Comparative Genomics Reveals Specific Genetic Architectures In
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Downloaded 10/5/2021 9:43:05 AM
Chemical Science View Article Online EDGE ARTICLE View Journal | View Issue Aromatic side-chain flips orchestrate the conformational sampling of functional loops in Cite this: Chem. Sci.,2021,12,9318 † All publication charges for this article human histone deacetylase 8 have been paid for by the Royal Society a a bcd of Chemistry Vaibhav Kumar Shukla, ‡ Lucas Siemons, ‡ Francesco L. Gervasio and D. Flemming Hansen *a Human histone deacetylase 8 (HDAC8) is a key hydrolase in gene regulation and an important drug-target. High-resolution structures of HDAC8 in complex with substrates or inhibitors are available, which have provided insights into the bound state of HDAC8 and its function. Here, using long all-atom unbiased molecular dynamics simulations and Markov state modelling, we show a strong correlation between the conformation of aromatic side chains near the active site and opening and closing of the surrounding functional loops of HDAC8. We also investigated two mutants known to allosterically downregulate the enzymatic activity of HDAC8. Based on experimental data, we hypothesise that I19S-HDAC8 is unable to Received 6th April 2021 Creative Commons Attribution 3.0 Unported Licence. release the product, whereas both product release and substrate binding are impaired in the S39E- Accepted 27th May 2021 HDAC8 mutant. The presented results deliver detailed insights into the functional dynamics of HDAC8 DOI: 10.1039/d1sc01929e and provide a mechanism for the substantial downregulation caused by allosteric mutations, including rsc.li/chemical-science a disease causing one. Introduction II (HDAC-4, -5, -6, -7, -9, and HDAC10), and class III (SIRT1-7) have sequence similarity to yeast Rpd3, Hda1, and Sir2, Acetylation of lysine side chains occurs as a co-translation or respectively, whereas class IV (HDAC11) shares sequence simi- 2 This article is licensed under a post-translational modication of proteins and was rst iden- larity with both class I and II proteins. -
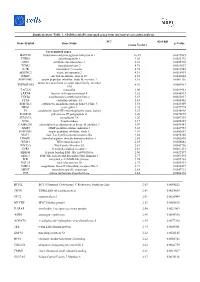
Supplementary Table 1. All Differentially Expressed Genes from Microarray Screening Analysis
Supplementary Table 1. All differentially expressed genes from microarray screening analysis. FCa (Gal-KD Gene Symbol Gene Name p-Value versus Vector) Up-regulated genes HAPLN1 hyaluronan and proteoglycan link protein 1 10.49 0.0027085 THBS1 thrombospondin 1 9.20 0.0022192 ODC1 ornithine decarboxylase 1 6.38 0.0055776 TGM2 transglutaminase 2 4.76 0.0015627 IL7R interleukin 7 receptor 4.75 0.0017245 SERINC2 serine incorporator 2 4.51 0.0014919 ITM2C integral membrane protein 2C 4.32 0.0044644 SERPINB7 serpin peptidase inhibitor, clade B, member 7 4.18 0.0081136 tumor necrosis factor receptor superfamily, member TNFRSF10D 4.01 0.0085561 10d TAGLN transgelin 3.90 0.0099963 LRRN4 leucine rich repeat neuronal 4 3.82 0.0046513 TGFB2 transforming growth factor beta 2 3.51 0.0035017 CPA4 carboxypeptidase A4 3.43 0.0008452 EPB41L3 erythrocyte membrane protein band 4.1-like 3 3.34 0.0025309 NRG1 neuregulin 1 3.28 0.0079724 F3 coagulation factor III (thromboplastin, tissue factor) 3.27 0.0038968 POLR3G polymerase III polypeptide G 3.26 0.0070675 SEMA7A semaphorin 7A 3.20 0.0087335 NT5E 5-nucleotidase 3.17 0.0036353 CAMK2N1 calmodulin-dependent protein kinase II inhibitor 1 3.07 0.0090141 TIMP3 TIMP metallopeptidase inhibitor 3 3.03 0.0047953 SERPINE1 serpin peptidase inhibitor, clade E 2.97 0.0053652 MALL mal, T-cell differentiation protein-like 2.88 0.0078205 DDAH1 dimethylarginine dimethylaminohydrolase 1 2.86 0.0002895 WDR3 WD repeat domain 3 2.85 0.0058842 WNT5A Wnt Family Member 5A 2.81 0.0043796 GPR1 G protein-coupled receptor 1 2.81 0.0021313 -

