Autoimmunity in a Mouse Model of Lupus Deficiency Promotes B1a
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Cyclin D2 Activates Cdk2 in Preference to Cdk4 in Human Breast Epithelial Cells
Oncogene (1997) 14, 1329 ± 1340 1997 Stockton Press All rights reserved 0950 ± 9232/97 $12.00 Cyclin D2 activates Cdk2 in preference to Cdk4 in human breast epithelial cells Kimberley J Sweeney, Boris Sarcevic, Robert L Sutherland and Elizabeth A Musgrove Cancer Research Program, Garvan Institute of Medical Research, St Vincent's Hospital, Sydney, NSW 2010, Australia To investigate the possibility of diering roles for cyclins Similarly, overexpression of cyclin D2 in myeloid cells D1 and D2 in breast epithelial cells, we examined the results in a decrease in the duration of G1 and an expression, cell cycle regulation and activity of these two increase in the percentage of cells in S-phase (Ando et G1 cyclins in both 184 normal breast epithelial cells and al., 1993; Kato and Sherr, 1993). Microinjection or T-47D breast cancer cells. Synchronisation studies in 184 electroporation of cyclin D1 or cyclin D2 antibodies cells demonstrated that cyclin D1 and cyclin D2 were demonstrated that these proteins were not only rate- dierentially regulated during G1, with cyclin D2 limiting but essential for progress through G1 (Baldin abundance increasing by 3.7-fold but only small changes et al., 1993; Quelle et al., 1993; Lukas et al., 1995b). in cyclin D1 abundance observed. The functional These eects are thought to be mediated by activation consequences of increased cyclin D2 expression were of cyclin-dependent kinases (CDKs) and consequent examined in T-47D cells, which express no detectable phosphorylation of the product of the retinoblastoma cyclin D2. Induced expression of cyclin D2 resulted in susceptibility gene, pRB (Hunter and Pines, 1994; increases in cyclin E expression, pRB phosphorylation Sherr, 1994). -

P27 and P21 Collaborate in the Regulation of Transcription By
6860–6873 Nucleic Acids Research, 2015, Vol. 43, No. 14 Published online 13 June 2015 doi: 10.1093/nar/gkv593 p27Kip1 and p21Cip1 collaborate in the regulation of transcription by recruiting cyclin–Cdk complexes on the promoters of target genes Serena Orlando1, Edurne Gallastegui1, Arnaud Besson2,3,4, Gabriel Abril1, Rosa Aligue´ 1, Maria Jesus Pujol1 and Oriol Bachs1,* 1Department of Cell Biology, Immunology and Neurosciences, University of Barcelona, 08036-Barcelona, Spain, 2INSERM UMR1037, Cancer Research Center of Toulouse, Toulouse, France, 3Universite´ de Toulouse, Toulouse, France and 4CNRS ERL5294, Toulouse, France Received December 22, 2014; Revised May 20, 2015; Accepted May 23, 2015 ABSTRACT through Cdk20 (2). Many members of this family, includ- ing Cdk4, Cdk6, Cdk2 and Cdk1, are involved in cell-cycle Transcriptional repressor complexes containing regulation (3). Cdk4, Cdk6 and Cdk2 regulate progression p130 and E2F4 regulate the expression of genes in- through the G1 phase although additionally, Cdk2 also reg- volved in DNA replication. During the G1 phase of ulates S phase. Finally, Cdk1 regulates mitosis. Binding the cell cycle, sequential phosphorylation of p130 of Cdks to specific cyclins confers a functional specializa- by cyclin-dependent kinases (Cdks) disrupts these tion to each complex (3). In response to mitogenic stim- complexes allowing gene expression. The Cdk in- uli, the synthesis of the D-type cyclins is induced in early– Kip1 hibitor and tumor suppressor p27 associates with mid G1 phase. These cyclins associate with Cdk4 and Cdk6, p130 and E2F4 by its carboxyl domain on the pro- forming complexes that phosphorylate and inactivate mem- moters of target genes but its role in the regulation bers of the retinoblastoma family of pocket proteins (pRb, of transcription remains unclear. -

