Are There Mutator Polymerases?
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Yeast DNA Polymerase Zeta ()Is Essential for Error-Free Replication Past Thymine Glycol
Downloaded from genesdev.cshlp.org on September 24, 2021 - Published by Cold Spring Harbor Laboratory Press Yeast DNA polymerase zeta ()is essential for error-free replication past thymine glycol Robert E. Johnson, Sung-Lim Yu, Satya Prakash, and Louise Prakash1 Sealy Center for Molecular Science, University of Texas Medical Branch at Galveston, Galveston, Texas 77555-1061, USA DNA polymerase zeta (Pol) promotes the mutagenic bypass of DNA lesions in eukaryotes. Genetic studies in Saccharomyces cerevisiae have indicated that relative to the contribution of other pathways, Pol makes only a modest contribution to lesion bypass. Intriguingly, however, disruption of the REV3 gene, which encodes the catalytic subunit of Pol, causes early embryonic lethality in mice. Here, we present genetic and biochemical evidence for the requirement of yeast Pol for predominantly error-free replication past thymine glycol (Tg), a DNA lesion formed frequently by free radical attack. These results raise the possibility that, as in yeast, in higher eukaryotes also, Pol makes a major contribution to the replicative bypass of Tgs as well as other lesions that block synthesis by replicative DNA polymerases. Such a preeminent role of Pol in lesion bypass would ensure that rapid cell divisions continue unabated during early embryonic development, thereby minimizing the generation of DNA strand breaks, chromosome aberrations, and the ensuing apoptotic response. [Keywords: DNApolymerase ; thymine glycol; translesion DNAsynthesis; Pol as an extender; error-free translesion DNAsynthesis by Pol ; yeast] Received October 4, 2002; revised version accepted October 31, 2002. Genetic studies in the yeast Saccharomyces cerevisiae et al. 2000b; Washington et al. 2000). Genetic studies in have indicated the requirement of Rad6–Rad18-depen- yeast have additionally indicated a role for Pol in the dent processes in promoting replication of damaged error-free bypass of cyclobutane pyrimidine dimers DNA. -

Table SI. Genes Upregulated ≥ 2-Fold by MIH 2.4Bl Treatment Affymetrix ID
Table SI. Genes upregulated 2-fold by MIH 2.4Bl treatment Fold UniGene ID Description Affymetrix ID Entrez Gene Change 1558048_x_at 28.84 Hs.551290 231597_x_at 17.02 Hs.720692 238825_at 10.19 93953 Hs.135167 acidic repeat containing (ACRC) 203821_at 9.82 1839 Hs.799 heparin binding EGF like growth factor (HBEGF) 1559509_at 9.41 Hs.656636 202957_at 9.06 3059 Hs.14601 hematopoietic cell-specific Lyn substrate 1 (HCLS1) 202388_at 8.11 5997 Hs.78944 regulator of G-protein signaling 2 (RGS2) 213649_at 7.9 6432 Hs.309090 serine and arginine rich splicing factor 7 (SRSF7) 228262_at 7.83 256714 Hs.127951 MAP7 domain containing 2 (MAP7D2) 38037_at 7.75 1839 Hs.799 heparin binding EGF like growth factor (HBEGF) 224549_x_at 7.6 202672_s_at 7.53 467 Hs.460 activating transcription factor 3 (ATF3) 243581_at 6.94 Hs.659284 239203_at 6.9 286006 Hs.396189 leucine rich single-pass membrane protein 1 (LSMEM1) 210800_at 6.7 1678 translocase of inner mitochondrial membrane 8 homolog A (yeast) (TIMM8A) 238956_at 6.48 1943 Hs.741510 ephrin A2 (EFNA2) 242918_at 6.22 4678 Hs.319334 nuclear autoantigenic sperm protein (NASP) 224254_x_at 6.06 243509_at 6 236832_at 5.89 221442 Hs.374076 adenylate cyclase 10, soluble pseudogene 1 (ADCY10P1) 234562_x_at 5.89 Hs.675414 214093_s_at 5.88 8880 Hs.567380; far upstream element binding protein 1 (FUBP1) Hs.707742 223774_at 5.59 677825 Hs.632377 small nucleolar RNA, H/ACA box 44 (SNORA44) 234723_x_at 5.48 Hs.677287 226419_s_at 5.41 6426 Hs.710026; serine and arginine rich splicing factor 1 (SRSF1) Hs.744140 228967_at 5.37 -