Functional Identification of a Novel Gene, Moae, for 3
www.nature.com/scientificreports OPEN Functional Identification of a Novel Gene, moaE, for 3-Succinoylpyridine Degradation in Received: 29 May 2015 Accepted: 28 July 2015 Pseudomonas putida S16 Published: 25 August 2015 Yi Jiang1,2, Hongzhi Tang1,2, Geng Wu1,2 & Ping Xu1,2 Microbial degradation of N-heterocyclic compounds, including xanthine, quinoline, nicotinate, and nicotine, frequently requires molybdenum hydroxylases. The intramolecular electron transfer chain of molybdenum hydroxylases consists of a molybdenum cofactor, two distinct [2Fe-2S] clusters, and flavin adenine dinucleotide. 3-Succinoylpyridine monooxygenase (Spm), responsible for the transformation from 3-succinoylpyridine to 6-hydroxy-3-succinoylpyridine, is a crucial enzyme in the pyrrolidine pathway of nicotine degradation in Pseudomonas. Our previous work revealed that the heterotrimeric enzyme (SpmA, SpmB, and SpmC) requires molybdopterin cytosine dinucleotide as a cofactor for their activities. In this study, we knocked out four genes, including PPS_1556, PPS_2936, PPS_4063, and PPS_4397, and found that a novel gene, PPS_4397 encoding moaE, is necessary for molybdopterin cytosine dinucleotide biosynthesis. Resting cell reactions of the moaE deletion mutant incubated with 3 g l−1 nicotine at 30 °C resulted in accumulation of 3-succinoylpyridine, and the strain complemented by the moaE gene regained the ability to convert 3-succinoylpyridine. In addition, reverse transcription-quantitative polymerase chain reaction analysis indicated that the transcriptional levels of the genes of moaE, spmA, and spmC of Pseudomonas putida S16 were distinctly higher when grown in nicotine medium than in glycerol medium. Large quantities of tobacco wastes containing high concentration of nicotine are produced during tobacco manufacturing yearly1. Nicotine is well known to be harmful to human health and can cross biological membranes and blood-brain barrier easily2. -

Biochemical Characterization and Comparison of Aspartylglucosaminidases Secreted in Venom of the Parasitoid Wasps Asobara Tabida and Leptopilina Heterotoma
RESEARCH ARTICLE Biochemical characterization and comparison of aspartylglucosaminidases secreted in venom of the parasitoid wasps Asobara tabida and Leptopilina heterotoma Quentin Coulette1, SeÂverine Lemauf2, Dominique Colinet2, Geneviève PreÂvost1, Caroline Anselme1, Marylène Poirie 2, Jean-Luc Gatti2* a1111111111 1 Unite ªEcologie et Dynamique des Systèmes AnthropiseÂsº (EDYSAN, FRE 3498 CNRS-UPJV), Universite de Picardie Jules Verne, Amiens, France, 2 Universite CoÃte d'Azur, INRA, CNRS, ISA, Sophia Antipolis, a1111111111 France a1111111111 a1111111111 * [email protected] a1111111111 Abstract Aspartylglucosaminidase (AGA) is a low-abundance intracellular enzyme that plays a key OPEN ACCESS role in the last stage of glycoproteins degradation, and whose deficiency leads to human Citation: Coulette Q, Lemauf S, Colinet D, PreÂvost aspartylglucosaminuria, a lysosomal storage disease. Surprisingly, high amounts of AGA- G, Anselme C, Poirie M, et al. (2017) Biochemical characterization and comparison of like proteins are secreted in the venom of two phylogenetically distant hymenopteran para- aspartylglucosaminidases secreted in venom of the sitoid wasp species, Asobara tabida (Braconidae) and Leptopilina heterotoma (Cynipidae). parasitoid wasps Asobara tabida and Leptopilina These venom AGAs have a similar domain organization as mammalian AGAs. They share heterotoma. PLoS ONE 12(7): e0181940. https:// with them key residues for autocatalysis and activity, and the mature - and -subunits also doi.org/10.1371/journal.pone.0181940 α β form an (αβ)2 structure in solution. Interestingly, only one of these AGAs subunits (α for Editor: Erjun Ling, Institute of Plant Physiology and AtAGA and for LhAGA) is glycosylated instead of the two subunits for lysosomal human Ecology Shanghai Institutes for Biological β Sciences, CHINA AGA (hAGA), and these glycosylations are partially resistant to PGNase F treatment. -