Expression Profiling of KLF4
Expression Profiling of KLF4 AJCR0000006 Supplemental Data Figure S1. Snapshot of enriched gene sets identified by GSEA in Klf4-null MEFs. Figure S2. Snapshot of enriched gene sets identified by GSEA in wild type MEFs. 98 Am J Cancer Res 2011;1(1):85-97 Table S1: Functional Annotation Clustering of Genes Up-Regulated in Klf4 -Null MEFs ILLUMINA_ID Gene Symbol Gene Name (Description) P -value Fold-Change Cell Cycle 8.00E-03 ILMN_1217331 Mcm6 MINICHROMOSOME MAINTENANCE DEFICIENT 6 40.36 ILMN_2723931 E2f6 E2F TRANSCRIPTION FACTOR 6 26.8 ILMN_2724570 Mapk12 MITOGEN-ACTIVATED PROTEIN KINASE 12 22.19 ILMN_1218470 Cdk2 CYCLIN-DEPENDENT KINASE 2 9.32 ILMN_1234909 Tipin TIMELESS INTERACTING PROTEIN 5.3 ILMN_1212692 Mapk13 SAPK/ERK/KINASE 4 4.96 ILMN_2666690 Cul7 CULLIN 7 2.23 ILMN_2681776 Mapk6 MITOGEN ACTIVATED PROTEIN KINASE 4 2.11 ILMN_2652909 Ddit3 DNA-DAMAGE INDUCIBLE TRANSCRIPT 3 2.07 ILMN_2742152 Gadd45a GROWTH ARREST AND DNA-DAMAGE-INDUCIBLE 45 ALPHA 1.92 ILMN_1212787 Pttg1 PITUITARY TUMOR-TRANSFORMING 1 1.8 ILMN_1216721 Cdk5 CYCLIN-DEPENDENT KINASE 5 1.78 ILMN_1227009 Gas2l1 GROWTH ARREST-SPECIFIC 2 LIKE 1 1.74 ILMN_2663009 Rassf5 RAS ASSOCIATION (RALGDS/AF-6) DOMAIN FAMILY 5 1.64 ILMN_1220454 Anapc13 ANAPHASE PROMOTING COMPLEX SUBUNIT 13 1.61 ILMN_1216213 Incenp INNER CENTROMERE PROTEIN 1.56 ILMN_1256301 Rcc2 REGULATOR OF CHROMOSOME CONDENSATION 2 1.53 Extracellular Matrix 5.80E-06 ILMN_2735184 Col18a1 PROCOLLAGEN, TYPE XVIII, ALPHA 1 51.5 ILMN_1223997 Crtap CARTILAGE ASSOCIATED PROTEIN 32.74 ILMN_2753809 Mmp3 MATRIX METALLOPEPTIDASE -

Charles M. Perou, Phd Associate Professor Departments of Genetics
Charles M. Perou, PhD Associate Professor Departments of Genetics and Pathology Carolina Center for Genome Sciences Lineberger Comprehensive Cancer Center University of North Carolina at Chapel Hill Identification of GBM Subtypes using Gene Expression Profiling Agilent Custom Affymetrix Human Affymetrix HT-HG- 244k Exon 1.0 Array U133A Array Microarray Identify Samples and Genes represented on all 3 platforms Factor Analysis Single unified gene expression measurement for each gene 202 patients 11,681 genes Identification of GBM Subtypes Census Clustering of the 202 samples X 1740 Unsupervised clustering of 1740 genes suggests that 4 subtypes of GBM exist variably expressed genes selected using a unified gene expression measure across 3 expression platforms Core TCGA Samples (173) Gene Ontology/Pathway: with Subtype-defining genes • ProNeural: 1. nervous system development ProNeural Normal-like EGFR Mesenchymal 2. neuron differentiation (SOXs) 3. cell cycle = proliferation 4. cell adhesion molecules 5. ErbB signaling pathway FBXO3 GABRB2 • Normal‐like: SNCG NTSR2 1. nucleotide metabolic process MBP 2. neurological system process DLL3 3. axon NKX2-2, NRXN1, NRXN2 TOP2B, CDC7, MYB 4. neuron projection SOX2, SOX4, SOX10,SOX11 5. synaptic transmission ERBB3, PAK3, PAK7 NCAM1 OLIG2 • EGFR: 1. regulation of transcription 2. cell migration 3. nervous system development FGFR3 4. cell proliferation PDGFRA 5. metal ion binding EGFR AKT2 GLI2 TGFB3 • Mesenchymal: CASP8, CASP5, CASP4, CASP1 1. immune response COL8A2, COL5A1, COL1A1,COL1A2 2. receptor activity ILR4 CHI3L1 3. wound healing TRADD TLR2, TLR4, 4. cytokine and chemokine IGFBI mediated signaling pathway RELB 5. NF‐B Signaling Pathway Correlations between gene expression subtypes and clinical parameters Survival Analysis of Subtypes n=196 Subtypes are correlated with: 1. -
