Targeting DNA Repair, Cell Cycle, and Tumor Microenvironment in B Cell Lymphoma
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Aurora Kinase a in Gastrointestinal Cancers: Time to Target Ahmed Katsha1, Abbes Belkhiri1, Laura Goff3 and Wael El-Rifai1,2,4*
Katsha et al. Molecular Cancer (2015) 14:106 DOI 10.1186/s12943-015-0375-4 REVIEW Open Access Aurora kinase A in gastrointestinal cancers: time to target Ahmed Katsha1, Abbes Belkhiri1, Laura Goff3 and Wael El-Rifai1,2,4* Abstract Gastrointestinal (GI) cancers are a major cause of cancer-related deaths. During the last two decades, several studies have shown amplification and overexpression of Aurora kinase A (AURKA) in several GI malignancies. These studies demonstrated that AURKA not only plays a role in regulating cell cycle and mitosis, but also regulates a number of key oncogenic signaling pathways. Although AURKA inhibitors have moved to phase III clinical trials in lymphomas, there has been slower progress in GI cancers and solid tumors. Ongoing clinical trials testing AURKA inhibitors as a single agent or in combination with conventional chemotherapies are expected to provide important clinical information for targeting AURKA in GI cancers. It is, therefore, imperative to consider investigations of molecular determinants of response and resistance to this class of inhibitors. This will improve evaluation of the efficacy of these drugs and establish biomarker based strategies for enrollment into clinical trials, which hold the future direction for personalized cancer therapy. In this review, we will discuss the available data on AURKA in GI cancers. We will also summarize the major AURKA inhibitors that have been developed and tested in pre-clinical and clinical settings. Keywords: Aurora kinases, Therapy, AURKA inhibitors, MNL8237, Alisertib, Gastrointestinal, Cancer, Signaling pathways Introduction stage [9-11]. Furthermore, AURKA is critical for Mitotic kinases are the main proteins that coordinate ac- bipolar-spindle assembly where it interacts with Ran- curate mitotic processing [1]. -

The Interactions of DNA Repair, Telomere Homeostasis, and P53 Mutational Status in Solid Cancers: Risk, Prognosis, and Prediction
cancers Review The Interactions of DNA Repair, Telomere Homeostasis, and p53 Mutational Status in Solid Cancers: Risk, Prognosis, and Prediction Pavel Vodicka 1,2,3, Ladislav Andera 4,5, Alena Opattova 1,2,3,† and Ludmila Vodickova 1,2,3,*,† 1 Institute of Experimental Medicine, AS CR, 142 20 Prague, Czech Republic; [email protected] (P.V.); [email protected] (A.O.) 2 First Medical Faculty, Charles University, 121 08 Prague, Czech Republic 3 Biomedical Center, Faculty of Medicine in Pilsen, Charles University, 301 00 Pilsen, Czech Republic 4 Institute of Biotechnology, AS CR, 252 50 Vestec, Czech Republic; [email protected] 5 Institute of Molecular Genetics, AS CR, 142 20 Prague, Czech Republic * Correspondence: [email protected] † These authors contribured eaqually to this paper. Simple Summary: p53 is a nuclear transcription factor with a pro-apoptotic function. Somatic mutations in this gene represent one of the most critical events in human carcinogenesis. The disruption of genomic integrity due to the accumulation of various kinds of DNA damage, deficient DNA repair capacity, and alteration of telomere homeostasis constitute the hallmarks of malignant diseases. The main aim of our review was to accentuate a complex comprehension of the interactions between fundamental players in carcinogenesis of solid malignancies such as DNA damage response, telomere homeostasis and TP53. Abstract: The disruption of genomic integrity due to the accumulation of various kinds of DNA Citation: Vodicka, P.; Andera, L.; Opattova, A.; Vodickova, L. The damage, deficient DNA repair capacity, and telomere shortening constitute the hallmarks of malig- Interactions of DNA Repair, Telomere nant diseases. -