Biosynthesis of Natural Products Containing Β-Amino Acids
Natural Product Reports Biosynthesis of natural products containing β -amino acids Journal: Natural Product Reports Manuscript ID: NP-REV-01-2014-000007.R1 Article Type: Review Article Date Submitted by the Author: 21-Apr-2014 Complete List of Authors: Kudo, Fumitaka; Tokyo Institute Of Technology, Department of Chemistry Miyanaga, Akimasa; Tokyo Institute Of Technology, Department of Chemistry Eguchi, T; Tokyo Institute Of Technology, Department of Chemistry and Materials Science Page 1 of 20 Natural Product Reports NPR RSC Publishing REVIEW Biosynthesis of natural products containing βββ- amino acids Cite this: DOI: 10.1039/x0xx00000x Fumitaka Kudo, a Akimasa Miyanaga, a and Tadashi Eguchi *b Received 00th January 2014, We focus here on β-amino acids as components of complex natural products because the presence of β-amino acids Accepted 00th January 2014 produces structural diversity in natural products and provides characteristic architectures beyond that of ordinary DOI: 10.1039/x0xx00000x α-L-amino acids, thus generating significant and unique biological functions in nature. In this review, we first survey the known bioactive β-amino acid-containing natural products including nonribosomal peptides, www.rsc.org/ macrolactam polyketides, and nucleoside-β-amino acid hybrids. Next, the biosynthetic enzymes that form β-amino acids from α-amino acids and de novo synthesis of β-amino acids are summarized. Then, the mechanisms of β- amino acid incorporation into natural products are reviewed. Because it is anticipated that the rational swapping of the β-amino acid moieties with various side chains and stereochemistries by biosynthetic engineering should lead to the creation of novel architectures and bioactive compounds, the accumulation of knowledge regarding β- amino acid-containing natural product biosynthetic machinery could have a significant impact in this field. -

Active Site Tyrosine Is Essential for Amidohydrolase but Not for Esterase
Active site tyrosine is essential for amidohydrolase but not for esterase activity of a class 2 histone deacetylase-like bacterial enzyme Kristin Moreth, Daniel Riester, Christian Hildmann, René Hempel, Dennis Wegener, Andreas Schober, Andreas Schwienhorst To cite this version: Kristin Moreth, Daniel Riester, Christian Hildmann, René Hempel, Dennis Wegener, et al.. Ac- tive site tyrosine is essential for amidohydrolase but not for esterase activity of a class 2 histone deacetylase-like bacterial enzyme. Biochemical Journal, Portland Press, 2006, 401 (3), pp.659-665. 10.1042/BJ20061239. hal-00478649 HAL Id: hal-00478649 https://hal.archives-ouvertes.fr/hal-00478649 Submitted on 30 Apr 2010 HAL is a multi-disciplinary open access L’archive ouverte pluridisciplinaire HAL, est archive for the deposit and dissemination of sci- destinée au dépôt et à la diffusion de documents entific research documents, whether they are pub- scientifiques de niveau recherche, publiés ou non, lished or not. The documents may come from émanant des établissements d’enseignement et de teaching and research institutions in France or recherche français ou étrangers, des laboratoires abroad, or from public or private research centers. publics ou privés. Biochemical Journal Immediate Publication. Published on 12 Oct 2006 as manuscript BJ20061239 Active site tyrosine is essential for amidohydrolase but not for esterase activity of a class 2 histone deacetylase-like bacterial enzyme Kristin Moreth¶, Daniel Riester¶, Christian Hildmann¶, René Hempel¶, Dennis Wegener¶,§,‡, -