Ubiquitin and Ubiquitin-Like Proteins Are Essential Regulators of DNA Damage Bypass
cancers Review Ubiquitin and Ubiquitin-Like Proteins Are Essential Regulators of DNA Damage Bypass Nicole A. Wilkinson y, Katherine S. Mnuskin y, Nicholas W. Ashton * and Roger Woodgate * Laboratory of Genomic Integrity, National Institute of Child Health and Human Development, National Institutes of Health, 9800 Medical Center Drive, Rockville, MD 20850, USA; [email protected] (N.A.W.); [email protected] (K.S.M.) * Correspondence: [email protected] (N.W.A.); [email protected] (R.W.); Tel.: +1-301-435-1115 (N.W.A.); +1-301-435-0740 (R.W.) Co-first authors. y Received: 29 August 2020; Accepted: 29 September 2020; Published: 2 October 2020 Simple Summary: Ubiquitin and ubiquitin-like proteins are conjugated to many other proteins within the cell, to regulate their stability, localization, and activity. These modifications are essential for normal cellular function and the disruption of these processes contributes to numerous cancer types. In this review, we discuss how ubiquitin and ubiquitin-like proteins regulate the specialized replication pathways of DNA damage bypass, as well as how the disruption of these processes can contribute to cancer development. We also discuss how cancer cell survival relies on DNA damage bypass, and how targeting the regulation of these pathways by ubiquitin and ubiquitin-like proteins might be an effective strategy in anti-cancer therapies. Abstract: Many endogenous and exogenous factors can induce genomic instability in human cells, in the form of DNA damage and mutations, that predispose them to cancer development. Normal cells rely on DNA damage bypass pathways such as translesion synthesis (TLS) and template switching (TS) to replicate past lesions that might otherwise result in prolonged replication stress and lethal double-strand breaks (DSBs). -

Endogenous Overexpression of an Active Phosphorylated Form of DNA Polymerase Β Under Oxidative Stress in Trypanosoma Cruzi
RESEARCH ARTICLE Endogenous overexpression of an active phosphorylated form of DNA polymerase β under oxidative stress in Trypanosoma cruzi Diego A. Rojas1☯, Fabiola Urbina2☯, Sandra Moreira-Ramos2, Christian Castillo3, Ulrike Kemmerling3, Michel Lapier4, Juan Diego Maya4, Aldo Solari2, Edio Maldonado2¤* 1 Microbiology and Micology Program, ICBM, Faculty of Medicine, University of Chile, Santiago, Chile, 2 Cellular and Molecular Biology Program, ICBM, Faculty of Medicine, University of Chile, Santiago, Chile, 3 Anatomy and Developmental Biology Program, ICBM, Faculty of Medicine, University of Chile, Santiago, a1111111111 Chile, 4 Molecular and Clinical Pharmacology Program, ICBM, Faculty of Medicine, University of Chile, a1111111111 Santiago, Chile a1111111111 a1111111111 ☯ These authors contributed equally to this work. a1111111111 ¤ Current address: Independencia, Santiago, Chile * [email protected] Abstract OPEN ACCESS Trypanosoma cruzi is exposed during its life to exogenous and endogenous oxidative Citation: Rojas DA, Urbina F, Moreira-Ramos S, Castillo C, Kemmerling U, Lapier M, et al. (2018) stress, leading to damage of several macromolecules such as DNA. There are many DNA Endogenous overexpression of an active repair pathways in the nucleus and mitochondria (kinetoplast), where specific protein com- phosphorylated form of DNA polymerase β under plexes detect and eliminate damage to DNA. One group of these proteins is the DNA poly- oxidative stress in Trypanosoma cruzi. PLoS Negl Trop Dis 12(2): e0006220. https://doi.org/10.1371/ merases. In particular, Tc DNA polymerase β participates in kinetoplast DNA replication and journal.pntd.0006220 repair. However, the mechanisms which control its expression under oxidative stress are Editor: Joachim Clos, Bernhard Nocht Institute for still unknown. -
