The Tumor Microenvironment Shapes Lineage, Transcriptional, and Functional Diversity of Infiltrating Myeloid Cells
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Type I Interferons and the Development of Impaired Vascular Function and Repair in Human and Murine Lupus
Type I Interferons and the Development of Impaired Vascular Function and Repair in Human and Murine Lupus by Seth G Thacker A dissertation submitted in partial fulfillment of the requirements for the degree of Doctor of Philosophy (Immunology) in The University of Michigan 2011 Doctoral Committee: Associate Professor Mariana J. Kaplan, Chair Professor David A. Fox Professor Alisa E. Koch Professor Matthias Kretzler Professor Nicholas W. Lukacs Associate Professor Daniel T. Eitzman © Seth G Thacker 2011 Sharon, this work is dedicated to you. This achievement is as much yours as it is mine. Your support through all six years of this Ph.D. process has been incredible. You put up with my countless miscalculations on when I would finish experiments, and still managed to make me and our kids feel loved and special. Without you this would have no meaning. Sharon, you are the safe harbor in my life. ii Acknowledgments I have been exceptionally fortunate in my time here at the University of Michigan. I have been able to interact with so many supportive people over the years. I would like to express my thanks and admiration for my mentor. Mariana has taught me so much about writing, experimental design and being a successful scientist in general. I could never have made it here without her help. I would also like to thank Mike Denny. He had a hand in the beginning of all of my projects in one way or another, and was always quick and eager to help in whatever way he could. He really made my first year in the lab successful. -

Supplemental Materials ZNF281 Enhances Cardiac Reprogramming
Supplemental Materials ZNF281 enhances cardiac reprogramming by modulating cardiac and inflammatory gene expression Huanyu Zhou, Maria Gabriela Morales, Hisayuki Hashimoto, Matthew E. Dickson, Kunhua Song, Wenduo Ye, Min S. Kim, Hanspeter Niederstrasser, Zhaoning Wang, Beibei Chen, Bruce A. Posner, Rhonda Bassel-Duby and Eric N. Olson Supplemental Table 1; related to Figure 1. Supplemental Table 2; related to Figure 1. Supplemental Table 3; related to the “quantitative mRNA measurement” in Materials and Methods section. Supplemental Table 4; related to the “ChIP-seq, gene ontology and pathway analysis” and “RNA-seq” and gene ontology analysis” in Materials and Methods section. Supplemental Figure S1; related to Figure 1. Supplemental Figure S2; related to Figure 2. Supplemental Figure S3; related to Figure 3. Supplemental Figure S4; related to Figure 4. Supplemental Figure S5; related to Figure 6. Supplemental Table S1. Genes included in human retroviral ORF cDNA library. Gene Gene Gene Gene Gene Gene Gene Gene Symbol Symbol Symbol Symbol Symbol Symbol Symbol Symbol AATF BMP8A CEBPE CTNNB1 ESR2 GDF3 HOXA5 IL17D ADIPOQ BRPF1 CEBPG CUX1 ESRRA GDF6 HOXA6 IL17F ADNP BRPF3 CERS1 CX3CL1 ETS1 GIN1 HOXA7 IL18 AEBP1 BUD31 CERS2 CXCL10 ETS2 GLIS3 HOXB1 IL19 AFF4 C17ORF77 CERS4 CXCL11 ETV3 GMEB1 HOXB13 IL1A AHR C1QTNF4 CFL2 CXCL12 ETV7 GPBP1 HOXB5 IL1B AIMP1 C21ORF66 CHIA CXCL13 FAM3B GPER HOXB6 IL1F3 ALS2CR8 CBFA2T2 CIR1 CXCL14 FAM3D GPI HOXB7 IL1F5 ALX1 CBFA2T3 CITED1 CXCL16 FASLG GREM1 HOXB9 IL1F6 ARGFX CBFB CITED2 CXCL3 FBLN1 GREM2 HOXC4 IL1F7 -

Cellular and Molecular Signatures in the Disease Tissue of Early
