSIS DISEASE MECHANISMS BY CEREBROSPINAL FLUID PROTEOMIC PROFILING by Mahlon Angus Collins B.S. in Neuroscience/Psychology, Central Michigan University, 2006 M.S. in Experimental Psychology, Central Michigan University, 2009 Submitted to the Graduate Faculty of the School of Medicine in partial fulfillment of the requirements for the degree of PhD in Neurobiology University of Pittsburgh 2016 f UNIVERSITY OF PITTSBURGH SCHOOL OF MEDICINE This dissertation was presented by Mahlon Angus Collins It was defended on April 14, 2016 and approved by Carey Balaban, PhD, Professor Michael Gold, PhD, Professor Teresa Hastings, PhD, Associate Professor Janice Robertson, PhD, Associate Professor Rebecca Seal, PhD, Assistant Professor Dissertation Advisor: Robert Bowser, PhD, Adjunct Professor ii Copyright © by Mahlon A. Collins 2016 iii IDENTIFICATION OF AMYOTROPHIC LATERAL SCLEROSIS DISEASE MECHANISMS BY CEREBROSPINAL FLUID PROTEOMIC PROFILING Mahlon Angus Collins, PhD University of Pittsburgh, 2016 Amyotrophic lateral sclerosis (ALS) is the most common form of adult-onset motor neuron disease. Heterogeneity in clinical, genetic, and pathological features of ALS suggest the disease is a spectrum of disorders each resulting in motor neuron degeneration. Molecular profiling of ALS patients is, therefore, a useful means of characterizing and stratifying the ALS population. To this end, mass spectrometric proteomic profiling was performed on cerebrospinal fluid (CSF) from ALS, healthy control (HC), and other -
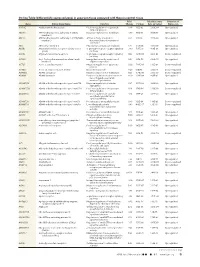
Differentially Expressed Genes in Aneurysm Tissue Compared With
On-line Table: Differentially expressed genes in aneurysm tissue compared with those in control tissue Fold False Discovery Direction of Gene Entrez Gene Name Function Change P Value Rate (q Value) Expression AADAC Arylacetamide deacetylase Positive regulation of triglyceride 4.46 1.33E-05 2.60E-04 Up-regulated catabolic process ABCA6 ATP-binding cassette, subfamily A (ABC1), Integral component of membrane 3.79 9.15E-14 8.88E-12 Up-regulated member 6 ABCC3 ATP-binding cassette, subfamily C (CFTR/MRP), ATPase activity, coupled to 6.63 1.21E-10 7.33E-09 Up-regulated member 3 transmembrane movement of substances ABI3 ABI family, member 3 Peptidyl-tyrosine phosphorylation 6.47 2.47E-05 4.56E-04 Up-regulated ACKR1 Atypical chemokine receptor 1 (Duffy blood G-protein–coupled receptor signaling 3.80 7.95E-10 4.18E-08 Up-regulated group) pathway ACKR2 Atypical chemokine receptor 2 G-protein–coupled receptor signaling 0.42 3.29E-04 4.41E-03 Down-regulated pathway ACSM1 Acyl-CoA synthetase medium-chain family Energy derivation by oxidation of 9.87 1.70E-08 6.52E-07 Up-regulated member 1 organic compounds ACTC1 Actin, ␣, cardiac muscle 1 Negative regulation of apoptotic 0.30 7.96E-06 1.65E-04 Down-regulated process ACTG2 Actin, ␥2, smooth muscle, enteric Blood microparticle 0.29 1.61E-16 2.36E-14 Down-regulated ADAM33 ADAM domain 33 Integral component of membrane 0.23 9.74E-09 3.95E-07 Down-regulated ADAM8 ADAM domain 8 Positive regulation of tumor necrosis 4.69 2.93E-04 4.01E-03 Up-regulated factor (ligand) superfamily member 11 production ADAMTS18 -

Transcriptome Profiling Reveals the Complexity of Pirfenidone Effects in IPF
ERJ Express. Published on August 30, 2018 as doi: 10.1183/13993003.00564-2018 Early View Original article Transcriptome profiling reveals the complexity of pirfenidone effects in IPF Grazyna Kwapiszewska, Anna Gungl, Jochen Wilhelm, Leigh M. Marsh, Helene Thekkekara Puthenparampil, Katharina Sinn, Miroslava Didiasova, Walter Klepetko, Djuro Kosanovic, Ralph T. Schermuly, Lukasz Wujak, Benjamin Weiss, Liliana Schaefer, Marc Schneider, Michael Kreuter, Andrea Olschewski, Werner Seeger, Horst Olschewski, Malgorzata Wygrecka Please cite this article as: Kwapiszewska G, Gungl A, Wilhelm