Biochemical Mechanism of RNA Interference in Higher Organisms: a Dissertation
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

2008-Annual-Report.Pdf
whitehead institute 2008 AnnuAl RepoRt. a year in the life of a scientific community empowered to explore biology’s most fundamental questions for the betterment of human health. whitehead institute 2008 annual report a Preserving the mission, contents facing the future 1 preserving the mission, it’s customary in this space to recount and reflect facing the future on the accomplishments of the year gone by. i’ll certainly do so here—proudly—but in many 5 scientific achievement important ways, 2008 was about positioning the institute for years to come. 15 principal investigators many colleges, universities, and independent 30 whitehead fellows research institutions found themselves in dire fiscal positions at the close of 2008 and entered 34 community evolution 2009 in operational crisis reflective of the global economic environment. hiring freezes and large- 40 honor roll of donors scale workforce reductions have become the norm. although whitehead institute is certainly not 46 financial summary insulated from the impact of the downturn, i am 48 leadership pleased and somewhat humbled to report that the institute remains financially strong and no less 49 sited for science committed to scientific excellence. david page, director over the past two years, we have been engaged in a focused effort to increase efficiency and editor & direCtor reduce our administrative costs, with the explicit goal of ensuring that as much of the institute’s matt fearer revenue as possible directly supports Whitehead research. Our approach, which has resulted in assoCiate editor nicole giese a 10-percent reduction in operational expense, has been carefully considered. every decision offiCe of CommuniCation and PubliC affairs has been evaluated not just for its potential effects on our scientific mission, but also for 617.258.5183 www.whitehead.mit.edu possible consequences to the whitehead community and its unique culture. -

Dr. David P. Bartel Journal Interviews
Home About Scientific Press Room Contact Us ● ScienceWatch Home ● Interviews Featured Interviews Author Commentaries Institutional Interviews 2008 : June 2008 - Author Commentaries : Dr. David P. Bartel Journal Interviews Podcasts AUTHOR COMMENTARIES - 2008 ● Analyses June 2008 Featured Analyses Dr. David P. Bartel A Featured Scientist from Essential Science IndicatorsSM What's Hot In... Special Topics Earlier this year, ScienceWatch.com published an analysis of high-impact research in Molecular Biology & Genetics over the past five years. The ● Data & Rankings analysis ranked David Bartel #2 among authors publishing high-impact papers in this field, based on 19 papers cited a total of 4,542 times. Sci-Bytes In Essential Science Indicators from Thomson Reuters, Dr. Bartel's record Fast Breaking Papers includes 76 original articles and review papers published between January 1, New Hot Papers 1998 and February 29, 2008, with a total of 10,386 cites. These papers can be found in the fields of Molecular Biology & Genetics, Biology & Emerging Research Fronts Biochemistry, and Plant & Animal Science. Fast Moving Fronts Research Front Maps Current Classics Dr. Bartel is a Howard Hughes Medical Institute investigator whose lab is based at the Whitehead Institute Top Topics for Biomedical Research in Cambridge, Massachusetts. He is also a Professor in the Department of Biology at MIT. Rising Stars New Entrants In the interview below, he talks with correspondent Gary Taubes about his highly cited research. Country Profiles What motivated your research originally on these small, non-coding RNAs, and particularly the 2000 Cell paper (Zamore PD, et al., RNAi: double-stranded RNA directs the ATP-dependent ● About Science Watch cleavage of mRNA at 21 to 23 nucleotide intervals," 101[1]: 25-33, 31 March 200), which is your most-highly cited paper that’s not a review? Methodology Tom Tuschl and Phil Zamore [see also] were postdocs in my lab working in collaboration with Phil Sharp Archives [see also] on the biochemistry of RNAi. -