Cellular and Molecular Signatures in the Disease Tissue of Early Rheumatoid Arthritis Stratify Clinical Response to csDMARD-Therapy and Predict Radiographic Progression Frances Humby1,* Myles Lewis1,* Nandhini Ramamoorthi2, Jason Hackney3, Michael Barnes1, Michele Bombardieri1, Francesca Setiadi2, Stephen Kelly1, Fabiola Bene1, Maria di Cicco1, Sudeh Riahi1, Vidalba Rocher-Ros1, Nora Ng1, Ilias Lazorou1, Rebecca E. Hands1, Desiree van der Heijde4, Robert Landewé5, Annette van der Helm-van Mil4, Alberto Cauli6, Iain B. McInnes7, Christopher D. Buckley8, Ernest Choy9, Peter Taylor10, Michael J. Townsend2 & Costantino Pitzalis1 1Centre for Experimental Medicine and Rheumatology, William Harvey Research Institute, Barts and The London School of Medicine and Dentistry, Queen Mary University of London, Charterhouse Square, London EC1M 6BQ, UK. Departments of 2Biomarker Discovery OMNI, 3Bioinformatics and Computational Biology, Genentech Research and Early Development, South San Francisco, California 94080 USA 4Department of Rheumatology, Leiden University Medical Center, The Netherlands 5Department of Clinical Immunology & Rheumatology, Amsterdam Rheumatology & Immunology Center, Amsterdam, The Netherlands 6Rheumatology Unit, Department of Medical Sciences, Policlinico of the University of Cagliari, Cagliari, Italy 7Institute of Infection, Immunity and Inflammation, University of Glasgow, Glasgow G12 8TA, UK 8Rheumatology Research Group, Institute of Inflammation and Ageing (IIA), University of Birmingham, Birmingham B15 2WB, UK 9Institute of -
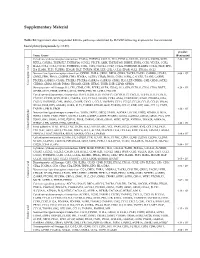
Supplementary Material
Supplementary Material Table S1: Significant downregulated KEGGs pathways identified by DAVID following exposure to five cinnamon- based phenylpropanoids (p < 0.05). p-value Term: Genes (Benjamini) Cytokine-cytokine receptor interaction: FASLG, TNFSF14, CXCL11, IL11, FLT3LG, CCL3L1, CCL3L3, CXCR6, XCR1, 2.43 × 105 RTEL1, CSF2RA, TNFRSF17, TNFRSF14, CCNL2, VEGFB, AMH, TNFRSF10B, INHBE, IFNB1, CCR3, VEGFA, CCR2, IL12A, CCL1, CCL3, CXCL5, TNFRSF25, CCR1, CSF1, CX3CL1, CCL7, CCL24, TNFRSF1B, IL12RB1, CCL21, FIGF, EPO, IL4, IL18R1, FLT1, TGFBR1, EDA2R, HGF, TNFSF8, KDR, LEP, GH2, CCL13, EPOR, XCL1, IFNA16, XCL2 Neuroactive ligand-receptor interaction: OPRM1, THRA, GRIK1, DRD2, GRIK2, TACR2, TACR1, GABRB1, LPAR4, 9.68 × 105 GRIK5, FPR1, PRSS1, GNRHR, FPR2, EDNRA, AGTR2, LTB4R, PRSS2, CNR1, S1PR4, CALCRL, TAAR5, GABRE, PTGER1, GABRG3, C5AR1, PTGER3, PTGER4, GABRA6, GABRA5, GRM1, PLG, LEP, CRHR1, GH2, GRM3, SSTR2, Chlorogenic acid Chlorogenic CHRM3, GRIA1, MC2R, P2RX2, TBXA2R, GHSR, HTR2C, TSHR, LHB, GLP1R, OPRD1 Hematopoietic cell lineage: IL4, CR1, CD8B, CSF1, FCER2, GYPA, ITGA2, IL11, GP9, FLT3LG, CD38, CD19, DNTT, 9.29 × 104 GP1BB, CD22, EPOR, CSF2RA, CD14, THPO, EPO, HLA-DRA, ITGA2B Cytokine-cytokine receptor interaction: IL6ST, IL21R, IL19, TNFSF15, CXCR3, IL15, CXCL11, TGFB1, IL11, FLT3LG, CXCL10, CCR10, XCR1, RTEL1, CSF2RA, IL21, CCNL2, VEGFB, CCR8, AMH, TNFRSF10C, IFNB1, PDGFRA, EDA, CXCL5, TNFRSF25, CSF1, IFNW1, CNTFR, CX3CL1, CCL5, TNFRSF4, CCL4, CCL27, CCL24, CCL25, CCL23, IFNA6, IFNA5, FIGF, EPO, AMHR2, IL2RA, FLT4, TGFBR2, EDA2R, -

Jimmunol.0903453.Full.Pdf