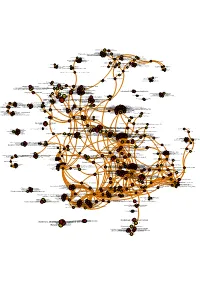
Mcf7specificall
Glycoprotein hormones alpha chain Hormone ligand-binding receptors Glycoprotein hormones Mineralocorticoid biosynthesis TFAP2 (AP-2) family regulates transcriptionAndrogenThyroid of biosynthesis growth stimulating factors hormone and their receptor receptors Reactions specificThyroxine to the complex biosynthesis N-glycan synthesis pathway Multidrug resistance-associated proteinGlucagon-like 5 peptide 1 receptor 78 kDa glucose-regulated protein Hyaluronan biosynthesis and export Glucagon-type ligand receptors Platelet degranulation G alpha (s)ABC-family signalling proteinsevents mediated transport PERKATF6 regulates(ATF6-alpha) gene activates expression chaperones Guanine nucleotide-binding protein G(s), subunit alpha Regulation of HSF1-mediated heat shock response Glucagon-like Peptide-1 (GLP1) regulates insulin secretion ATF6 (ATF6-alpha)IRE1alpha activates activates chaperone chaperones genes Parathyroid hormone receptor Serotonin (5-HT)Prostacyclin receptorG alpha signalling (z) signalling through events prostacyclin receptor Ras-related protein Rab-9A G alpha (q) signalling events GlucagonHedgehog signaling 'off' in metabolic state regulation PKA activation in glucagon signalling RAB geranylgeranylation Ubiquitin carboxyl-terminal hydrolase 1 Vasopressin regulates renal water homeostasis via Aquaporins Rap guanine nucleotide exchange factor 3 Class B/2 (Secretin Neuropeptidefamily receptors) S receptor Rap guanine nucleotide exchange factor 4 RetrogradeRAB transportGEFs exchange at the Trans-Golgi-Network GTP for GDP on RABs Serotonin -

Download Tool
by Submitted in partial satisfaction of the requirements for degree of in in the GRADUATE DIVISION of the UNIVERSITY OF CALIFORNIA, SAN FRANCISCO Approved: ______________________________________________________________________________ Chair ______________________________________________________________________________ ______________________________________________________________________________ ______________________________________________________________________________ ______________________________________________________________________________ Committee Members Copyright 2019 by Adolfo Cuesta ii Acknowledgements For me, completing a doctoral dissertation was a huge undertaking that was only possible with the support of many people along the way. First, I would like to thank my PhD advisor, Jack Taunton. He always gave me the space to pursue my own ideas and interests, while providing thoughtful guidance. Nearly every aspect of this project required a technique that was completely new to me. He trusted that I was up to the challenge, supported me throughout, helped me find outside resources when necessary. I remain impressed with his voracious appetite for the literature, and ability to recall some of the most subtle, yet most important details in a paper. Most of all, I am thankful that Jack has always been so generous with his time, both in person, and remotely. I’ve enjoyed our many conversations and hope that they will continue. I’d also like to thank my thesis committee, Kevan Shokat and David Agard for their valuable support, insight, and encouragement throughout this project. My lab mates in the Taunton lab made this such a pleasant experience, even on the days when things weren’t working well. I worked very closely with Tangpo Yang on the mass spectrometry aspects of this project. Xiaobo Wan taught me almost everything I know about protein crystallography. Thank you as well to Geoff Smith, Jordan Carelli, Pat Sharp, Yazmin Carassco, Keely Oltion, Nicole Wenzell, Haoyuan Wang, Steve Sethofer, and Shyam Krishnan, Shawn Ouyang and Qian Zhao. -