Structure-Based Discovery and Bioactivity Evaluation of Novel
molecules Article Structure-Based Discovery and Bioactivity Evaluation of Novel Aurora-A Kinase Inhibitors as Anticancer Agents via Docking-Based Comparative Intermolecular Contacts Analysis (dbCICA) Majd S. Hijjawi 1 , Reem Fawaz Abutayeh 2 and Mutasem O. Taha 3,* 1 Department of Pharmacology, Faculty of Medicine, The University of Jordan, Amman 11942, Jordan; [email protected] 2 Department of Pharmaceutical Chemistry and Pharmacognosy, Faculty of Pharmacy, Applied Science Private University, Amman 11931, Jordan; [email protected] 3 Department of Pharmaceutical Sciences, Faculty of Pharmacy, University of Jordan, Amman 11942, Jordan * Correspondence: [email protected]; Tel.: +962-6-535-5000 Academic Editors: Helen Osborn, Mohammad Najlah, Jean Jacques Vanden Eynde, Annie Mayence and Tien L. Huang Received: 15 October 2020; Accepted: 11 December 2020; Published: 18 December 2020 Abstract: Aurora-A kinase plays a central role in mitosis, where aberrant activation contributes to cancer by promoting cell cycle progression, genomic instability, epithelial-mesenchymal transition, and cancer stemness. Aurora-A kinase inhibitors have shown encouraging results in clinical trials but have not gained Food and Drug Administration (FDA) approval. An innovative computational workflow named Docking-based Comparative Intermolecular Contacts Analysis (dbCICA) was applied—aiming to identify novel Aurora-A kinase inhibitors—using seventy-nine reported Aurora-A kinase inhibitors to specify the best possible docking settings needed to fit into the active-site binding pocket of Aurora-A kinase crystal structure, in a process that only potent ligands contact critical binding-site spots, distinct from those occupied by less-active ligands. Optimal dbCICA models were transformed into two corresponding pharmacophores. -

Spatiotemporal Dynamics of Aurora B-PLK1-MCAK Signaling Axis
www.nature.com/scientificreports OPEN Spatiotemporal dynamics of Aurora B-PLK1-MCAK signaling axis orchestrates kinetochore Received: 21 October 2014 Accepted: 13 May 2015 bi-orientation and faithful Published: 24 July 2015 chromosome segregation Hengyi Shao1, Yuejia Huang2,3, Liangyu Zhang1,3, Kai Yuan1, Youjun Chu1,3, Zhen Dou1,2, Changjiang Jin1, Minerva Garcia-Barrio3, Xing Liu1,2,3 & Xuebiao Yao1 Chromosome segregation in mitosis is orchestrated by the dynamic interactions between the kinetochore and spindle microtubules. The microtubule depolymerase mitotic centromere-associated kinesin (MCAK) is a key regulator for an accurate kinetochore-microtubule attachment. However, the regulatory mechanism underlying precise MCAK depolymerase activity control during mitosis remains elusive. Here, we describe a novel pathway involving an Aurora B-PLK1 axis for regulation of MCAK activity in mitosis. Aurora B phosphorylates PLK1 on Thr210 to activate its kinase activity at the kinetochores during mitosis. Aurora B-orchestrated PLK1 kinase activity was examined in real- time mitosis using a fluorescence resonance energy transfer-based reporter and quantitative analysis of native PLK1 substrate phosphorylation. Active PLK1, in turn, phosphorylates MCAK at Ser715 which promotes its microtubule depolymerase activity essential for faithful chromosome segregation. Importantly, inhibition of PLK1 kinase activity or expression of a non-phosphorylatable MCAK mutant prevents correct kinetochore-microtubule attachment, resulting in abnormal anaphase with chromosome bridges. We reason that the Aurora B-PLK1 signaling at the kinetochore orchestrates MCAK activity, which is essential for timely correction of aberrant kinetochore attachment to ensure accurate chromosome segregation during mitosis. During cell division, accurate chromosome segregation requires dynamic interactions between kineto- chores and spindle microtubules (MTs), which results in accurate chromosome bi-orientation1–4. -