Antiviral Response Dictated by Choreographed Cascade of Transcription Factors This information is current as Elena Zaslavsky, Uri Hershberg, Jeremy Seto, Alissa M. of October 2, 2021. Pham, Susanna Marquez, Jamie L. Duke, James G. Wetmur, Benjamin R. tenOever, Stuart C. Sealfon and Steven H. Kleinstein J Immunol published online 17 February 2010 http://www.jimmunol.org/content/early/2010/02/17/jimmun ol.0903453 Downloaded from Supplementary http://www.jimmunol.org/content/suppl/2010/02/15/jimmunol.090345 Material 3.DC1 http://www.jimmunol.org/ Why The JI? Submit online. • Rapid Reviews! 30 days* from submission to initial decision • No Triage! Every submission reviewed by practicing scientists • Fast Publication! 4 weeks from acceptance to publication by guest on October 2, 2021 *average Subscription Information about subscribing to The Journal of Immunology is online at: http://jimmunol.org/subscription Permissions Submit copyright permission requests at: http://www.aai.org/About/Publications/JI/copyright.html Email Alerts Receive free email-alerts when new articles cite this article. Sign up at: http://jimmunol.org/alerts The Journal of Immunology is published twice each month by The American Association of Immunologists, Inc., 1451 Rockville Pike, Suite 650, Rockville, MD 20852 All rights reserved. Print ISSN: 0022-1767 Online ISSN: 1550-6606. Published February 17, 2010, doi:10.4049/jimmunol.0903453 The Journal of Immunology Antiviral Response Dictated by Choreographed Cascade of Transcription Factors Elena Zaslavsky,*,†,1 Uri Hershberg,‡,1 Jeremy Seto,*,† Alissa M. Pham,x Susanna Marquez,‡ Jamie L. Duke,{ James G. Wetmur,x Benjamin R. tenOever,x Stuart C. Sealfon,*,† and Steven H. -

WO 2018/067991 Al 12 April 2018 (12.04.2018) W !P O PCT
(12) INTERNATIONAL APPLICATION PUBLISHED UNDER THE PATENT COOPERATION TREATY (PCT) (19) World Intellectual Property Organization International Bureau (10) International Publication Number (43) International Publication Date WO 2018/067991 Al 12 April 2018 (12.04.2018) W !P O PCT (51) International Patent Classification: achusetts 021 15 (US). THE BROAD INSTITUTE, A61K 51/10 (2006.01) G01N 33/574 (2006.01) INC. [US/US]; 415 Main Street, Cambridge, Massachu C07K 14/705 (2006.01) A61K 47/68 (2017.01) setts 02142 (US). MASSACHUSETTS INSTITUTE OF G01N 33/53 (2006.01) TECHNOLOGY [US/US]; 77 Massachusetts Avenue, Cambridge, Massachusetts 02139 (US). (21) International Application Number: PCT/US2017/055625 (72) Inventors; and (71) Applicants: KUCHROO, Vijay K. [IN/US]; 30 Fairhaven (22) International Filing Date: Road, Newton, Massachusetts 02149 (US). ANDERSON, 06 October 2017 (06.10.2017) Ana Carrizosa [US/US]; 110 Cypress Street, Brookline, (25) Filing Language: English Massachusetts 02445 (US). MADI, Asaf [US/US]; c/o The Brigham and Women's Hospital, Inc., 75 Francis (26) Publication Language: English Street, Boston, Massachusetts 021 15 (US). CHIHARA, (30) Priority Data: Norio [US/US]; c/o The Brigham and Women's Hospital, 62/405,835 07 October 2016 (07.10.2016) US Inc., 75 Francis Street, Boston, Massachusetts 021 15 (US). REGEV, Aviv [US/US]; 15a Ellsworth Ave, Cambridge, (71) Applicants: THE BRIGHAM AND WOMEN'S HOSPI¬ Massachusetts 02139 (US). SINGER, Meromit [US/US]; TAL, INC. [US/US]; 75 Francis Street, Boston, Mass c/o The Broad Institute, Inc., 415 Main Street, Cambridge, (54) Title: MODULATION OF NOVEL IMMUNE CHECKPOINT TARGETS CD4 FIG. -

Diverse Functional Autoantibodies in Patients with COVID-19
medRxiv preprint doi: https://doi.org/10.1101/2020.12.10.20247205; this version posted December 11, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted medRxiv a license to display the preprint in perpetuity. It is made available under a CC-BY-NC-ND 4.0 International license . Diverse Functional Autoantibodies in Patients with COVID-19 Eric Y. Wang1,*, Tianyang Mao1,*, Jon Klein1,*, Yile Dai1,*, John D. Huck1, Feimei Liu1, Neil S. Zheng1, Ting Zhou1, Benjamin Israelow1, Patrick Wong1, Carolina Lucas1, Julio Silva1, Ji Eun Oh1, Eric Song1, Emily S. Perotti1, Suzanne Fischer1, Melissa