Genome-Wide Analyses of Repeat-Induced Point Mutations in the Ascomycota
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Molecular Identification of Fungi
Molecular Identification of Fungi Youssuf Gherbawy l Kerstin Voigt Editors Molecular Identification of Fungi Editors Prof. Dr. Youssuf Gherbawy Dr. Kerstin Voigt South Valley University University of Jena Faculty of Science School of Biology and Pharmacy Department of Botany Institute of Microbiology 83523 Qena, Egypt Neugasse 25 [email protected] 07743 Jena, Germany [email protected] ISBN 978-3-642-05041-1 e-ISBN 978-3-642-05042-8 DOI 10.1007/978-3-642-05042-8 Springer Heidelberg Dordrecht London New York Library of Congress Control Number: 2009938949 # Springer-Verlag Berlin Heidelberg 2010 This work is subject to copyright. All rights are reserved, whether the whole or part of the material is concerned, specifically the rights of translation, reprinting, reuse of illustrations, recitation, broadcasting, reproduction on microfilm or in any other way, and storage in data banks. Duplication of this publication or parts thereof is permitted only under the provisions of the German Copyright Law of September 9, 1965, in its current version, and permission for use must always be obtained from Springer. Violations are liable to prosecution under the German Copyright Law. The use of general descriptive names, registered names, trademarks, etc. in this publication does not imply, even in the absence of a specific statement, that such names are exempt from the relevant protective laws and regulations and therefore free for general use. Cover design: WMXDesign GmbH, Heidelberg, Germany, kindly supported by ‘leopardy.com’ Printed on acid-free paper Springer is part of Springer Science+Business Media (www.springer.com) Dedicated to Prof. Lajos Ferenczy (1930–2004) microbiologist, mycologist and member of the Hungarian Academy of Sciences, one of the most outstanding Hungarian biologists of the twentieth century Preface Fungi comprise a vast variety of microorganisms and are numerically among the most abundant eukaryotes on Earth’s biosphere. -

Thermophilic Fungi: Taxonomy and Biogeography
Journal of Agricultural Technology Thermophilic Fungi: Taxonomy and Biogeography Raj Kumar Salar1* and K.R. Aneja2 1Department of Biotechnology, Chaudhary Devi Lal University, Sirsa – 125 055, India 2Department of Microbiology, Kurukshetra University, Kurukshetra – 136 119, India Salar, R. K. and Aneja, K.R. (2007) Thermophilic Fungi: Taxonomy and Biogeography. Journal of Agricultural Technology 3(1): 77-107. A critical reappraisal of taxonomic status of known thermophilic fungi indicating their natural occurrence and methods of isolation and culture was undertaken. Altogether forty-two species of thermophilic fungi viz., five belonging to Zygomycetes, twenty-three to Ascomycetes and fourteen to Deuteromycetes (Anamorphic Fungi) are described. The taxa delt with are those most commonly cited in the literature of fundamental and applied work. Latest legal valid names for all the taxa have been used. A key for the identification of thermophilic fungi is given. Data on geographical distribution and habitat for each isolate is also provided. The specimens deposited at IMI bear IMI number/s. The document is a sound footing for future work of indentification and nomenclatural interests. To solve residual problems related to nomenclatural status, further taxonomic work is however needed. Key Words: Biodiversity, ecology, identification key, taxonomic description, status, thermophile Introduction Thermophilic fungi are a small assemblage in eukaryota that have a unique mechanism of growing at elevated temperature extending up to 60 to 62°C. During the last four decades many species of thermophilic fungi sporulating at 45oC have been reported. The species included in this account are only those which are thermophilic in the sense of Cooney and Emerson (1964). -

1Hjs Lichtarge Lab 2006