Cadmium Induced Cell Apoptosis, DNA Damage, De
Int. J. Med. Sci. 2013, Vol. 10 1485 Ivyspring International Publisher International Journal of Medical Sciences 2013; 10(11):1485-1496. doi: 10.7150/ijms.6308 Research Paper Cadmium Induced Cell Apoptosis, DNA Damage, De- creased DNA Repair Capacity, and Genomic Instability during Malignant Transformation of Human Bronchial Epithelial Cells Zhiheng Zhou1*, Caixia Wang2*, Haibai Liu1, Qinhai Huang1, Min Wang1 , Yixiong Lei1 1. School of public health, Guangzhou Medical University, Guangzhou 510182, People’s Republic of China 2. Department of internal medicine of Guangzhou First People’s Hospital affiliated to Guangzhou Medical University, Guangzhou 510180, People’s Republic of China * These authors contributed equally to this work. Corresponding author: Yixiong Lei, School of public health, Guangzhou Medical University, 195 Dongfengxi Road, Guangzhou 510182, People’s Republic of China. Fax: +86-20-81340196. E-mail: [email protected] © Ivyspring International Publisher. This is an open-access article distributed under the terms of the Creative Commons License (http://creativecommons.org/ licenses/by-nc-nd/3.0/). Reproduction is permitted for personal, noncommercial use, provided that the article is in whole, unmodified, and properly cited. Received: 2013.03.23; Accepted: 2013.08.12; Published: 2013.08.30 Abstract Cadmium and its compounds are well-known human carcinogens, but the mechanisms underlying the carcinogenesis are not entirely understood. Our study was designed to elucidate the mech- anisms of DNA damage in cadmium-induced malignant transformation of human bronchial epi- thelial cells. We analyzed cell cycle, apoptosis, DNA damage, gene expression, genomic instability, and the sequence of exons in DNA repair genes in several kinds of cells. -

Expression Profiling of KLF4
Expression Profiling of KLF4 AJCR0000006 Supplemental Data Figure S1. Snapshot of enriched gene sets identified by GSEA in Klf4-null MEFs. Figure S2. Snapshot of enriched gene sets identified by GSEA in wild type MEFs. 98 Am J Cancer Res 2011;1(1):85-97 Table S1: Functional Annotation Clustering of Genes Up-Regulated in Klf4 -Null MEFs ILLUMINA_ID Gene Symbol Gene Name (Description) P -value Fold-Change Cell Cycle 8.00E-03 ILMN_1217331 Mcm6 MINICHROMOSOME MAINTENANCE DEFICIENT 6 40.36 ILMN_2723931 E2f6 E2F TRANSCRIPTION FACTOR 6 26.8 ILMN_2724570 Mapk12 MITOGEN-ACTIVATED PROTEIN KINASE 12 22.19 ILMN_1218470 Cdk2 CYCLIN-DEPENDENT KINASE 2 9.32 ILMN_1234909 Tipin TIMELESS INTERACTING PROTEIN 5.3 ILMN_1212692 Mapk13 SAPK/ERK/KINASE 4 4.96 ILMN_2666690 Cul7 CULLIN 7 2.23 ILMN_2681776 Mapk6 MITOGEN ACTIVATED PROTEIN KINASE 4 2.11 ILMN_2652909 Ddit3 DNA-DAMAGE INDUCIBLE TRANSCRIPT 3 2.07 ILMN_2742152 Gadd45a GROWTH ARREST AND DNA-DAMAGE-INDUCIBLE 45 ALPHA 1.92 ILMN_1212787 Pttg1 PITUITARY TUMOR-TRANSFORMING 1 1.8 ILMN_1216721 Cdk5 CYCLIN-DEPENDENT KINASE 5 1.78 ILMN_1227009 Gas2l1 GROWTH ARREST-SPECIFIC 2 LIKE 1 1.74 ILMN_2663009 Rassf5 RAS ASSOCIATION (RALGDS/AF-6) DOMAIN FAMILY 5 1.64 ILMN_1220454 Anapc13 ANAPHASE PROMOTING COMPLEX SUBUNIT 13 1.61 ILMN_1216213 Incenp INNER CENTROMERE PROTEIN 1.56 ILMN_1256301 Rcc2 REGULATOR OF CHROMOSOME CONDENSATION 2 1.53 Extracellular Matrix 5.80E-06 ILMN_2735184 Col18a1 PROCOLLAGEN, TYPE XVIII, ALPHA 1 51.5 ILMN_1223997 Crtap CARTILAGE ASSOCIATED PROTEIN 32.74 ILMN_2753809 Mmp3 MATRIX METALLOPEPTIDASE -

Charles M. Perou, Phd Associate Professor Departments of Genetics