The Key to the Nornicotine Enantiomeric Composition in Tobacco Leaf
University of Kentucky UKnowledge Theses and Dissertations--Plant and Soil Sciences Plant and Soil Sciences 2012 ENANTIOSELECTIVE DEMETHYLATION: THE KEY TO THE NORNICOTINE ENANTIOMERIC COMPOSITION IN TOBACCO LEAF Bin Cai University of Kentucky, [email protected] Right click to open a feedback form in a new tab to let us know how this document benefits ou.y Recommended Citation Cai, Bin, "ENANTIOSELECTIVE DEMETHYLATION: THE KEY TO THE NORNICOTINE ENANTIOMERIC COMPOSITION IN TOBACCO LEAF" (2012). Theses and Dissertations--Plant and Soil Sciences. 5. https://uknowledge.uky.edu/pss_etds/5 This Doctoral Dissertation is brought to you for free and open access by the Plant and Soil Sciences at UKnowledge. It has been accepted for inclusion in Theses and Dissertations--Plant and Soil Sciences by an authorized administrator of UKnowledge. For more information, please contact [email protected]. STUDENT AGREEMENT: I represent that my thesis or dissertation and abstract are my original work. Proper attribution has been given to all outside sources. I understand that I am solely responsible for obtaining any needed copyright permissions. I have obtained and attached hereto needed written permission statements(s) from the owner(s) of each third-party copyrighted matter to be included in my work, allowing electronic distribution (if such use is not permitted by the fair use doctrine). I hereby grant to The University of Kentucky and its agents the non-exclusive license to archive and make accessible my work in whole or in part in all forms of media, now or hereafter known. I agree that the document mentioned above may be made available immediately for worldwide access unless a preapproved embargo applies. -

Characterization of Aspartylglucosaminidase Activation and Aspartylglucosaminuria Mutations
Arto Pennanen Publications of the National Public Health Institute A 1 / 2004 — INDOOR AIR POLLUTION AND HEALTH RISKS IN FINNISH ICE ARENAS RISKSINFINNISHICE AND HEALTH AIR POLLUTION INDOOR Jani Saarela CHARACTERIZATION OF ASPARTYLGLUCOSAMINIDASE ACTIVATION AND ASPARTYLGLUCOSAMINURIA MUTATIONS ISBN 951-740-485-9 ISSN 0359-3584 ISBN 951-740-486-7 (pdf) Department of Molecular Medicine, ISSN 1458-6290 (pdf) National Public Health Institute, Helsinki, Finland and http://www.ktl.fi /portal/suomi/julkaisut/julkaisusarjat/ Department of Medical Genetics, kansanterveyslaitoksen_julkaisuja_a/ University of Helsinki, Finland Kopijyvä Kuopio 2005 Helsinki 2004 PPennanen_kansi.inddennanen_kansi.indd 1 117.2.20057.2.2005 115:26:195:26:19 CHARACTERIZATION OF ASPARTYLGLUCOSAMINIDASE ACTIVATION AND ASPARTYLGLUCOSAMINURIA MUTATIONS Jani Saarela Department of Molecular Medicine, National Public Health Institute, Helsinki, Finland and Department of Medical Genetics, University of Helsinki, Finland Academic Dissertation To be publicly discussed with the permission of the Medical Faculty of the University of Helsinki, in the lecture room 3 of Biomedicum Helsinki, Haartmaninkatu 8, Helsinki, on January 30th, 2004, at 12 o’clock noon. Helsinki 2004 Supervised by Professor Leena Peltonen-Palotie National Public Health Institute and Department of Medical Genetics University of Helsinki, Helsinki, Finland Reviewed by Professor Ole Kristian Tollersrud and Docent Marc Baumann Department of Medical Biochemistry Protein Chemistry/Proteomics Unit University of Tromsoe and Neuroscience Research Program Tromsoe, Norway University of Helsinki, Helsinki, Finland To be publicly discussed with Professor Marja Makarow Institute of Biotechnology and Department of Applied Biochemistry and Molecular Biology University of Helsinki, Helsinki, Finland Julkaisija-Utgivare-Publisher Kansanterveyslaitos (KTL) Mannerheimintie 166 00300 Helsinki puh. vaihde 09-47441, felefax 09-4744 8408 Folkhälsoinstitutet Mannerheimvägen 166 00300, Helsinki tel. -