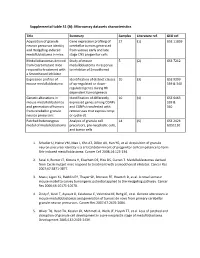
Supplemental Table S1 (A): Microarray Datasets Characteristics
Supplemental table S1 (A): Microarray datasets characteristics Title Summary Samples Literature ref. GEO ref. Acquisition of granule Gene expression profiling of 27 (1) GSE 11859 neuron precursor identity cerebellar tumors generated and Hedgehog‐induced from various early and late medulloblastoma in mice. stage CNS progenitor cells Medulloblastomas derived Study of mouse 5 (2) GSE 7212 from Cxcr6 mutant mice medulloblastoma in response respond to treatment with to inhibitor of Smoothened a Smoothened inhibitor Expression profiles of Identification of distinct classes 10 (3) GSE 9299 mouse medulloblastoma of up‐regulated or down‐ 339 & 340 regulated genes during Hh dependent tumorigenesis Genetic alterations in Identification of differently 10 (4) GSE 6463 mouse medulloblastomas expressed genes among CGNPs 339 & and generation of tumors and CGNPs transfected with 340 from cerebellar granule retroviruses that express nmyc neuron precursors or cyclin‐d1 Patched heterozygous Analysis of granule cell 14 (5) GSE 2426 model of medulloblastoma precursors, pre‐neoplastic cells, GDS1110 and tumor cells 1. Schuller U, Heine VM, Mao J, Kho AT, Dillon AK, Han YG, et al. Acquisition of granule neuron precursor identity is a critical determinant of progenitor cell competence to form Shh‐induced medulloblastoma. Cancer Cell 2008;14:123‐134. 2. Sasai K, Romer JT, Kimura H, Eberhart DE, Rice DS, Curran T. Medulloblastomas derived from Cxcr6 mutant mice respond to treatment with a smoothened inhibitor. Cancer Res 2007;67:3871‐3877. 3. Mao J, Ligon KL, Rakhlin EY, Thayer SP, Bronson RT, Rowitch D, et al. A novel somatic mouse model to survey tumorigenic potential applied to the Hedgehog pathway. Cancer Res 2006;66:10171‐10178. -

Tagging Single Nucleotide Polymorphisms in Cell Cycle Control Genes and Susceptibility to Invasive Epithelial Ovarian Cancer
Research Article Tagging Single Nucleotide Polymorphisms in Cell Cycle Control Genes and Susceptibility to Invasive Epithelial Ovarian Cancer Simon A. Gayther,1 Honglin Song,2 Susan J. Ramus,1 Susan Kru¨ger Kjaer,4 Alice S. Whittemore,5 Lydia Quaye,1 Jonathan Tyrer,2 Danielle Shadforth,2 Estrid Hogdall,4 Claus Hogdall,6 Jan Blaeker,7 Richard DiCioccio,8 Valerie McGuire,5 Penelope M. Webb,9 Jonathan Beesley,9 Adele C. Green,9 David C. Whiteman,9 The Australian Ovarian Cancer Study Group,9 The Australian Cancer Study (Ovarian Cancer),20 Marc T. Goodman,10 Galina Lurie,10 Michael E. Carney,10 Francesmary Modugno,10 Roberta B. Ness,11 Robert P. Edwards,12 Kirsten B. Moysich,7 Ellen L. Goode,13 Fergus J. Couch,13 Julie M. Cunningham,13 Thomas A. Sellers,14 Anna H. Wu,15 Malcolm C. Pike,15 Edwin S. Iversen,16 Jeffrey R. Marks,16 Montserrat Garcia-Closas,17 Louise Brinton,17 Jolanta Lissowska,18 Beata Peplonska,19 Douglas F. Easton,3 Ian Jacobs,1 Bruce A.J. Ponder,2 Joellen Schildkraut,16 C. Leigh Pearce,15 Georgia Chenevix-Trench,9 Andrew Berchuck,16 Paul D.P. Pharoah,2 and on behalf of the Ovarian Cancer Association Consortium 1Translational Research Laboratories, University College London, London, United Kingdom; 2Cancer Research United Kingdom Department of Oncology and 3Cancer Research United Kingdom Genetic Epidemiology Unit, University of Cambridge, Strangeways Research Laboratory, Cambridge, United Kingdom; 4Danish Cancer Society, Copenhagen, Denmark; 5Stanford University School of Medicine, Stanford, California; 6University of Copenhagen, -

CDKN2C-Null Leiomyosarcoma: a Novel