Charles M. Perou, PhD Associate Professor Departments of Genetics and Pathology Carolina Center for Genome Sciences Lineberger Comprehensive Cancer Center University of North Carolina at Chapel Hill Identification of GBM Subtypes using Gene Expression Profiling Agilent Custom Affymetrix Human Affymetrix HT-HG- 244k Exon 1.0 Array U133A Array Microarray Identify Samples and Genes represented on all 3 platforms Factor Analysis Single unified gene expression measurement for each gene 202 patients 11,681 genes Identification of GBM Subtypes Census Clustering of the 202 samples X 1740 Unsupervised clustering of 1740 genes suggests that 4 subtypes of GBM exist variably expressed genes selected using a unified gene expression measure across 3 expression platforms Core TCGA Samples (173) Gene Ontology/Pathway: with Subtype-defining genes • ProNeural: 1. nervous system development ProNeural Normal-like EGFR Mesenchymal 2. neuron differentiation (SOXs) 3. cell cycle = proliferation 4. cell adhesion molecules 5. ErbB signaling pathway FBXO3 GABRB2 • Normal‐like: SNCG NTSR2 1. nucleotide metabolic process MBP 2. neurological system process DLL3 3. axon NKX2-2, NRXN1, NRXN2 TOP2B, CDC7, MYB 4. neuron projection SOX2, SOX4, SOX10,SOX11 5. synaptic transmission ERBB3, PAK3, PAK7 NCAM1 OLIG2 • EGFR: 1. regulation of transcription 2. cell migration 3. nervous system development FGFR3 4. cell proliferation PDGFRA 5. metal ion binding EGFR AKT2 GLI2 TGFB3 • Mesenchymal: CASP8, CASP5, CASP4, CASP1 1. immune response COL8A2, COL5A1, COL1A1,COL1A2 2. receptor activity ILR4 CHI3L1 3. wound healing TRADD TLR2, TLR4, 4. cytokine and chemokine IGFBI mediated signaling pathway RELB 5. NF‐B Signaling Pathway Correlations between gene expression subtypes and clinical parameters Survival Analysis of Subtypes n=196 Subtypes are correlated with: 1. -
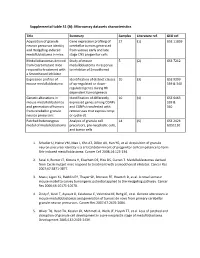
Supplemental Table S1 (A): Microarray Datasets Characteristics
Supplemental table S1 (A): Microarray datasets characteristics Title Summary Samples Literature ref. GEO ref. Acquisition of granule Gene expression profiling of 27 (1) GSE 11859 neuron precursor identity cerebellar tumors generated and Hedgehog‐induced from various early and late medulloblastoma in mice. stage CNS progenitor cells Medulloblastomas derived Study of mouse 5 (2) GSE 7212 from Cxcr6 mutant mice medulloblastoma in response respond to treatment with to inhibitor of Smoothened a Smoothened inhibitor Expression profiles of Identification of distinct classes 10 (3) GSE 9299 mouse medulloblastoma of up‐regulated or down‐ 339 & 340 regulated genes during Hh dependent tumorigenesis Genetic alterations in Identification of differently 10 (4) GSE 6463 mouse medulloblastomas expressed genes among CGNPs 339 & and generation of tumors and CGNPs transfected with 340 from cerebellar granule retroviruses that express nmyc neuron precursors or cyclin‐d1 Patched heterozygous Analysis of granule cell 14 (5) GSE 2426 model of medulloblastoma precursors, pre‐neoplastic cells, GDS1110 and tumor cells 1. Schuller U, Heine VM, Mao J, Kho AT, Dillon AK, Han YG, et al. Acquisition of granule neuron precursor identity is a critical determinant of progenitor cell competence to form Shh‐induced medulloblastoma. Cancer Cell 2008;14:123‐134. 2. Sasai K, Romer JT, Kimura H, Eberhart DE, Rice DS, Curran T. Medulloblastomas derived from Cxcr6 mutant mice respond to treatment with a smoothened inhibitor. Cancer Res 2007;67:3871‐3877. 3. Mao J, Ligon KL, Rakhlin EY, Thayer SP, Bronson RT, Rowitch D, et al. A novel somatic mouse model to survey tumorigenic potential applied to the Hedgehog pathway. Cancer Res 2006;66:10171‐10178. -

Herpes Simplex Virus Type 2: Bystander Or Active Player in Cervical Carcinogenesis?