Pages 1–8 1hjs Evolutionary trace report by report maker April 15, 2010 4.3.1 Alistat 7 4.3.2 CE 7 4.3.3 DSSP 7 4.3.4 HSSP 7 4.3.5 LaTex 8 4.3.6 Muscle 8 4.3.7 Pymol 8 4.4 Note about ET Viewer 8 4.5 Citing this work 8 4.6 About report maker 8 4.7 Attachments 8 1 INTRODUCTION From the original Protein Data Bank entry (PDB id 1hjs): Title: Structure of two fungal beta-1,4-galactanases: searching for the basis for temperature and ph optimum. Compound: Mol id: 1; molecule: beta-1,4-galactanase; chain: a, b, c, d; ec: 3.2.1.89; engineered: yes; other details: 2-n-acetyl-beta-d- glucose(residue 601) linked to asn 111 in the four molecules Organism, scientific name: Thielavia Heterothallica; 1hjs contains a single unique chain 1hjsA (332 residues long) and CONTENTS its homologues 1hjsD, 1hjsC, and 1hjsB. 1 Introduction 1 2 CHAIN 1HJSA 2.1 P83692 overview 2 Chain 1hjsA 1 2.1 P83692 overview 1 From SwissProt, id P83692, 96% identical to 1hjsA: 2.2 Multiple sequence alignment for 1hjsA 1 Description: Arabinogalactan endo-1,4-beta-galactosidase (EC 2.3 Residue ranking in 1hjsA 1 3.2.1.89) (Endo-1,4- beta-galactanase) (Galactanase). 2.4 Top ranking residues in 1hjsA and their position on Organism, scientific name: Thielavia heterothallica (Mycelio- the structure 2 phthora thermophila). 2.4.1 Clustering of residues at 25% coverage. 2 Taxonomy: Eukaryota; Fungi; Ascomycota; Pezizomycotina; 2.4.2 Overlap with known functional surfaces at Sordariomycetes; Sordariomycetidae; Sordariales; Chaetomiaceae; 25% coverage. -

The Phylogeny of Plant and Animal Pathogens in the Ascomycota
Physiological and Molecular Plant Pathology (2001) 59, 165±187 doi:10.1006/pmpp.2001.0355, available online at http://www.idealibrary.com on MINI-REVIEW The phylogeny of plant and animal pathogens in the Ascomycota MARY L. BERBEE* Department of Botany, University of British Columbia, 6270 University Blvd, Vancouver, BC V6T 1Z4, Canada (Accepted for publication August 2001) What makes a fungus pathogenic? In this review, phylogenetic inference is used to speculate on the evolution of plant and animal pathogens in the fungal Phylum Ascomycota. A phylogeny is presented using 297 18S ribosomal DNA sequences from GenBank and it is shown that most known plant pathogens are concentrated in four classes in the Ascomycota. Animal pathogens are also concentrated, but in two ascomycete classes that contain few, if any, plant pathogens. Rather than appearing as a constant character of a class, the ability to cause disease in plants and animals was gained and lost repeatedly. The genes that code for some traits involved in pathogenicity or virulence have been cloned and characterized, and so the evolutionary relationships of a few of the genes for enzymes and toxins known to play roles in diseases were explored. In general, these genes are too narrowly distributed and too recent in origin to explain the broad patterns of origin of pathogens. Co-evolution could potentially be part of an explanation for phylogenetic patterns of pathogenesis. Robust phylogenies not only of the fungi, but also of host plants and animals are becoming available, allowing for critical analysis of the nature of co-evolutionary warfare. Host animals, particularly human hosts have had little obvious eect on fungal evolution and most cases of fungal disease in humans appear to represent an evolutionary dead end for the fungus. -

Fungal Planet Description Sheets: 400–468
Persoonia 36, 2016: 316– 458 www.ingentaconnect.com/content/nhn/pimj RESEARCH ARTICLE http://dx.doi.org/10.3767/003158516X692185 Fungal Planet description sheets: 400–468 P.W. Crous1,2, M.J. Wingfield3, D.M. Richardson4, J.J. Le Roux4, D. Strasberg5, J. Edwards6, F. Roets7, V. Hubka8, P.W.J. Taylor9, M. Heykoop10, M.P. Martín11, G. Moreno10, D.A. Sutton12, N.P. Wiederhold12, C.W. Barnes13, J.R. Carlavilla10, J. Gené14, A. Giraldo1,2, V. Guarnaccia1, J. Guarro14, M. Hernández-Restrepo1,2, M. Kolařík15, J.L. Manjón10, I.G. Pascoe6, E.S. Popov16, M. Sandoval-Denis14, J.H.C. Woudenberg1, K. Acharya17, A.V. Alexandrova18, P. Alvarado19, R.N. Barbosa20, I.G. Baseia21, R.A. Blanchette22, T. Boekhout3, T.I. Burgess23, J.F. Cano-Lira14, A. Čmoková8, R.A. Dimitrov24, M.Yu. Dyakov18, M. Dueñas11, A.K. Dutta17, F. Esteve- Raventós10, A.G. Fedosova16, J. Fournier25, P. Gamboa26, D.E. Gouliamova27, T. Grebenc28, M. Groenewald1, B. Hanse29, G.E.St.J. Hardy23, B.W. Held22, Ž. Jurjević30, T. Kaewgrajang31, K.P.D. Latha32, L. Lombard1, J.J. Luangsa-ard33, P. Lysková34, N. Mallátová35, P. Manimohan32, A.N. Miller36, M. Mirabolfathy37, O.V. Morozova16, M. Obodai38, N.T. Oliveira20, M.E. Ordóñez39, E.C. Otto22, S. Paloi17, S.W. Peterson40, C. Phosri41, J. Roux3, W.A. Salazar 39, A. Sánchez10, G.A. Sarria42, H.-D. Shin43, B.D.B. Silva21, G.A. Silva20, M.Th. Smith1, C.M. Souza-Motta44, A.M. Stchigel14, M.M. Stoilova-Disheva27, M.A. Sulzbacher 45, M.T. Telleria11, C. Toapanta46, J.M. Traba47, N. -