original reports CDKN2C-Null Leiomyosarcoma: A Novel, Genomically Distinct Class of TP53/ RB1–Wild-Type Tumor With Frequent CIC Genomic Alterations and 1p/19q-Codeletion Erik A. Williams, MD1; Radwa Sharaf, PhD1; Brennan Decker, MD, PhD2; Adrienne J. Werth, MD3; Helen Toma, MD3; Meagan Montesion, PhD1; Ethan S. Sokol, PhD1; Dean C. Pavlick, BS1; Nikunj Shah, BS1; Kevin Jon Williams, MD4; Jeffrey M. Venstrom, MD1; Brian M. Alexander, MD, MPH1; Jeffrey S. Ross, MD1,5; Lee A. Albacker, PhD1; Douglas I. Lin, MD, PhD1; Shakti H. Ramkissoon, MD, PhD1,6; and Julia A. Elvin, MD, PhD1 abstract PURPOSE Leiomyosarcoma (LMS) harbors frequent mutations in TP53 and RB1 but few actionable genomic alterations. Here, we searched for recurrent actionable genomic alterations in LMS that occur in the absence of common untreatable oncogenic drivers. METHODS Tissues from 276,645 unique advanced cancers, including 2,570 uterine and soft tissue LMS, were sequenced by hybrid-capture–based next-generation DNA and RNA sequencing/comprehensive genomic profiling of up to 406 genes. We characterized clinicopathologic features of relevant patient cases. RESULTS Overall, 77 LMS exhibited homozygous copy loss of CDKN2C at chromosome 1p32.3 (3.0% of LMS). Genomic alterations (GAs) in TP53, RB1, and ATRX were rare compared with the remainder of the LMS cohort (11.7% v 73.4%, 0% v 54.5%, 2.6% v 24.5%, respectively; all P , .0001). CDKN2C-null LMS patient cases were significantly enriched for GAs in CIC (40.3% v 1.4%) at 19q13.2, CDKN2A (46.8% v 7.0%), and RAD51B (16.9% v 1.7%; all P , .0001). -

Increased Expression of Unmethylated CDKN2D by 5-Aza-2'-Deoxycytidine in Human Lung Cancer Cells
Oncogene (2001) 20, 7787 ± 7796 ã 2001 Nature Publishing Group All rights reserved 0950 ± 9232/01 $15.00 www.nature.com/onc Increased expression of unmethylated CDKN2D by 5-aza-2'-deoxycytidine in human lung cancer cells Wei-Guo Zhu1, Zunyan Dai2,3, Haiming Ding1, Kanur Srinivasan1, Julia Hall3, Wenrui Duan1, Miguel A Villalona-Calero1, Christoph Plass3 and Gregory A Otterson*,1 1Division of Hematology/Oncology, Department of Internal Medicine, The Ohio State University-Comprehensive Cancer Center, Columbus, Ohio, OH 43210, USA; 2Department of Pathology, The Ohio State University-Comprehensive Cancer Center, Columbus, Ohio, OH 43210, USA; 3Division of Human Cancer Genetics, Department of Molecular Virology, Immunology and Medical Genetics, The Ohio State University-Comprehensive Cancer Center, Columbus, Ohio, OH 43210, USA DNA hypermethylation of CpG islands in the promoter Introduction region of genes is associated with transcriptional silencing. Treatment with hypo-methylating agents can Methylation of cytosine residues in CpG sequences is a lead to expression of these silenced genes. However, DNA modi®cation that plays a role in normal whether inhibition of DNA methylation in¯uences the mammalian development (Costello and Plass, 2001; expression of unmethylated genes has not been exten- Li et al., 1992), imprinting (Li et al., 1993) and X sively studied. We analysed the methylation status of chromosome inactivation (Pfeifer et al., 1990). To date, CDKN2A and CDKN2D in human lung cancer cell lines four mammalian DNA methyltransferases (DNMT) and demonstrated that the CDKN2A CpG island is have been identi®ed (Bird and Wole, 1999). Disrup- methylated, whereas CDKN2D is unmethylated. Treat- tion of the balance in methylated DNA is a common ment of cells with 5-aza-2'-deoxycytidine (5-Aza-CdR), alteration in cancer (Costello et al., 2000; Costello and an inhibitor of DNA methyltransferase 1, induced a dose Plass, 2001; Issa et al., 1993; Robertson et al., 1999). -