Journal of Gynecology and Women’s Health ISSN 2474-7602 Mini Review J Gynecol Women’s Health Volume 12 Issue 2 - October 2018 Copyright © All rights are reserved by Jude Ogechukwu Okoye DOI: 10.19080/JGWH.2018.12.555832 Herpes Simplex Virus Type 2: Bystander or Active Player in Cervical Carcinogenesis? Jude Ogechukwu Okoye* Department of Medical Laboratory Science, Babcock University, Nigeria Submission: September 19, 2018 ; Published: October 05, 2018 *Corresponding author: Jude Ogechukwu Okoye, Department of Medical Laboratory Science, Babcock University, Nigeria, Tel: ; Email: Abstract More emphasis have been placed on producing vaccines that prevent host cell entry by Human Papillomavirus (HPV) while less attention andhas beenits potential given to to Herpes up-modulate Simplex both virus HPV type regulatory 2 (HSV-2) andwhich oncogenic potentially gene promotes and downregulate acquisition hostand co-habitationtumour suppressor of human suggest immunodeficiency that it is not a virus (HIV), HPV and Epstein-Barr virus (EBV). The tendency for HSV-2 to induce ulcerations and chronic inflammation following reactivation, to reduce cervical cancer related mortality. bystander in cervical carcinogenesis. Thus, efforts geared towards finding HSV-2 effective microbicide and vaccine should be intensified so as Keywords: Herpes Simplex virus type 2; Human papillomavirus; Epstein-Barr virus; Cervical cancer Abbreviations: HPV: Human Papillomavirus; HIV: Human Immune virus; SSC: Squamous Cell Carcinoma; LMP1: Latent Membrane Protein 1; EBV: Epstein-Barr virus; CIN: Cervical Intraepithelial Neoplasia Introduction Cervical cancer is the second most common type of cancer in a woman’s lifetime and may also facilitate EBV infection, a (9.8%) affecting women worldwide [1-3]. -

Malignant Transformation of Human Fibroblasts Caused by Expression Of
Proc. Nati. Acad. Sci. USA Vol. 86, pp. 187-191, January 1989 Cell Biology Malignant transformation of human fibroblasts caused by expression of a transfected T24 HRAS oncogene (human cell transformation/infinite life-span human flbroblasts/pHO6T1/T24 HRAS expression/malignant fibrosarcoma) PETER J. HURLIN*, VERONICA M. MAHER, AND J. JUSTIN MCCORMICKt Carcinogenesis Laboratory, Fee Hall, Department of Microbiology and Department of Biochemistry, Michigan State University, East Lansing, MI 48824-1316 Communicated by G. N. Wogan, September 19, 1988 ABSTRACT We showed previously that diploid human passage rodent cells were transfected with an HRAS onco- fibroblasts that express a transfected HRAS oncogene from the gene and any one of a number of genes considered to be human bladder carcinoma cell line T24 exhibit several char- involved in causing cellular immortalization, malignant trans- acteristics of transformed cells but do not acquire an infinite formation resulted (5-7). Not surprisingly, transfection of life-span and are not tumorigenic. To extend these studies ofthe HRAS oncogenes into infinite life-span rodent fibroblast cell T24 HRAS in human cells, we have utilized an infinite life-span, lines resulted in malignant transformation (5, 8) in the but otherwise phenotypically normal, human fibroblast cell majority of the studies (9). strain, MSU-1.1, developed in this laboratory after transfec- To test models proposed for ras transformation generated tion of diploid fibroblasts with a viral v-myc oncogene. Trans- from experiments with rodent fibroblasts for their relevance fection of MSU-1.1 cells with the T24 HRAS flanked by two to human cell transformation, we and our colleagues (10-12) transcriptional enhancer elements (pHO6T1) yielded foci of and several other groups (13-21) have used human fibroblasts morphologically transformed cells. -