Fire Departments of Pathology and Genetics, Stanford University School of Medicine, 300 Pasteur Drive, Room L235, Stanford, CA 94305-5324, USA
GENE SILENCING BY DOUBLE STRANDED RNA Nobel Lecture, December 8, 2006 by Andrew Z. Fire Departments of Pathology and Genetics, Stanford University School of Medicine, 300 Pasteur Drive, Room L235, Stanford, CA 94305-5324, USA. I would like to thank the Nobel Assembly of the Karolinska Institutet for the opportunity to describe some recent work on RNA-triggered gene silencing. First a few disclaimers, however. Telling the full story of gene silencing would be a mammoth enterprise that would take me many years to write and would take you well into the night to read. So we’ll need to abbreviate the story more than a little. Second (and as you will see) we are only in the dawn of our knowledge; so consider the following to be primer... the best we could do as of December 8th, 2006. And third, please understand that the story that I am telling represents the work of several generations of biologists, chemists, and many shades in between. I’m pleased and proud that work from my labo- ratory has contributed to the field, and that this has led to my being chosen as one of the messengers to relay the story in this forum. At the same time, I hope that there will be no confusion of equating our modest contributions with those of the much grander RNAi enterprise. DOUBLE STRANDED RNA AS A BIOLOGICAL ALARM SIGNAL These disclaimers in hand, the story can now start with a biography of the first main character. Double stranded RNA is probably as old (or almost as old) as life on earth. -

Whitehead Faculty and Fellows Members
Whitehead Faculty and Fellows Members Ask a visiting scientist to describe Whitehead Institute and three themes David Bartel studies microRNAs and other small RNAs emerge immediately: the exceptional quality of the scientific staff, the that specify the destruction and/or translational collaborative spirit, and the ethos that encourages researchers at every level repression of mRNAs. He also studies mRNAs, focusing to share new ideas and benefit from the insights and experience of their on their untranslated regions and poly(A) tails, and colleagues. how these regions recruit and mediate regulatory processes. His lab found that microRNAs affect most human protein-coding genes, either by regulating The key to this combination of excellence and accessibility is, of course, the them or by shaping their evolution. Faculty. “From the beginning, we sought researchers who had terrific scientific instincts, but we were also looking for people who would feel comfortable in Iain Cheeseman investigates the process of our open environment,” says Founding Member Gerald Fink. “The exchange chromosome segregation and cell division. In of energy and new ideas never stops, from the formal research retreat, to the particular, he uses proteomics, biochemistry, cell faculty lunches, to the countless informal conversations in hallways and biology, and functional approaches to examine the lounges.” composition, structure, organization and function of the kinetochore — the group of proteins that assemble Whitehead Institute faculty, known as Members, are selected through a joint at the centromere and are required for chromosome appointment process with the Massachusetts Institute of Technology (MIT) segregation and cell division. Department of Biology. Whitehead Institute is solely responsible for their salary and research. -

Micrornas Generate Gene Expression Thresholds with Ultrasensitive Transitions
MicroRNAs generate gene expression thresholds with ultrasensitive transitions Shankar Mukherji1,*, Margaret S. Ebert2,3,*, Grace X. Zheng2,4, John S. Tsang5, Phillip A. Sharp2,3, and Alexander van Oudenaarden2,6,# dramatically. In particular, we show that regulation by Short Abstract — MicroRNAs are an abundant class of post- miRNAs establishes a threshold level of target mRNA below transcriptional regulator whose repressive effects are thought which protein production is highly repressed. Beyond this to have broad reach in terms of number of targets but modest threshold, there is a regime in which expression responds effect for any given target. In this study we performed a single- cell analysis of the effects of microRNA-mediated repression. ultrasensitively to target mRNA input until reaching high We observe that microRNAs generate gene expression enough mRNA levels to almost escape repression by thresholds by which target protein production is robustly miRNA. We constructed a mathematical model, similar to silenced below a given transcriptional activity and is activated models used to describe small RNA (sRNA) reguation in in an ultrasensitive fashion as the transcriptional activity is bacteria [7] and protein-protein titration effects [8,9], increased. The threshold can be tuned by modulating the describing repression of target gene expression by both non- microRNA concentration and the number of microRNA binding sites in the target 3’-UTR. catalytic and catalytic activity of miRNA. The model predicted, and experiments confirmed, that the ultrasensitive Keywords — microRNA, gene expression, post- regime could be shifted to higher target mRNA levels by transcriptional regulation transfecting additional miRNA or by increasing the number of miRNA binding sites in the 3’ UTR of the target mRNA. -