Cyclin-Dependent Kinases and CDK Inhibitors in Virus-Associated Cancers Shaian Tavakolian, Hossein Goudarzi and Ebrahim Faghihloo*
Tavakolian et al. Infectious Agents and Cancer (2020) 15:27 https://doi.org/10.1186/s13027-020-00295-7 REVIEW Open Access Cyclin-dependent kinases and CDK inhibitors in virus-associated cancers Shaian Tavakolian, Hossein Goudarzi and Ebrahim Faghihloo* Abstract The role of several risk factors, such as pollution, consumption of alcohol, age, sex and obesity in cancer progression is undeniable. Human malignancies are mainly characterized by deregulation of cyclin-dependent kinases (CDK) and cyclin inhibitor kinases (CIK) activities. Viruses express some onco-proteins which could interfere with CDK and CIKs function, and induce some signals to replicate their genome into host’scells.By reviewing some studies about the function of CDK and CIKs in cells infected with oncoviruses, such as HPV, HTLV, HERV, EBV, KSHV, HBV and HCV, we reviewed the mechanisms of different onco-proteins which could deregulate the cell cycle proteins. Keywords: CDK, CIKs, Cancer, Virus Introduction the key role of the phosphorylation in the entrance of Cell division is controlled by various elements [1–10], the cells to the S phase of the cell cycle [19]. especially serine/ threonine protein kinase complexes, CDK genes are classified in mammalian cells into differ- called cyclin-dependent kinases (CDKs), and cyclins, ent classes of CDKs, especially some important regulatory whose expression is prominently regulated by the bind- ones (The regulatory CDKs play important roles in medi- ing to CDK inhibitors [11, 12]. In all eukaryotic species, ating cell cycle). Each of these CDKs could interact with a these genes are classified into different families. It is specific cyclin and thereby regulating the expression of well-established that the complexes of cyclin and CDK different genes [20, 21]. -

Co-Deletion of P18 and P16 /P14 /P15 in Glioblastoma Multiforme
Review Conspirators in a Capital Crime: Co-deletion of p18INK4c and p16INK4a/p14ARF/p15INK4b in Glioblastoma Multiforme David A. Solomon,1,2 Jung-Sik Kim,2 Walter Jean,3 and Todd Waldman2 1Tumor Biology Training Program, 2Department of Oncology, Lombardi Comprehensive Cancer Center, and 3Department of Neurosurgery, Georgetown University School of Medicine, Washington, District of Columbia Abstract and showing that p18INK4c-deficient mice are tumor prone, these INK4c Glioblastoma multiforme (GBM) is one of the most dreaded recent studies have suggested that inactivation of p18 may cancer diagnoses due to its poor prognosis and the limited play a perhaps underappreciated role in human cancer patho- INK4a genesis. As such, this review will provide a brief history of the treatment options. Homozygous deletion of the p16 / INK4c p14ARF/p15INK4b locus is among the most common genetic discovery of p18 , summarize the studies that link it to cancers in mice and humans, and provide a forward-looking alterations in GBM. Two recent studies have shown that INK4c deletion and mutation of another INK4 family member, assessment of the role of p18 inactivation in the pathogenesis p18INK4c, also drives the pathogenesis of GBM. This minireview of GBM (Fig. 1). INK4c will discuss the known roles for p18 in the initiation and INK4c progression of cancer and suggest opportunities for future Discovery of p18 studies. [Cancer Res 2008;68(21):8657–60] The human p18INK4c gene was initially discovered in 1994 by Guan and colleagues (8) in a yeast two-hybrid screen to identify Introduction proteins that interacted with human cdk6. The mouse homologue was reported the following year by Hirai and colleagues (9). -