On a New Species of Chaetomidium, C. Vicugnae, with a Cephalothecoid
On a new species of Chaetomidium, C. vicugnae, with a cepha- lothecoid peridium and its relationships with Chaetomiaceae (Sordariales) Francesco DOVERI Abstract: a sample of vicuña dung from a Chilean coastal desert was submitted to the attention of the au- thor, who at first sight noticed the presence of different pyrenomycetes. several hairy cleistothecia particu- larly caught his attention and were subjected to a morphological study that proved them to belong to a new species of Chaetomidium. after mentioning the main features of Sordariales and Chaetomiaceae, the author describes in detail the macro-and microscopic characters of the new species Chaetomidium vicugnae Ascomycete.org, 10 (2) : 86–96 and compares it with all the other Chaetomidium spp. with a cephalothecoid peridium. The extensive dis- Mise en ligne le 22/04/2018 cussion focuses on the characterization and relationships of the genus Chaetomidium and Chaetomidium 10.25664/ART-0231 vicugnae within the complex family Chaetomiaceae. all collections of the related species are recorded and dung is regarded as the preferential substrate. Keys are provided to sexual morph genera of Chaetomiaceae and to Chaetomidium species with a cephalothecoid peridium. Keywords: ascomycota, coprophily, germination, homoplasy, morphology, peridial frame, systematics. Introduction zing the importance of a future systematic study of vicuña dung for a better knowledge of the generic relationships in this family. My studies on coprophilous ascomycetes (Doveri, 2004, 2011) al- lowed me to meet with several representatives of Sordariales Cha- Materials and methods def. ex D. Hawksw. & o.e. erikss., an order identifiable with the so called “pyrenomycetes” s.str., i.e. -

Coprophilous Ascomycetes with Passive Ascospore Liberation from Brazil
Phytotaxa 295 (2): 159–172 ISSN 1179-3155 (print edition) http://www.mapress.com/j/pt/ PHYTOTAXA Copyright © 2017 Magnolia Press Article ISSN 1179-3163 (online edition) https://doi.org/10.11646/phytotaxa.295.2.4 Coprophilous ascomycetes with passive ascospore liberation from Brazil ROGER FAGNER RIBEIRO MELO1*, LEONOR COSTA MAIA1 & ANDREW NICHOLAS MILLER2 1Universidade Federal de Pernambuco, Centro de Biociências, Departamento de Micologia, Av. da Engenharia, s/n, 50740‒600, Recife, Pernambuco, Brazil 2University of Illinois at Urbana-Champaign, Illinois Natural History Survey, 1816 South Oak Street, Champaign, IL 61820, USA Correspondence: [email protected] Abstract Ascomycetes with passive ascospore liberation fruiting on herbivore dung are discussed. A total of 270 samples of cattle, goat and horse dung were collected for 20 months along an edaphic and climatic gradient from the Atlantic Forest complex to the semi-arid Caatinga complex in Pernambuco, northeastern Brazil. Thirteen species were identified and described. Lophot- richus bartlettii and Kernia nitida were the most frequently recorded species. Corynascus sepedonium, Leuconeurospora pulcherrima, Melanospora damnosa, M. zamiae, Mycoarachis inversa, Zopfiella erostrata and Zopfiella longicaudata are reported for the first time in Brazil. Descriptions, a photographic plate and an identification key to the studied species, along with a table with key characters of the most common genera of coprophilous ascomycetes with passive ascospore liberation are provided. Key words: Ascomycota, coprophilous fungi, Microascales, non-ostiolate ascomycetes Introduction Coprophilous fungi form a collective group of saprobes able to live, feed and reproduce in dung, especially from herbivores (Webster 1970, Krug et al. 2004, Kirk et al. 2008). These fungi are associated with various animals (most notably mammals), domesticated or wild (Richardson 2001), presenting an array of morphologic and physiologic life strategies to efficiently exploit their substrate (Ingold 1961, Dix & Webster 1995, Kirschner et al. -

