Deletions of CDKN2C in Multiple Myeloma: Biological and Clinical Implications Paola E
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Expression Profiling of KLF4
Expression Profiling of KLF4 AJCR0000006 Supplemental Data Figure S1. Snapshot of enriched gene sets identified by GSEA in Klf4-null MEFs. Figure S2. Snapshot of enriched gene sets identified by GSEA in wild type MEFs. 98 Am J Cancer Res 2011;1(1):85-97 Table S1: Functional Annotation Clustering of Genes Up-Regulated in Klf4 -Null MEFs ILLUMINA_ID Gene Symbol Gene Name (Description) P -value Fold-Change Cell Cycle 8.00E-03 ILMN_1217331 Mcm6 MINICHROMOSOME MAINTENANCE DEFICIENT 6 40.36 ILMN_2723931 E2f6 E2F TRANSCRIPTION FACTOR 6 26.8 ILMN_2724570 Mapk12 MITOGEN-ACTIVATED PROTEIN KINASE 12 22.19 ILMN_1218470 Cdk2 CYCLIN-DEPENDENT KINASE 2 9.32 ILMN_1234909 Tipin TIMELESS INTERACTING PROTEIN 5.3 ILMN_1212692 Mapk13 SAPK/ERK/KINASE 4 4.96 ILMN_2666690 Cul7 CULLIN 7 2.23 ILMN_2681776 Mapk6 MITOGEN ACTIVATED PROTEIN KINASE 4 2.11 ILMN_2652909 Ddit3 DNA-DAMAGE INDUCIBLE TRANSCRIPT 3 2.07 ILMN_2742152 Gadd45a GROWTH ARREST AND DNA-DAMAGE-INDUCIBLE 45 ALPHA 1.92 ILMN_1212787 Pttg1 PITUITARY TUMOR-TRANSFORMING 1 1.8 ILMN_1216721 Cdk5 CYCLIN-DEPENDENT KINASE 5 1.78 ILMN_1227009 Gas2l1 GROWTH ARREST-SPECIFIC 2 LIKE 1 1.74 ILMN_2663009 Rassf5 RAS ASSOCIATION (RALGDS/AF-6) DOMAIN FAMILY 5 1.64 ILMN_1220454 Anapc13 ANAPHASE PROMOTING COMPLEX SUBUNIT 13 1.61 ILMN_1216213 Incenp INNER CENTROMERE PROTEIN 1.56 ILMN_1256301 Rcc2 REGULATOR OF CHROMOSOME CONDENSATION 2 1.53 Extracellular Matrix 5.80E-06 ILMN_2735184 Col18a1 PROCOLLAGEN, TYPE XVIII, ALPHA 1 51.5 ILMN_1223997 Crtap CARTILAGE ASSOCIATED PROTEIN 32.74 ILMN_2753809 Mmp3 MATRIX METALLOPEPTIDASE -

Charles M. Perou, Phd Associate Professor Departments of Genetics
Charles M. Perou, PhD Associate Professor Departments of Genetics and Pathology Carolina Center for Genome Sciences Lineberger Comprehensive Cancer Center University of North Carolina at Chapel Hill Identification of GBM Subtypes using Gene Expression Profiling Agilent Custom Affymetrix Human Affymetrix HT-HG- 244k Exon 1.0 Array U133A Array Microarray Identify Samples and Genes represented on all 3 platforms Factor Analysis Single unified gene expression measurement for each gene 202 patients 11,681 genes Identification of GBM Subtypes Census Clustering of the 202 samples X 1740 Unsupervised clustering of 1740 genes suggests that 4 subtypes of GBM exist variably expressed genes selected using a unified gene expression measure across 3 expression platforms Core TCGA Samples (173) Gene Ontology/Pathway: with Subtype-defining genes • ProNeural: 1. nervous system development ProNeural Normal-like EGFR Mesenchymal 2. neuron differentiation (SOXs) 3. cell cycle = proliferation 4. cell adhesion molecules 5. ErbB signaling pathway FBXO3 GABRB2 • Normal‐like: SNCG NTSR2 1. nucleotide metabolic process MBP 2. neurological system process DLL3 3. axon NKX2-2, NRXN1, NRXN2 TOP2B, CDC7, MYB 4. neuron projection SOX2, SOX4, SOX10,SOX11 5. synaptic transmission ERBB3, PAK3, PAK7 NCAM1 OLIG2 • EGFR: 1. regulation of transcription 2. cell migration 3. nervous system development FGFR3 4. cell proliferation PDGFRA 5. metal ion binding EGFR AKT2 GLI2 TGFB3 • Mesenchymal: CASP8, CASP5, CASP4, CASP1 1. immune response COL8A2, COL5A1, COL1A1,COL1A2 2. receptor activity ILR4 CHI3L1 3. wound healing TRADD TLR2, TLR4, 4. cytokine and chemokine IGFBI mediated signaling pathway RELB 5. NF‐B Signaling Pathway Correlations between gene expression subtypes and clinical parameters Survival Analysis of Subtypes n=196 Subtypes are correlated with: 1. -