J, et al. Transcriptome profiling reveals the complexity of pirfenidone effects in IPF. Eur Respir J 2018; in press (https://doi.org/10.1183/13993003.00564-2018). This manuscript has recently been accepted for publication in the European Respiratory Journal. It is published here in its accepted form prior to copyediting and typesetting by our production team. After these production processes are complete and the authors have approved the resulting proofs, the article will move to the latest issue of the ERJ online. Copyright ©ERS 2018 Copyright 2018 by the European Respiratory Society. Transcriptome profiling reveals the complexity of pirfenidone effects in IPF Grazyna Kwapiszewska1,2, Anna Gungl2, Jochen Wilhelm3†, Leigh M. Marsh1, Helene Thekkekara Puthenparampil1, Katharina Sinn4, Miroslava Didiasova5, Walter Klepetko4, Djuro Kosanovic3, Ralph T. Schermuly3†, Lukasz Wujak5, Benjamin Weiss6, Liliana Schaefer7, Marc Schneider8†, Michael Kreuter8†, Andrea Olschewski1, -

Analysis of Gene Expression from Systemic Lupus Erythematosus Synovium Reveals Myeloid Cell‑Driven Pathogenesis of Lupus Arthritis Erika L
www.nature.com/scientificreports OPEN Analysis of gene expression from systemic lupus erythematosus synovium reveals myeloid cell‑driven pathogenesis of lupus arthritis Erika L. Hubbard1*, Michelle D. Catalina1,2, Sarah Heuer1,3, Prathyusha Bachali1, Robert Robl1, Nicholas S. Geraci1,4, Amrie C. Grammer1,5 & Peter E. Lipsky1,5 Arthritis is a common manifestation of systemic lupus erythematosus (SLE) yet understanding of the underlying pathogenic mechanisms remains incomplete. We, therefore, interrogated gene expression profles of SLE synovium to gain insight into the nature of lupus arthritis (LA), using osteoarthritis (OA) and rheumatoid arthritis (RA) as comparators. Knee synovia from SLE, OA, and RA patients were analyzed for diferentially expressed genes (DEGs) and also by Weighted Gene Co‑expression Network Analysis (WGCNA) to identify modules of highly co‑expressed genes. Genes upregulated and/or co‑expressed in LA revealed numerous immune/infammatory cells dominated by a myeloid phenotype, in which pathogenic macrophages, myeloid‑lineage cells, and their secreted products perpetuate infammation, whereas OA was characterized by fbroblasts and RA of lymphocytes. Genes governing trafcking of immune cells into the synovium by chemokines were identifed, but not in situ generation of germinal centers (GCs). Gene Set Variation Analysis (GSVA) confrmed activation of specifc immune cell types in LA. Numerous therapies were predicted to target LA, including TNF, NFκB, MAPK, and CDK inhibitors. Detailed gene expression analysis identifed a unique pattern of cellular components and physiologic pathways operative in LA, as well as drugs potentially able to target this common manifestation of SLE. Systemic lupus erythematosus (SLE) is a complex autoimmune disease in which loss of self-tolerance gives rise to pathogenic autoantibodies causing widespread infammation and tissue damage 1. -
Single-Cell Transcriptome Analysis Reveals Mesenchymal Stem Cells In
bioRxiv preprint doi: https://doi.org/10.1101/2021.09.02.458742; this version posted September 3, 2021. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY 4.0 International license. 1 Single-cell transcriptome analysis reveals mesenchymal stem cells 2 in cavernous hemangioma 3 Fulong Ji1$, Yong Liu2$, Jinsong Shi3$, Chunxiang Liu1, Siqi Fu1 4 Heng Wang1, Bingbing Ren1, Dong Mi4, Shan Gao2*, Daqing Sun1* 5 1 Department of Paediatric Surgery, Tianjin Medical University General Hospital, Tianjin 300052, P.R. China. 6 China. 7 2 College of Life Sciences, Nankai University, Tianjin, Tianjin 300071, P.R. China; 8 3 National Clinical Research Center of Kidney Disease, Jinling Hospital, Nanjing University School of 9 Medicine, Nanjing, Jiangsu 210016, P.R. China; 10 4 School of Mathematical Sciences, Nankai University, Tianjin, Tianjin 300071, P.R. China; 11 12 13 $ These authors contributed equally to this paper. 