Campbell5, John B. Fournier5, Anne L. Wyllie3, Chantal B. F. Vogels3, Isabel M. Ott3, Chaney C. Kalinich3, Mary E. Petrone3, Anne E. Watkins3, Yale IMPACT Team¶, Charles Dela Cruz4, Shelli F. Farhadian5, Wade L. Schulz6,7, Nathan D. Grubaugh3, Albert I. Ko3,5, Akiko Iwasaki1,3,8,#, Aaron M. Ring1,2,# 1 Department of Immunobiology, Yale School of Medicine, New Haven, CT, USA 2 Department of Pharmacology, Yale School of Medicine, New Haven, CT, USA 3 Department of Epidemiology of Microbial Diseases, Yale School of Public Health, New Haven, CT, USA 4 Department of Medicine, Section of Pulmonary and Critical Care Medicine, Yale School of Medicine, New Haven, CT, USA 5 Department of Internal Medicine (Infectious Diseases), Yale School of Medicine, New Haven, CT, USA 6 Department of Laboratory Medicine, Yale School of Medicine, New Haven, CT, USA 7 Center for Outcomes Research and Evaluation, Yale-New Haven Hospital, New Haven, CT, USA 8 Howard Hughes Medical Institute, Chevy Chase, MD, USA * These authors contributed equally to this work ¶ A list of authors and their affiliations appears at the end of the paper # Correspondence: [email protected] (A.M.R.); [email protected] (A.I.) 1 NOTE: This preprint reports new research that has not been certified by peer review and should not be used to guide clinical practice. -

Prevalent Homozygous Deletions of Type I Interferon and Defensin Genes in Human Cancers Associate with Immunotherapy Resistance
Author Manuscript Published OnlineFirst on April 4, 2018; DOI: 10.1158/1078-0432.CCR-17-3008 Author manuscripts have been peer reviewed and accepted for publication but have not yet been edited. Prevalent Homozygous Deletions of Type I Interferon and Defensin Genes in Human Cancers Associate with Immunotherapy Resistance Zhenqing Ye1; Haidong Dong2,3; Ying Li1; Tao Ma1,4; Haojie Huang4; Hon-Sing Leong4; Jeanette Eckel-Passow1; Jean-Pierre A. Kocher1; Han Liang5; LiguoWang1,4,* 1Division of Biomedical Statistics and Informatics, Department of Health Sciences, Mayo Clinic, Rochester, Minnesota 55905, United States of America 2Department of Immunology, College of Medicine, Mayo Clinic, Rochester, Minnesota 55905, United States of America 3Department of Urology, Mayo Clinic, Rochester, Minnesota 55905, United States of America 4Department of Biochemistry and Molecular Biology, Mayo Clinic, Rochester, Minnesota 55905, United States of America 5Department of Bioinformatics and Computational Biology, The University of Texas MD Anderson Cancer Center, Houston, Texas 77030, United States of America Running title: Pervasive Deletion of Interferon/Defensin in Human Cancers Keywords: Homozygous deletion, Type-I interferon, Defensin, immunotherapy resistance, cancer * Correspondence to: Liguo Wang, Ph.D. Associate Professor Division of Biomedical Statistics and Informatics, Mayo Clinic 200 1st St SW Rochester, MN 55905, USA Phone: +1-507-284-8728 Fax: +1-507-284-0745 Email: [email protected] The authors declare no potential conflicts of interest. 1 Downloaded from clincancerres.aacrjournals.org on September 29, 2021. © 2018 American Association for Cancer Research. Author Manuscript Published OnlineFirst on April 4, 2018; DOI: 10.1158/1078-0432.CCR-17-3008 Author manuscripts have been peer reviewed and accepted for publication but have not yet been edited. -

Gene Set Size Count Z-Score P-Value Q-Value List of Genes