A Reevaluation of Predatory Orbiliaceous Fungi. II. a New Generic Concept
©Verlag Ferdinand Berger & Söhne Ges.m.b.H., Horn, Austria, download unter www.biologiezentrum.at A reevaluation of predatory orbiliaceous fungi. II. A new generic concept Markus Scholler1, Gregor Hagedorn2 & A. Rubner1 Fachrichtung Biologie, Mykologisches Labor, Universität Greifswald, Jahn-Str. 15, 17487 Greifswald, Germany 2Institut für Pflanzenvirologie, Mikrobiologie und Biologische Sicherheit, Biologische Bundesanstalt für Land- und Forstwirtschaft, Königin-Luise-Str. 19, 14195 Berlin, Germany M. Scholler, G. Hagedorn & A. Rubner (1999). A reevaluation of predatory orbiliaceous fungi. II. A new generic concept. - Sydowia 51(1): 89-113. A new genus concept is proposed for predatory anamorphic Orbiliaceae in which the trapping device is the main morphological criterion for the delimitation of the genera. Molecular, ecological, physiological, biological, and further mor- phological features are taken into account as well. Following the groups identified by Hagedorn & Scholler (1999), these predatory fungi are divided into four genera: Arthrobotrys Corda forming adhesive networks, Drechslerella Subram. forming constricting rings, Dactylellina M. Morelet forming stalked adhesive knobs, and Gamsylella gen. nov. for species producing adhesive columns and unstalked knobs. Eighty-two species are accepted, for 51 of which new combinations are proposed. Keywords: Nematophagous fungi, Orbiliaceae, Arlhrobolrys, Daclylellina, Drechslerella, Gamsylella gen. nov, trapping devices, taxonomy. Present generic and phylogenetic concepts for predatory -

1 the Genus Thielavia Zopf (Chaetomiaceae, Sordariales) Is Characterized By
UNIVERSITAT ROVIRA I VIRGILI ESTUDIO TAXONOMICO DE LOS ASCOMYCETES DEL SUELO Akberto Miguel Stchingel Glikman ISBN:978-84-691-1881-8 /DL:T-342-2008 1 The genus Thielavia Zopf (Chaetomiaceae, Sordariales) is characterized by 2 spherical, non ostiolate ascomata, with a thin peridium, and one-celled, darkly 3 pigmented ascospores. Revisions of this genus are due to Mouchacca (1973), 4 Malloch and Cain (1973) and Arx et al (1988). Arx (1973) excluded T. sepedonium 5 Emmons and other similar species with ascospores with two germ pores and 6 Myceliophthora Constantin anamorphs, and erected the genus Corynascus Arx to 7 accommodate these species. The last, most comprehensive, revision is due to Arx 8 et al (1988) who accepted 14 species. Later, two more species have been 9 described (Chen and Chen 1996, Ito et al 1998). Here we describe two new 10 species recently isolated from Easter Island and Indian soils respectively. 11 12 MATERIALS AND METHODS 13 14 Indian soil samples were collected close Jaipur, state of Rajasthan. It is a tropical 15 semiarid region dominated by a hot climate. The average temperature is 23 C in 16 winter and 35 C in summer, and the total annual precipitation is about 900 mm. 17 Vegetation is mainly grasses and shrubs. Easter Island soil samples were collected 18 near Hanga-Roa city. It is a triangular, volcanic, semiarid island of 162 km2 located 19 in the middle of the Pacific Ocean. The vegetation is composed mainly of grasses, 20 with reduced forest masses which are composed of a few imported trees, such as 21 Eucaliptus spp. -
A Stable Backbone for the Fungi
A stable backbone for the fungi Ingo Ebersberger1, Matthias Gube2, Sascha Strauss1, Anne Kupczok1, Martin Eckart2,3, Kerstin Voigt2,3, Erika Kothe2, and Arndt von Haeseler1 1 Center for Integrative Bioinformatics Vienna (CIBIV), University of Vienna, Medical University of Vienna, University of Veterinary Medicine Vienna, Austria 2 Friedrich Schiller University, Institute for Microbiology, Jena, Germany 3 current address: Fungal Reference Centre, Jena, Germany 1 Fungi are abundant in the biosphere. They have fascinated mankind as far as written history goes and have considerably influenced our culture. In biotechnology, cell biology, genetics, and life sciences in general fungi constitute relevant model organisms. Once the phylogenetic relationships of fungi are stably resolved individual results from fungal research can be combined into a holistic picture of biology. However, and despite recent progress1-3, the backbone of the fungal phylogeny is not yet fully