Eira Kelo | Kelo | 156 | Eira
View metadata, citation and similar papers at core.ac.uk brought to you by CORE dissertations provided by UEF Electronic Publications | 156 | Eira| 156 Kelo | Eira Kelo Catalytic and Therapeutic Characteristics of Human Aspartylglycosaminuria (AGU), Recombinant Glycosylasparaginase an inherited lysosomal storage Catalytic and Therapeutic Characteristics Human of Recombinant Glycosylasparaginase and Bacterial L-asparaginases Eira Kelo disease, is caused by the deficient and Bacterial L-asparaginases activity of a lysosomal enzyme glycosylasparaginase (GA). Catalytic and Therapeutic Loss of GA activity results in the Characteristics of Human accumulation of glycoasparagines in tissues leading to progressive Recombinant Glycosylasparaginase psychomotor retardation and a shortened life span. Currently there is no cure for and Bacterial L-asparaginases AGU. In this study, the effectiveness of enzyme replacement therapy (ERT) with human recombinant GA was evaluated in the mouse model of AGU. Furthermore, the enzymatic properties of GA and bacterial asparaginases were compared. This study reveals new therapeutic and catalytic properties of human GA and bacterial asparaginases. Publications of the University of Eastern Finland Dissertations in Health Sciences Publications of the University of Eastern Finland Dissertations in Health Sciences isbn 978-952-61-1045-5 EIRA KELO EIRA KELO Catalytic and therapeutic characteristics of Catalytic and therapeutic characteristics of human recombinant glycosylasparaginase human recombinant glycosylasparaginase -

Chemistry of Acetyl Transfer by Histone Modifying Enzymes: Structure, Mechanism and Implications for Effector Design
Oncogene (2007) 26, 5528–5540 & 2007 Nature Publishing Group All rights reserved 0950-9232/07 $30.00 www.nature.com/onc REVIEW Chemistry of acetyl transfer by histone modifying enzymes: structure, mechanism and implications for effector design SC Hodawadekar and R Marmorstein The Wistar Institute and The Department of Chemistry, University of Pennsylvania, Philadelphia, PA, USA The post-translational modification of histones plays an remodeling proteins that mobilize the histone proteins important role in chromatin regulation, a process that within chromatin (Varga-Weisz and Becker, 2006); insures the fidelity of gene expression and other DNA histone chaperone proteins that assemble, disassemble transactions. Of the enzymes that mediate post-transla- or replace variant histones within chromatin (Loyola tion modification, the histone acetyltransferase (HAT) and Almouzni, 2004); and post-translational modifica- and histone deacetylase (HDAC) proteins that add and tion enzymes that add or remove functional groups to or remove acetyl groups to and from target lysine residues from the histone proteins (Nightingale et al., 2006). within histones, respectively, have been the most exten- The post-translational modifications of histones sively studied at both the functional and structural levels. involve the addition or removal of acetyl, methyl or Not surprisingly, the aberrant activity of several of these phosphate groups as well as the reversible transfer of the enzymes have been implicated in human diseases such as ubiquitin and sumo proteins. -

Structural Analysis of Quinoline 2-Oxidoreductase from Pseudomonas Putida 86