WO 2015/048577 A2 April 2015 (02.04.2015) W P O P C T
(12) INTERNATIONAL APPLICATION PUBLISHED UNDER THE PATENT COOPERATION TREATY (PCT) (19) World Intellectual Property Organization International Bureau (10) International Publication Number (43) International Publication Date WO 2015/048577 A2 April 2015 (02.04.2015) W P O P C T (51) International Patent Classification: (81) Designated States (unless otherwise indicated, for every A61K 48/00 (2006.01) kind of national protection available): AE, AG, AL, AM, AO, AT, AU, AZ, BA, BB, BG, BH, BN, BR, BW, BY, (21) International Application Number: BZ, CA, CH, CL, CN, CO, CR, CU, CZ, DE, DK, DM, PCT/US20 14/057905 DO, DZ, EC, EE, EG, ES, FI, GB, GD, GE, GH, GM, GT, (22) International Filing Date: HN, HR, HU, ID, IL, IN, IR, IS, JP, KE, KG, KN, KP, KR, 26 September 2014 (26.09.2014) KZ, LA, LC, LK, LR, LS, LU, LY, MA, MD, ME, MG, MK, MN, MW, MX, MY, MZ, NA, NG, NI, NO, NZ, OM, (25) Filing Language: English PA, PE, PG, PH, PL, PT, QA, RO, RS, RU, RW, SA, SC, (26) Publication Language: English SD, SE, SG, SK, SL, SM, ST, SV, SY, TH, TJ, TM, TN, TR, TT, TZ, UA, UG, US, UZ, VC, VN, ZA, ZM, ZW. (30) Priority Data: 61/883,925 27 September 2013 (27.09.2013) US (84) Designated States (unless otherwise indicated, for every 61/898,043 31 October 2013 (3 1. 10.2013) US kind of regional protection available): ARIPO (BW, GH, GM, KE, LR, LS, MW, MZ, NA, RW, SD, SL, ST, SZ, (71) Applicant: EDITAS MEDICINE, INC. -

Ablation of XP-V Gene Causes Adipose Tissue Senescence and Metabolic Abnormalities
Ablation of XP-V gene causes adipose tissue PNAS PLUS senescence and metabolic abnormalities Yih-Wen Chena, Robert A. Harrisb, Zafer Hatahetc, and Kai-ming Choua,1 aDepartment of Pharmacology and Toxicology, Indiana University School of Medicine, Indianapolis, IN 46202; bRichard Roudebush Veterans Affairs Medical Center and the Department of Biochemistry and Molecular Biology, Indiana University School of Medicine, Indianapolis, IN 46202; and cDepartment of Biological and Physical Sciences, Northwestern State University of Louisiana, Natchitoches, LA 71497 Edited by James E. Cleaver, University of California, San Francisco, CA, and approved June 26, 2015 (received for review April 12, 2015) Obesity and the metabolic syndrome have evolved to be major DNA polymerase η (pol η) is a specialized lesion bypass poly- health issues throughout the world. Whether loss of genome merase that faithfully replicates across UV-induced cyclobutane integrity contributes to this epidemic is an open question. DNA pyrimidine dimers (9) to rescue stalled DNA replication forks polymerase η (pol η), encoded by the xeroderma pigmentosum from potential breakages and mutations. Defects in the gene (XP-V) gene, plays an essential role in preventing cutaneous cancer encoding pol η produce a variant form of the autosomal recessive caused by UV radiation-induced DNA damage. Herein, we demon- disease xeroderma pigmentosum (XP-V) (9). Patients with XP-V strate that pol η deficiency in mice (pol η−/−) causes obesity with are highly sensitive to sunlight and prone to cutaneous cancer (9). visceral fat accumulation, hepatic steatosis, hyperleptinemia, In addition to skin, pol η is expressed in most tissues (10). The hyperinsulinemia, and glucose intolerance. -

Genome-Wide Investigation of Cellular Functions for Trna Nucleus