Signature of Author
Computational and experimental analysis of plant microRNAs by Matthew W. Jones-Rhoades B.A., Chemistry (1999) Grinnell College Submitted to the Department of Biology in Partial Fulfillment of the Requirement for the Degree of Doctor of Philosophy in Biology MASSACHUSETTSINSTATE at the OF TECHNOLOGY Massachusetts Institute of Technology MAY 2 7 2005 June 2005 LIBRARIES © 2005 Massachusetts Institute of Technology All rights reserved Signatureof Author.................................... ................. ................. Department of Biology x.. '"---.~ May 20, 2005 Certified by ......... · .................................................... David P. Bartel Professor of Biology Thesis Supervisor Acceptedby ... ......... Stephen P. Bell Professor of Biology Chairman, Graduate Student Committee ,' tti v ;S Computational and experimental analysis of plant microRNAs Matthew W. Jones-Rhoades Submitted to the Department of Biology on May 20, 2005 in Partial Fulfillment of the Requirement for the Degree of Doctor of Philosophy in Biology ABSTRACT MicroRNAs (miRNAs) are small, endogenous, non-coding RNAs that mediate gene regulation in plants and animals. We demonstrated that Arabidopsis thaliana miRNAs are highly complementary (0-3 mispairs in an ungapped alignment) to more mRNAs than would be expected by chance. These mRNAs are therefore putative regulatory targets of their complementary miRNAs. Many miRNA complementary sites are conserved to the monocot Oryza sativa (rice), implying evolutionary conservation based on function at the nucleotide level. The majority of predicted miRNA targets encode for transcription factors and other proteins with known or inferred roles in developmental patterning, implying that the miRNAs themselves are high-level regulators of development. Our findings indicated that miRNAs are key components of numerous regulatory circuits in plants and set the stage for numerous additional experiments to investigate in depth the significance of miRNA-mediated regulation for particular target families and genes. -
Springer A++ Viewer
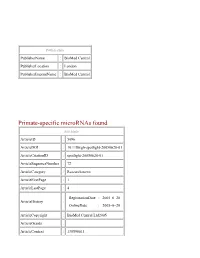
PublisherInfo PublisherName : BioMed Central PublisherLocation : London PublisherImprintName : BioMed Central Primate-specific microRNAs found ArticleInfo ArticleID : 5096 ArticleDOI : 10.1186/gb-spotlight-20050620-01 ArticleCitationID : spotlight-20050620-01 ArticleSequenceNumber : 72 ArticleCategory : Research news ArticleFirstPage : 1 ArticleLastPage : 4 RegistrationDate : 2005–6–20 ArticleHistory : OnlineDate : 2005–6–20 ArticleCopyright : BioMed Central Ltd2005 ArticleGrants : ArticleContext : 130596611 Charles Q Choi Email: [email protected] Israeli scientists have identified what may be the first microRNAs specific to primates, in research published online June 19 in Nature Genetics. They suggest these new microRNAs might mean that hundreds remain to be found in the human genome. "Finding a group of genes that is specific to primates is very important for understanding our evolution, and bears significant diagnostic and therapeutic potential," lead researcher Isaac Bentwich, at Rosetta Genomics in Rehovot, told The Scientist. MicroRNAs are single-stranded RNAs roughly 22 nucleotides long that regulate gene expression by binding to target gene mRNAs. The DNA sequence that codes for a microRNA gene includes the microRNA sequence and a nearby complementary sequence that, when transcribed, form a double- stranded RNA hairpin loop. Past studies have identified 222 human microRNAs, of which scientists had confirmed the sequences of only 86 in humans. To uncover more, Bentwich and colleagues computationally folded the entire human genome into hairpins 55 nucleotides or longer. Of 11 million potential hairpins, 434,239 appeared to be viable microRNA candidates. "Conventional approaches start with genome comparisons, to identify sequences important enough to be conserved across species. This new approach doesn't start with genome alignments. This allows them to find microRNAs not conserved beyond primates," Victor Ambros of Dartmouth Medical School in Hanover, N.H., who did not participate in this study, told The Scientist. -