resolved. Especially the early evolutionary history of fungi4-6 and the order or below-order relationships within the ascomycetes remain uncertain. Here we present the first phylogenomic study for a eukaryotic kingdom that merges all publicly available fungal genomes and expressed sequence tags (EST) to build a data set comprising 128 genes and 146 taxa. The resulting tree provides a stable phylogenetic backbone for the fungi. Moreover, we present the first formal supertree based on 161 fungal taxa and 128 gene trees. The combined evidences from the trees support the deep-level stability of the fungal groups towards a comprehensive natural system of the fungi. They indicate that the classification of the fungi, especially their alliance with the Microsporidia, requires careful revision. -
Ascomycota, Fungi)
Fungal Diversity DOI 10.1007/s13225-014-0296-3 Coprophilous contributions to the phylogeny of Lasiosphaeriaceae and allied taxa within Sordariales (Ascomycota, Fungi) Åsa Kruys & Sabine M. Huhndorf & Andrew N. Miller Received: 28 November 2013 /Accepted: 12 July 2014 # School of Science 2014 Abstract The phylogenetic relationships of Keywords Ascomal wall . Ascomycete . β-tubulin . LSU Lasiosphaeriaceae are complicated in that the family is nrDNA . Systematics,Taxonomy paraphyletic and includes Sordariaceae and Chaetomiaceae, as well as several polyphyletic genera. This study focuses on the phylogenetic relationships of the coprophilous genera, Introduction Anopodium, Apodospora, Arnium, Fimetariella and Zygospermella. They are traditionally circumscribed based Sordariales is a large group of microfungi that occur world- on ascospore characters, which have proven homoplasious wide as degraders of dung, wood, plant material, and soil in other genera within the family. Our results based on LSU (Cannon and Kirk 2007). The order includes several well- nrDNA and ß–tubulin sequences distinguish four lineages of known members including species of Chaetomium that are Lasiosphaeriaceae taxa. Anopodium joins the clade of mor- common indoor contaminants associated with high humidity, phologically similar, yellow-pigmented species of and the model organisms Neurospora crassa, Podospora Cercophora and Lasiosphaeria. Apodospora is monophyletic anserina, Sordaria fimicola,andS. macrospora and joins a larger group of taxa with unclear affinities to each -
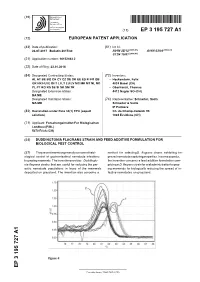
Duddingtonia Flagrans Strain and Feed Additive Formulation for Biological Pest Control
(19) TZZ¥_ _T (11) EP 3 195 727 A1 (12) EUROPEAN PATENT APPLICATION (43) Date of publication: (51) Int Cl.: 26.07.2017 Bulletin 2017/30 A01N 25/12 (2006.01) A01N 63/04 (2006.01) C12N 15/81 (2006.01) (21) Application number: 16152463.2 (22) Date of filing: 22.01.2016 (84) Designated Contracting States: (72) Inventors: AL AT BE BG CH CY CZ DE DK EE ES FI FR GB • Heckendorn, Felix GR HR HU IE IS IT LI LT LU LV MC MK MT NL NO 4024 Basel (CH) PL PT RO RS SE SI SK SM TR • Oberhänsli, Thomas Designated Extension States: 4412 Nuglar SO (CH) BA ME Designated Validation States: (74) Representative: Schneiter, Sorin MA MD Schneiter & Vuille IP Partners (83) Declaration under Rule 32(1) EPC (expert Ch. de Champ-Colomb 7B solution) 1024 Ecublens (CH) (71) Applicant: Forschungsinstitut Fur Biologischen Landbau (FiBL) 5070 Frick (CH) (54) DUDDINGTONIA FLAGRANS STRAIN AND FEED ADDITIVE FORMULATION FOR BIOLOGICAL PEST CONTROL (57) The present invention generally concerns the bi- method for selecting D. flagrans strains exhibiting im- ological control of gastrointestinal nematode infections proved nematode capturing properties. In some aspects, in grazing mammals. The invention provides Duddingto- the invention concerns a feed additive formulation com- nia flagrans strains that are useful for reducing the par- prising a D. flagrans strain for oral administration to graz- asitic nematode populations in feces of the mammals ing mammals for biologically reducing the spread of in- deposited on grassland. The invention also concerns a fective nematodes on grassland. EP 3 195 727 A1 Printed by Jouve, 75001 PARIS (FR) EP 3 195 727 A1 Description Technical Field 5 [0001] The present invention relates to new strains of Duddingtonia flagrans, to tools and methods for identifying a D.