Genome-wide Investigation of Cellular Functions for tRNA Nucleus- Cytoplasm Trafficking in the Yeast Saccharomyces cerevisiae DISSERTATION Presented in Partial Fulfillment of the Requirements for the Degree Doctor of Philosophy in the Graduate School of The Ohio State University By Hui-Yi Chu Graduate Program in Molecular, Cellular and Developmental Biology The Ohio State University 2012 Dissertation Committee: Anita K. Hopper, Advisor Stephen Osmani Kurt Fredrick Jane Jackman Copyright by Hui-Yi Chu 2012 Abstract In eukaryotic cells tRNAs are transcribed in the nucleus and exported to the cytoplasm for their essential role in protein synthesis. This export event was thought to be unidirectional. Surprisingly, several lines of evidence showed that mature cytoplasmic tRNAs shuttle between nucleus and cytoplasm and their distribution is nutrient-dependent. This newly discovered tRNA retrograde process is conserved from yeast to vertebrates. Although how exactly the tRNA nuclear-cytoplasmic trafficking is regulated is still under investigation, previous studies identified several transporters involved in tRNA subcellular dynamics. At least three members of the β-importin family function in tRNA nuclear-cytoplasmic intracellular movement: (1) Los1 functions in both the tRNA primary export and re-export processes; (2) Mtr10, directly or indirectly, is responsible for the constitutive retrograde import of cytoplasmic tRNA to the nucleus; (3) Msn5 functions solely in the re-export process. In this thesis I focus on the physiological role(s) of the tRNA nuclear retrograde pathway. One possibility is that nuclear accumulation of cytoplasmic tRNA serves to modulate translation of particular transcripts. To test this hypothesis, I compared expression profiles from non-translating mRNAs and polyribosome-bound translating mRNAs collected from msn5Δ and mtr10Δ mutants and wild-type cells, in fed or acute amino acid starvation conditions. -

Inhibition of Mutagenic Translesion Synthesis: a Possible Strategy for Improving Chemotherapy?
VIEWPOINTS Inhibition of mutagenic translesion synthesis: A possible strategy for improving chemotherapy? Kinrin Yamanaka1☯, Nimrat Chatterjee1☯, Michael T. Hemann1,2, Graham C. Walker1* 1 Department of Biology, Massachusetts Institute of Technology, Cambridge, Massachusetts, United States of America, 2 Koch Institute, Massachusetts Institute of Technology, Cambridge, Massachusetts, United States of America ☯ These authors contributed equally to this work. * [email protected] a1111111111 Overview a1111111111 DNA damaging chemotherapy is the first line of treatment for certain cancers, but its long- a1111111111 term success is often marred by the eventual acquisition of chemoresistance. Other cancers a1111111111 a1111111111 cannot be treated because they are intrinsically resistant to such chemotherapy. These 2 types of resistance are coupled in the context of translesion synthesis (TLS), which is carried out by specialized TLS DNA polymerases that can replicate past DNA lesions but in a lower fidelity manner. First, TLS DNA polymerases permit the bypass of modified DNA bases during DNA synthesis, thereby allowing proliferation to continue in the presence of chemotherapy, an issue OPEN ACCESS of particular relevance to intrinsic drug resistance. Second, mistakes introduced by TLS poly- Citation: Yamanaka K, Chatterjee N, Hemann MT, merases copying over DNA lesions introduced during the chemotherapy lead to mutations Walker GC (2017) Inhibition of mutagenic that contribute to acquired resistance. These dual functions of mutagenic TLS polymerases translesion synthesis: A possible strategy for with respect to chemoresistance make these proteins very promising targets for adjuvant ther- improving chemotherapy? PLoS Genet 13(8): apy. The major branch of mutagenic TLS requires REV1, a Y family DNA polymerase that e1006842. https://doi.org/10.1371/journal. -