(12) United States Patent (10) Patent No.: US 8,790,922 B2 Tuschl Et Al
USOO8790922B2 (12) United States Patent (10) Patent No.: US 8,790,922 B2 Tuschl et al. (45) Date of Patent: *Jul. 29, 2014 (54) RNASEQUENCE-SPECIFIC MEDIATORS OF 5,576,208 A 11/1996 Monia et al. RNA INTERFERENCE 5,578,716 A 1 1/1996 Szyfetal. 5,580,859 A 12/1996 Felgner et al. 5,594,122 A 1/1997 Friesen (75) Inventors: Thomas Tuschl, Brooklyn, NY (US); 5,624,803 A 4/1997 Noonberg et al. Phillip D. Zamore, Northborough, MA 5,624,808 A 4/1997 Thompson et al. (US); Phillip A. Sharp, Newton, MA 5,670,633. A 9, 1997 Cook et al. 5,672,695 A 9, 1997 Eckstein et al. (US); David P. Bartel, Brookline, MA 5,674,683 A 10, 1997 KOO1 (US) 5,712.257 A 1/1998 Carter 5,719,271 A 2f1998 Cook et al. (73) Assignees: Max-Planck-Gesellschaft zur 5,770,580 A 6/1998 Ledley et al. Forderung der Wissenschaften E.V. 5,795,715 A 8, 1998 Livache et al. Munich (DE); Massachusetts Institute 5,801,154 A 9, 1998 Baracchini et al. 5,814,500 A 9, 1998 Dietz of Technology, Cambridge, MA (US); 5,898,031 A 4/1999 Crooke Whitehead Institute for Biomedical 5,908,779 A 6/1999 Carmichael et al. Research, Cambridge, MA (US); 5,919,722 A 7/1999 Verduijn et al. University of Massachusetts, Boston, 5,919,772 A 7/1999 Szyfetal. 5,972,704 A 10/1999 Draper et al. MA (US) 5.998,203 A 12/1999 Matulic-Adamic et al. -

Molecular Characterization of Animal Micrornas: Sequence, Expression, and Stability
Molecular Characterization of Animal microRNAs: Sequence, Expression, and Stability. by Nelson C. Lau B.S. Biochemistry and Molecular Biology University at Albany, SUNY, 1999 SUBMITTED TO THE DEPARTMENT OF BIOLOGY IN PARTIAL FULFILLMENT OF THE REQUIREMENTS FOR THE DEGREE OF DOCTOR OF PHILOSOPHY AT THE MASSACHUSETTS INSTITUTE OF TECHNOLOGY SEPTEMBER 2004 ( 2004 Nelson C. Lau. All Rights Reserved. The author hereby grants to MIT permission to reproduce and to distribute publicly paper and electronic copies of this thesis document in whole or in part. Signature of Author: Department of Biology (June 30, 2004) Certified by: / David P. Bartel Professor of Biology Thesis Supervisor Accepted by: qtenhin P Rfll' Professor of Biology Chair, Biology Graduate Committee MASSACHUSETTS INSiEUTE OF TECHNOLOGY JUL 0 1 2004 1 LIBRARIES ARCHIVES Molecular Characterization of Animal microRNAs: Sequence, Expression, and Stability. by Nelson C. Lau Submitted to the Department of Biology On June 30, 2004 in Partial Fulfillment of the Requirements for the Degree of Doctor of Philosophy ABSTRACT Multicellular organisms possess natural gene-regulatory pathways that employ small RNAs to negatively regulate gene expression. In nematodes, the small temporal RNAs (stRNAs), lin-4 and let-7, negatively regulate genes important in specifying developmental timing. A gene- silencing pathway present in plants, fungi and animals called RNA interference, involves the conversion of long double-stranded RNA into short interfering RNAs, which can serve to negatively regulate endogenous genes or suppress the replication of viruses and transposons. To investigate how wide a role small RNAs play in regulating gene expression in animals, we developed a RNA cloning procedure and first applied it to the cloning of small RNAs from the nematode, Caenorhabditis elegans. -