Supplementary Data 2-1 Results from competitive pathway enrichment analysis based on genome-wide summary-level data of hsGWAS. Gene Set Size Count z-score p-value q-value List of Genes KEGG_ALLOGRAFT_ 38 36 12.6476 0.00E+00 0.00E+00 HLA-DRB5(4.11028); IL2(3.62922); HLA-E(3.27708); HLA-G(3.2749); HLA- REJECTION DQA2(3.16347); HLA-DRA(3.14994); HLA-DOB(3.11438); HLA-DMB(3.09917); HLA-DMA(3.09615); TNF(3.09414); HLA-DPB1(3.03922); HLA-DPA1(3.0355); HLA-A(3.02312); HLA-C(2.98835); HLA-DQB1(2.97025); HLA-B(2.95626); HLA- DQA1(2.94591); HLA-F(2.83254); HLA-DRB1(2.76738); HLA-DOA(2.72508); IFNG(1.74523); PRF1(1.46483); FASLG(1.10083); FAS(0.923795); HLA- DRB3(0.760622); CD80(0.436654); IL4(0.324564); CD40(0.271072); HLA- DRB4(0.0708609); IL12A(0.0685942); IL5(0.0659333); CD86(0.049911); CD28(- 0.16103); IL10(-0.165903); IL12B(-0.245281); GZMB(-0.268581); KEGG_GRAFT_VERSUS_HOS 42 37 12.6129 0.00E+00 0.00E+00 HLA-DRB5(4.11028); IL2(3.62922); HLA-E(3.27708); HLA-G(3.2749); HLA- T_DISEASE DQA2(3.16347); HLA-DRA(3.14994); HLA-DOB(3.11438); HLA-DMB(3.09917); HLA-DMA(3.09615); TNF(3.09414); HLA-DPB1(3.03922); HLA-DPA1(3.0355); HLA-A(3.02312); HLA-C(2.98835); HLA-DQB1(2.97025); HLA-B(2.95626); HLA- DQA1(2.94591); HLA-F(2.83254); HLA-DRB1(2.76738); HLA-DOA(2.72508); IL6(1.94139); IFNG(1.74523); PRF1(1.46483); FASLG(1.10083); FAS(0.923795); HLA-DRB3(0.760622); CD80(0.436654); IL1A(0.402186); KLRD1(0.29064); KIR3DL1(0.157683); HLA-DRB4(0.0708609); CD86(0.049911); CD28(-0.16103); GZMB(-0.268581); IL1B(-0.388308); KLRC1(-0.466394); KIR3DL2(-0.786806); KEGG_TYPE_I_DIABETES_ME -

David Thesis 2007
The Role of Nuclear Factor-B in -Cell Survival and Function David Liuwantara A submission to the University of New South Wales in candidature for the degree of Doctor of Philosophy Gene Therapy and Autoimmunity Research Program Department of Inflammation and Immunology Garvan Institute of Medical Research Darlinghurst, Sydney, Australia November 2007 Statement of Originality ‘I hereby declare that this submission is my own work and to the best of my knowledge it contains no materials previously published or written by another person, or substantial proportions of material which have been accepted for the award of any other degree or diploma at UNSW or any other educational institution, except where due acknowledgement is made in the thesis. Any contribution made to the research by others, with whom I have worked at UNSW or elsewhere, is explicitly acknowledged in the thesis. I also declare that the intellectual content of this thesis is the product of my own work, except to the extent that assistance from others in the project’s design and conception or in style, presentation and linguistic expression is acknowleged.’ Signed …………………………………………………………… In Christ alone will I glory Though I could pride myself in battles won For I've been blessed beyond measure And by His strength alone I overcome Oh I could stop and count successes Like diamonds in my hands But those trophies could not equal To the grace by which I stand In Christ alone will I glory For only by His grace I am redeemed And only His tender mercy Could reach beyond my weakness to my need And now I seek no greater honor Than just to know Him more And to count my gains But losses to the glory of my Lord In Christ alone I place my trust And find my glory in the power of the cross In every victory let it be said of me My source of strength, my source of hope is Christ alone In Christ Alone by: Don Koch and Shawn Craig Paragon Music Corp © 1990 BMG Music Pty. -

GEP Analysis Validates High Risk MDS and Acute Myeloid Leukemia Post