Deletions of CDKN2C in Multiple Myeloma: Biological and Clinical Implications Paola E
Human Cancer Biology Deletions of CDKN2C in Multiple Myeloma: Biological and Clinical Implications Paola E. Leone,1Brian A. Walker,1Matthew W. Jenner,1Laura Chiecchio,2 GianPaolo Dagrada,2 Rebecca K.M. Protheroe,2 David C. Johnson,1Nicholas J. Dickens,1Jose Luis Brito,1Monica Else,1 David Gonzalez,1Fiona M. Ross,2 Selina Chen-Kiang,3 Faith E. Davies,1and Gareth J. Morgan1 Abstract Purpose: Deletions of chromosome 1have been described in 7% to 40% of cases of myeloma with inconsistent clinical consequences. CDKN2C at 1p32.3 has been identified in myeloma cell lines as the potential target of the deletion.We tested the clinical impact of 1p deletion and used high-resolution techniques to define the role of CDKN2C in primary patient material. Experimental Design: We analyzed 515 cases of monoclonal gammopathy of undetermined significance (MGUS), smoldering multiple myeloma (SMM), and newly diagnosed multiple mye- loma using fluorescence in situ hybridization (FISH) for deletions of CDKN2C.In78myeloma cases, we carried out Affymetrix single nucleotide polymorphism mapping and U133 Plus 2.0 ex- pression arrays. In addition, we did mutation, methylation, andWestern blotting analysis. Results: By FISH we identified deletion of 1p32.3 (CDKN2C)in3of66MGUS(4.5%),4of39 SMM (10.3%), and 55 of 369 multiple myeloma cases (15%).We examined the impact of copy number change at CDKN2C on overall survival (OS), and found that the cases with either hemi- zygous or homozygous deletion of CDKN2C had a worse OS compared with cases that were intact at this region (22 months versus 38 months; P = 0.003). Using gene mapping we identified three homozygous deletions at 1p32.3, containing CDKN2C,allofwhichlackedexpressionof CDKN2C. -

Up-Regulation of Cyclin-Dependent Kinase 4/Cyclin D2 Expression but Down- Regulation of Cyclin-Dependent Kinase 2/Cyclin E in Testicular Germ Cell Tumors1
[CANCER RESEARCH 61, 4214–4221, May 15, 2001] Up-Regulation of Cyclin-dependent Kinase 4/Cyclin D2 Expression but Down- Regulation of Cyclin-dependent Kinase 2/Cyclin E in Testicular Germ Cell Tumors1 Bettina A. Schmidt,2 Achim Rose, Christine Steinhoff, T. Strohmeyer, Michael Hartmann, and Rolf Ackermann Department of Urology, Heinrich-Heine-University of Duesseldorf, 40225 Duesseldorf, Germany [B. A. S., A. R., C. S., R. A.], and Department of Urology, Hospital of the Armed Forces, 22099 Hamburg, Germany [M. H.] ABSTRACT causes death in about 20–30% of patients with highly advanced stages. The present state of research provides only marginal insights Testicular germ cell tumors (GCT) characteristically display two chro- into the molecular pathogenesis of these tumors and the mechanisms mosome 12 abnormalities: the isochromosome i(12p) and concomitant underlying the effectiveness of chemotherapy. deletions of the long arm. Some genes important in the control of the G -S 1 The most frequent cytogenetic observation indicative of testicular cell cycle checkpoint G1-S, i.e., cyclin-dependent kinases 2 and 4, cyclin D2 are located on this chromosomal region. Therefore, testicular GCTs were GCT is the presence of an isochromosome of the short arm of analyzed as to the expression of CDK2, CDK4, CDK6, and the expression chromosome 12, i12(p), in up to 80% of all tumors. Although this of their catalytic partners cyclins D1, D2 and E by semiquantitative finding has been confirmed by various investigators (6–8), its diag- reverse transcription-PCR. Cyclin D2, located on 12p, was overexpressed nostic and prognostic relevance is still under discussion (9, 10).