Identification of Novel DNA-Damage Tolerance Genes Reveals Regulation
ARTICLE Received 22 Apr 2014 | Accepted 1 Oct 2014 | Published 25 Nov 2014 DOI: 10.1038/ncomms6437 OPEN Identification of novel DNA-damage tolerance genes reveals regulation of translesion DNA synthesis by nucleophosmin Omer Ziv1, Amit Zeisel2, Nataly Mirlas-Neisberg1, Umakanta Swain1, Reinat Nevo1, Nir Ben-Chetrit3, Maria Paola Martelli4, Roberta Rossi4, Stefan Schiesser5, Christine E. Canman6, Thomas Carell5, Nicholas E. Geacintov7, Brunangelo Falini4, Eytan Domany2 & Zvi Livneh1 Cells cope with replication-blocking lesions via translesion DNA synthesis (TLS). TLS is carried out by low-fidelity DNA polymerases that replicate across lesions, thereby pre- venting genome instability at the cost of increased point mutations. Here we perform a two- stage siRNA-based functional screen for mammalian TLS genes and identify 17 validated TLS genes. One of the genes, NPM1, is frequently mutated in acute myeloid leukaemia (AML). We show that NPM1 (nucleophosmin) regulates TLS via interaction with the catalytic core of DNA polymerase-Z (polZ), and that NPM1 deficiency causes a TLS defect due to protea- somal degradation of polZ. Moreover, the prevalent NPM1c þ mutation that causes NPM1 mislocalization in B30% of AML patients results in excessive degradation of polZ. These results establish the role of NPM1 as a key TLS regulator, and suggest a mechanism for the better prognosis of AML patients carrying mutations in NPM1. 1 Department of Biological Chemistry, Weizmann Institute of Science, Rehovot 76100, Israel. 2 Department of Physics of Complex Systems, Weizmann Institute of Science, Rehovot 76100, Israel. 3 Department of Biological Regulation, Weizmann Institute of Science, Rehovot 76100, Israel. 4 Institute of Hematology, University of Perugia, Perugia 06100, Italy. -
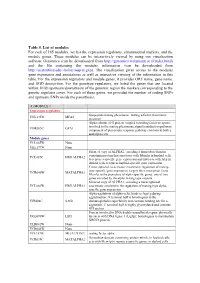
Table 5. List of Modules for Each of 165 Modules, We List the Expression Regulators, Chromosomal Markers, and the Module Genes
Table 5. List of modules For each of 165 modules, we list the expression regulators, chromosomal markers, and the module genes. These modules can be interactively viewed by using our visualization software Genomica (can be downloaded from http://genomica.weizmann.ac.il/index.html) and the file containing the modules information (can be downloaded from http://ai.stanford.edu/~koller/seqvar.gxp). The visualization gives access to the modules gene expression and annotations as well as interactive viewing of the information in this table. For the expression regulators and module genes, it provides ORF name, gene name, and SGD desciprtion. For the genotype regulators, we listed the genes that are located within 10 kb upstream/downstream of the genomic region the markers corresponding to the genetic regulator cover. For each of these genes, we provided the number of coding SNPs and upstream SNPs inside the parenthesis. (1) MODULE 1 Expression regulators lipopeptide mating pheromone; mating a-factor pheromone YNL145W MFA2 precursor Alpha subunit of G protein coupled to mating factor receptors, involved in the mating pheromone signal transduction pathway; YHR005C GPA1 component of pheromone response pathway common to both a and alpha cells Module genes YCL065W None YKL177W None Silenced copy of ALPHA2, encoding a homeobox-domain containing protein that associates with Mcm1p in haploid cells YCL067C HMLALPHA2 to repress a-specific gene expression and interacts with A1p in diploid cells to repress haploid-specific gene expression Transcriptional