Tagging Single Nucleotide Polymorphisms in Cell Cycle Control Genes and Susceptibility to Invasive Epithelial Ovarian Cancer
Research Article Tagging Single Nucleotide Polymorphisms in Cell Cycle Control Genes and Susceptibility to Invasive Epithelial Ovarian Cancer Simon A. Gayther,1 Honglin Song,2 Susan J. Ramus,1 Susan Kru¨ger Kjaer,4 Alice S. Whittemore,5 Lydia Quaye,1 Jonathan Tyrer,2 Danielle Shadforth,2 Estrid Hogdall,4 Claus Hogdall,6 Jan Blaeker,7 Richard DiCioccio,8 Valerie McGuire,5 Penelope M. Webb,9 Jonathan Beesley,9 Adele C. Green,9 David C. Whiteman,9 The Australian Ovarian Cancer Study Group,9 The Australian Cancer Study (Ovarian Cancer),20 Marc T. Goodman,10 Galina Lurie,10 Michael E. Carney,10 Francesmary Modugno,10 Roberta B. Ness,11 Robert P. Edwards,12 Kirsten B. Moysich,7 Ellen L. Goode,13 Fergus J. Couch,13 Julie M. Cunningham,13 Thomas A. Sellers,14 Anna H. Wu,15 Malcolm C. Pike,15 Edwin S. Iversen,16 Jeffrey R. Marks,16 Montserrat Garcia-Closas,17 Louise Brinton,17 Jolanta Lissowska,18 Beata Peplonska,19 Douglas F. Easton,3 Ian Jacobs,1 Bruce A.J. Ponder,2 Joellen Schildkraut,16 C. Leigh Pearce,15 Georgia Chenevix-Trench,9 Andrew Berchuck,16 Paul D.P. Pharoah,2 and on behalf of the Ovarian Cancer Association Consortium 1Translational Research Laboratories, University College London, London, United Kingdom; 2Cancer Research United Kingdom Department of Oncology and 3Cancer Research United Kingdom Genetic Epidemiology Unit, University of Cambridge, Strangeways Research Laboratory, Cambridge, United Kingdom; 4Danish Cancer Society, Copenhagen, Denmark; 5Stanford University School of Medicine, Stanford, California; 6University of Copenhagen, -

The RNA Helicase a in Malignant Transformation
www.impactjournals.com/oncotarget/ Oncotarget, Vol. 7, No. 19 The RNA helicase A in malignant transformation Marco Fidaleo1,2, Elisa De Paola1,2 and Maria Paola Paronetto1,2 1 Department of Movement, Human and Health Sciences, University of Rome “Foro Italico”, Rome, Italy 2 Laboratory of Cellular and Molecular Neurobiology, CERC, Fondazione Santa Lucia, Rome, Italy Correspondence to: Maria Paola Paronetto, email: [email protected] Keywords: RHA helicase, genomic stability, cancer Received: October 02, 2015 Accepted: January 29, 2016 Published: February 14, 2016 ABSTRACT The RNA helicase A (RHA) is involved in several steps of RNA metabolism, such as RNA processing, cellular transit of viral molecules, ribosome assembly, regulation of transcription and translation of specific mRNAs. RHA is a multifunctional protein whose roles depend on the specific interaction with different molecular partners, which have been extensively characterized in physiological situations. More recently, the functional implication of RHA in human cancer has emerged. Interestingly, RHA was shown to cooperate with both tumor suppressors and oncoproteins in different tumours, indicating that its specific role in cancer is strongly influenced by the cellular context. For instance, silencing of RHA and/or disruption of its interaction with the oncoprotein EWS-FLI1 rendered Ewing sarcoma cells more sensitive to genotoxic stresses and affected tumor growth and maintenance, suggesting possible therapeutic implications. Herein, we review the recent advances in the cellular functions of RHA and discuss its implication in oncogenesis, providing a perspective for future studies and potential translational opportunities in human cancer. INTRODUCTION females, which contain two X chromosomes [8]. The C. elegans RHA-1 displays about 60% of similarity with both RNA helicase A (RHA) is a DNA/RNA helicase human RHA and Drosophila MLE and is involved in gene involved in all the essential steps of RNA metabolism, silencing.