14 * Corresponding authors. 15 SG:[email protected] 16 DS:[email protected] 17 bioRxiv preprint doi: https://doi.org/10.1101/2021.09.02.458742; this version posted September 3, 2021. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY 4.0 International license. 18 Abstract 19 A cavernous hemangioma, well-known as vascular malformation, is present at birth, grows 20 proportionately with the child, and does not undergo regression. -

Gene Modules Associated with Human Diseases Revealed by Network
bioRxiv preprint doi: https://doi.org/10.1101/598151; this version posted June 15, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. Gene modules associated with human diseases revealed by network analysis Shisong Ma1,2*, Jiazhen Gong1†, Wanzhu Zuo1†, Haiying Geng1, Yu Zhang1, Meng Wang1, Ershang Han1, Jing Peng1, Yuzhou Wang1, Yifan Wang1, Yanyan Chen1 1. Hefei National Laboratory for Physical Sciences at the Microscale, School of Life Sciences, University of Science and Technology of China, Hefei, Anhui 230027, China 2. School of Data Science, University of Science and Technology of China, Hefei, Anhui 230027, China * To whom correspondence should be addressed. Email: [email protected] † These authors contribute equally. 1 bioRxiv preprint doi: https://doi.org/10.1101/598151; this version posted June 15, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. ABSTRACT Despite many genes associated with human diseases have been identified, disease mechanisms often remain elusive due to the lack of understanding how disease genes are connected functionally at pathways level. Within biological networks, disease genes likely map to modules whose identification facilitates etiology studies but remains challenging. We describe a systematic approach to identify disease-associated gene modules. -

DNA Methylation Profiling in Recurrent Miscarriage
DNA methylation profiling in recurrent miscarriage Li Pi1,*, Zhaofeng Zhang1,*, Yan Gu2, Xinyue Wang1, Jianmei Wang2, Jianhua Xu1, Junwei Liu1, Xuan Zhang1 and Jing Du1 1 NHC Key Lab. of Reproduction Regulation (Shanghai Institute of Planned Parenthood Research), Medical School, Fudan University, Shanghai, China 2 The Second Hospital of Tianjin Medical University, Tianjin, China * These authors contributed equally to this work. ABSTRACT Recurrent miscarriage (RM) is a complex clinical problem. However, specific diag- nostic biomarkers and candidate regulatory targets have not yet been identified. To explore RM-related biological markers and processes, we performed a genome-wide DNA methylation analysis using the Illumina Infinium HumanMethylation450 array platform. Methylation variable positions and differentially methylated regions (DMRs) were selected using the Limma package in R language. Thereafter, gene ontology (GO) enrichment analysis and pathway enrichment analysis were performed on these DMRs. A total of 1,799 DMRs were filtered out between patients with RM and healthy pregnant women. The GO terms were mainly related to system development, plasma membrane part, and sequence-specific DNA binding, while the enriched pathways included cell adhesion molecules, type I diabetes mellitus, and ECM–receptor interactions. In addition, genes, including ABR, ALCAM, HLA-E, HLA-G, and ISG15, were obtained. These genes may be potential candidates for diagnostic biomarkers and possible regulatory targets in RM. We then detected the mRNA expression levels of the candidate genes. The mRNA expression levels of the candidate genes in the RM group were significantly higher than those in the control group. However, additional research is still required to confirm their potential roles in the occurrence of RM.