Structural Analysis of Quinoline 2-Oxidoreductase from Pseudomonas putida 86 Structural and Biochemical Studies of Two Nus Family Proteins: NusB and NusA AR1-λN Complex NusA AR1-λN Complex NusB Quinoline 2-Oxidoreductase Irena Bonin Max-Planck-Institut für Biochemie Abteilung Strukturforschung D-82152 Martinsried, München 1 Max-Planck-Institut für Biochemie Abteilung Strukturforschung Structural Analysis of Quinoline 2-Oxidoreductase from Pseudomonas putida 86 Structural and Biochemical Studies of Two Nus Family Proteins: NusB and NusA AR1-λN Complex Irena Bonin Vollständiger Abdruck der von der Fakultät für Chemie der Technischen Universität München zur Erlangung des akademischen Grades eines Doktors der Naturwissenschaften genehmigten Dissertation. Vorsitzender: Univ.-Prof. Dr. St. J. Glaser Prüfer der Dissertation: 1. apl. Prof. Dr. Dr. h.c. R. Huber 2. Univ.-Prof. Dr. Dr. A. Bacher Die Dissertation wurde am 22.09.2004 bei der Technischen Universität München eingereicht und durch die Fakultät für Chemie am 11.11.2004 angenommen. 2 Ai miei genitori 3 ACKNOWLEDGEMENTS Acknowledgments The work, here presented, was carried out in the Abteilung für Strukturforschung at the Max-Planck-Institut für Biochemie, Martinsried bei München, under the supervision of Prof. Dr. Dr. h.c. Robert Huber. I would like to start thanking Prof. Dr. Robert Huber for giving me the opportunity to work in his department, his financial support, and for providing the necessary conditions for the success of my projects. To Dr. Markus C. Wahl who helped me in the initial struggle and promptly accepted to share his projects with me. A special thank goes to Dr. Holger Dobbek and Dr. -

Arthrobacter Nicotinovorans Pao1 Why Do We Need Its Proteome?
Arthrobacter nicotinovorans pAO1 Why do we need its proteome? Marius Mihășan, PhD. Faculty of Biology Alexandru Ioan Cuza University of Iaşi, Romania E-mail: [email protected] 4/2/17 Arthrobacter nicotinovorans - Why do we need its proteome? Slide 1 / 26 A bit about my home country Romania Alexandru Ioan Cuza University of Iasi Area: 92,043 sq mi Population: 20,121,641 Transylvania Flight history: Chemistry: Biology: Tr aian Vu ia Vintilă Ciocâlteu Victor Babeș first airplane to take off on its own power co-developed the Folin-Ciocalteu reagent more than 50 types of bacteria Physics: Costin Nenițescu Emil Racoviță Ştefan Procopiu many new classes of inorganic compounds founder of biospeleology the Bohr-Procopiu magneton Nicolae Teclu Emil Palade the Teclu burner Nobel Prize for contributions to cell biology 4/2/17 Arthrobacter nicotinovorans - Why do we need its proteome? Slide 2 / 26 Alexandru Ioan Cuza University of Iaşi Facts and figures ü The oldest Romanian university ü Diplomas recognized all over Europe Sciences Social Sciences & Humanities ü15 departments ü Biology ü Economics and Business Administration ü Chemistry ü History ü 93 Bachelor programs ü Computer Science ü Law ü Geography and Geology ü Letters ü Mathematics ü Orthodox Theology ü 176 Master programs ü Physics ü Philosophy and Social – Political Sciences ü Physical Education and Sports ü 26 PhD programs ü Psychology and Education Sciences ü Roman – Catholic Theology ü Center for European Studies 4/2/17 Arthrobacter nicotinovorans - Why do we need its proteome? Slide 3 / 26 Department of Biology Bulevardul Carol I nr. 20A, Iaşi, Romania, 700505 Tel.:+40(0)232201072 Fax: 40(0)232201472 www.bio.uaic.ro Founded in 1948 ü 758 students: • 20 PhD students ü46 full-time faculty members ü 16 technicians and administrative staff 4/2/17 Arthrobacter nicotinovorans - Why do we need its proteome? Slide 4 / 26 Programs of Study Biology BACHELOR Biochemistry semesters Environmental science Molecular genetics MASTER Biology Cellular and microbial biotech.