Guerenne et al. Journal of Hematology & Oncology (2016) 9:5 DOI 10.1186/s13045-016-0235-8 RESEARCH Open Access GEP analysis validates high risk MDS and acute myeloid leukemia post MDS mice models and highlights novel dysregulated pathways Laura Guerenne1,2, Stéphanie Beurlet1,2, Mohamed Said3, Petra Gorombei1,2, Carole Le Pogam1,2, Fabien Guidez1,2, Pierre de la Grange4, Nader Omidvar5, Valérie Vanneaux6, Ken Mills7, Ghulam J Mufti3, Laure Sarda-Mantel8,9, Maria Elena Noguera10, Marika Pla1,2,11, Pierre Fenaux1,2,10, Rose Ann Padua1,2,10†, Christine Chomienne1,2,10† and Patricia Krief1,2* Abstract Background: In spite of the recent discovery of genetic mutations in most myelodysplasic (MDS) patients, the pathophysiology of these disorders still remains poorly understood, and only few in vivo models are available to help unravel the disease. Methods: We performed global specific gene expression profiling and functional pathway analysis in purified Sca1+ cells of two MDS transgenic mouse models that mimic human high-risk MDS (HR-MDS) and acute myeloid leukemia (AML) post MDS, with NRASD12 and BCL2 transgenes under the control of different promoters MRP8NRASD12/tethBCL-2 or MRP8[NRASD12/hBCL-2], respectively. Results: Analysis of dysregulated genes that were unique to the diseased HR-MDS and AML post MDS mice and not their founder mice pointed first to pathways that had previously been reported in MDS patients, including DNA replication/damage/repair, cell cycle, apoptosis, immune responses, and canonical Wnt pathways, further validating these models at the gene expression level. Interestingly, pathways not previously reported in MDS were discovered. -

An Inflammatory Gene Signature Distinguishes Neurofibroma
www.nature.com/scientificreports OPEN An inflammatory gene signature distinguishes neurofibroma Schwann cells and macrophages Received: 26 May 2016 Accepted: 25 January 2017 from cells in the normal peripheral Published: 03 March 2017 nervous system Kwangmin Choi1, Kakajan Komurov1, Jonathan S. Fletcher1, Edwin Jousma1, Jose A. Cancelas1,2, Jianqiang Wu1 & Nancy Ratner1 Neurofibromas are benign peripheral nerve tumors driven byNF1 loss in Schwann cells (SCs). Macrophages are abundant in neurofibromas, and macrophage targeted interventions may have therapeutic potential in these tumors. We generated gene expression data from fluorescence- activated cell sorted (FACS) SCs and macrophages from wild-type and mutant nerve and neurofibroma to identify candidate pathways involved in SC-macrophage cross-talk. While in 1-month-old Nf1 mutant nerve neither SCs nor macrophages significantly differed from their normal counterparts, both macrophages and SCs showed significantly altered cytokine gene expression in neurofibromas. Computationally reconstructed SC-macrophage molecular networks were enriched for inflammation- associated pathways. We verified that neurofibroma SC conditioned medium contains macrophage chemo-attractants including colony stimulation factor 1 (CSF1). Network analysis confirmed previously implicated pathways and predict novel paracrine and autocrine loops involving cytokines, chemokines, and growth factors. Network analysis also predicted a central role for decreased type-I interferon signaling. We validated type-I interferon expression in neurofibroma by protein profiling, and show that treatment of neurofibroma-bearing mice with polyethylene glycolyated (PEGylated) type-I interferon- α2b reduces the expression of many cytokines overexpressed in neurofibroma. These studies reveal numerous potential targetable interactions between Nf1 mutant SCs and macrophages for further analyses. Neurofibromatosis type 1 (NF1) is one of the most common human monogenic disorders, affecting about 0.3% of the human population.