Discovering the First Microrna-Targeted Drug
Published October 29, 2012 JCB: Feature Discovering the first microRNA-targeted drug Morten Lindow1,2 and Sakari Kauppinen2,3 1Department of Biology, The Bioinformatics Centre, University of Copenhagen, DK-2200 Copenhagen, Denmark 2Santaris Pharma, DK-2970 Hørsholm, Denmark 3Department of Health Science and Technology, Aalborg University Copenhagen, DK-2450 Copenhagen SV, Denmark MicroRNAs (miRNAs) are important post-transcriptional first human oncogenic miRNAs (Eis et al., 2005; He et al., 2005; regulators of nearly every biological process in the cell Costinean et al., 2006). Since then, 18,226 miRNAs have been an- notated in animals, plants, and viruses, including 1,921 miRNAs and play key roles in the pathogenesis of human disease. encoded in the human genome (Kozomara and Griffiths-Jones, As a result, there are many drug discovery programs 2011). Indeed, miRNAs are predicted to repress a large fraction that focus on developing miRNA-based therapeutics. The of all protein-coding genes and to participate in the regulation most advanced of these programs targets the liver- of almost every biological process in the cell (Kloosterman and Downloaded from expressed miRNA-122 using the locked nucleic acid Plasterk, 2006; Bushati and Cohen, 2007; Bartel, 2009; Friedman (LNA)–modified antisense oligonucleotide miravirsen. et al., 2009; Ambros, 2011). Moreover, recent work implies that miRNA dysregulation is frequently associated with the patho- Here, we describe the discovery of miravirsen, which is genesis of human diseases (Gottwein and Cullen, 2008; Ventura currently in phase 2 clinical trials for treatment of hepatitis and Jacks, 2009; Williams et al., 2009; Mendell and Olson, C virus (HCV) infection. -
Xuebing Wu, Ph.D
Xuebing Wu, Ph.D. 630 West 168th Street Phone: (617) 301-1637 PS10-401 Email: [email protected] New York, NY 10032 Web: wu-lab.net EMPLOYMENT 11/1/2018- Assistant Professor Departments of Medicine and Systems Biology Columbia University Irving Medical Center 2014-2018 Helen Hay Whitney Postdoctoral Fellow Whitehead Institute for Biomedical Research Advisor: David P. Bartel, Ph.D. Research: Widespread influence of RNA structures on mRNA processing and stability EDUCATION 2009-2014 Ph.D. in Computational and Systems Biology Massachusetts Institute of Technology (MIT) Advisor: Phillip A. Sharp, Ph.D. (co-advisor: Christopher B. Burge, Ph.D.) Thesis: Mechanism and function of pervasive noncoding transcription in mammalian genomes 2007-2009 M.S. in Control Science and Engineering (Area: Bioinformatics) Department of Automation, Tsinghua University, China Advisor: Rui Jiang, Ph.D. Thesis: Disease gene and pathway identification using network alignment approaches 2003-2007 B.S. in Control Science and Engineering Department of Automation, Tsinghua University, China Advisor: Shao Li, M.D. (co-advisor: Michael Q. Zhang, Ph.D.) Thesis: Network-based global inference of human disease genes PUBLICATIONS * co-first authors; # co-corresponding authors 1. Zhang C*, Konermann S*, Brideau NJ, Lotfy P, Wu X, Novick SJ, Strutzenberg T, Griffin PR, Hsu PD, Lyumkis D. Structural basis for the RNA-guided ribonuclease activity of CRISPR-Cas13d. Cell. 2018, 175:212-223 2. Tycko J, Barrera LA, Huston N, Friedland AE, Wu X, Gootenberg JS, Abudayyeh OO, Myer VE, Wilson CJ, Hsu PD. Pairwise library screen systematically interrogates Staphylococcus aureus Cas9 specificity in human cells. Nature Communications. 2018, 9:2962 3. -

Singarna SYMPOSIUM – Speakers
SingaRNA SYMPOSIUM – Speakers Yue Wan Junior Principal Investigator, GIS, A*STAR, Singapore Title: In vivo mapping of eukaryotic RNA interactomes genome-wide using psoralen crosslinking Yue Wan received her B.Sc. in Cell Biology and Biochemistry from the Univer- sity of California, San Diego. She obtained her PhD in Cancer Biology from Stanford University, California, USA, under the mentorship of Howard Y. Chang. During her PhD, she developed the first high-throughput method for probing RNA structures genome-wide. Yue is a recipient of the NSS-PhD scholarship from Agency for Science, Technology and Research (A*STAR) in Singapore. She is currently a Junior Principal Investigator in the Genome Insti- tute of Singapore and a Society in Science-Branco Weiss Fellow. Yue is a recipient of the Young Scientist Award and a TR35 Asia Finalist. She is inter- ested in studying functional RNA structures and understanding their roles in regulating cellular biology. Luca Cartegni Assistant Professor, Rutgers University, USA Title: Therapeutic manipulation of RNA processing in cancer by antisense compounds The focus of our research is the understanding of the role of deregulated pre-mRNA splicing in cancer events, and the development of novel approaches to harness RNA processing to induce the expression of natural dominant-negative variants of active oncogenes in cancer, especially in the context of relapsing tumors which have become resistant to traditional thera- pies. In particular, we use antisense compounds to modulate alternative splicing and/or to activate intronic polyadenylation, in order to promote the expression of natural antagonists. Some of the targets we are focusing on include the induction of soluble decoy variants from VEGFR, EGFR, MET and other Receptor Tyrosine Kinases, as well as oncogenic transcription factors such as STAT3 and others.