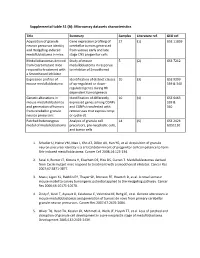
Supplemental Table S1 (A): Microarray Datasets Characteristics
Supplemental table S1 (A): Microarray datasets characteristics Title Summary Samples Literature ref. GEO ref. Acquisition of granule Gene expression profiling of 27 (1) GSE 11859 neuron precursor identity cerebellar tumors generated and Hedgehog‐induced from various early and late medulloblastoma in mice. stage CNS progenitor cells Medulloblastomas derived Study of mouse 5 (2) GSE 7212 from Cxcr6 mutant mice medulloblastoma in response respond to treatment with to inhibitor of Smoothened a Smoothened inhibitor Expression profiles of Identification of distinct classes 10 (3) GSE 9299 mouse medulloblastoma of up‐regulated or down‐ 339 & 340 regulated genes during Hh dependent tumorigenesis Genetic alterations in Identification of differently 10 (4) GSE 6463 mouse medulloblastomas expressed genes among CGNPs 339 & and generation of tumors and CGNPs transfected with 340 from cerebellar granule retroviruses that express nmyc neuron precursors or cyclin‐d1 Patched heterozygous Analysis of granule cell 14 (5) GSE 2426 model of medulloblastoma precursors, pre‐neoplastic cells, GDS1110 and tumor cells 1. Schuller U, Heine VM, Mao J, Kho AT, Dillon AK, Han YG, et al. Acquisition of granule neuron precursor identity is a critical determinant of progenitor cell competence to form Shh‐induced medulloblastoma. Cancer Cell 2008;14:123‐134. 2. Sasai K, Romer JT, Kimura H, Eberhart DE, Rice DS, Curran T. Medulloblastomas derived from Cxcr6 mutant mice respond to treatment with a smoothened inhibitor. Cancer Res 2007;67:3871‐3877. 3. Mao J, Ligon KL, Rakhlin EY, Thayer SP, Bronson RT, Rowitch D, et al. A novel somatic mouse model to survey tumorigenic potential applied to the Hedgehog pathway. Cancer Res 2006;66:10171‐10178. -

Tagging Single Nucleotide Polymorphisms in Cell Cycle Control Genes and Susceptibility to Invasive Epithelial Ovarian Cancer
Research Article Tagging Single Nucleotide Polymorphisms in Cell Cycle Control Genes and Susceptibility to Invasive Epithelial Ovarian Cancer Simon A. Gayther,1 Honglin Song,2 Susan J. Ramus,1 Susan Kru¨ger Kjaer,4 Alice S. Whittemore,5 Lydia Quaye,1 Jonathan Tyrer,2 Danielle Shadforth,2 Estrid Hogdall,4 Claus Hogdall,6 Jan Blaeker,7 Richard DiCioccio,8 Valerie McGuire,5 Penelope M. Webb,9 Jonathan Beesley,9 Adele C. Green,9 David C. Whiteman,9 The Australian Ovarian Cancer Study Group,9 The Australian Cancer Study (Ovarian Cancer),20 Marc T. Goodman,10 Galina Lurie,10 Michael E. Carney,10 Francesmary Modugno,10 Roberta B. Ness,11 Robert P. Edwards,12 Kirsten B. Moysich,7 Ellen L. Goode,13 Fergus J. Couch,13 Julie M. Cunningham,13 Thomas A. Sellers,14 Anna H. Wu,15 Malcolm C. Pike,15 Edwin S. Iversen,16 Jeffrey R. Marks,16 Montserrat Garcia-Closas,17 Louise Brinton,17 Jolanta Lissowska,18 Beata Peplonska,19 Douglas F. Easton,3 Ian Jacobs,1 Bruce A.J. Ponder,2 Joellen Schildkraut,16 C. Leigh Pearce,15 Georgia Chenevix-Trench,9 Andrew Berchuck,16 Paul D.P. Pharoah,2 and on behalf of the Ovarian Cancer Association Consortium 1Translational Research Laboratories, University College London, London, United Kingdom; 2Cancer Research United Kingdom Department of Oncology and 3Cancer Research United Kingdom Genetic Epidemiology Unit, University of Cambridge, Strangeways Research Laboratory, Cambridge, United Kingdom; 4Danish Cancer Society, Copenhagen, Denmark; 5Stanford University School of Medicine, Stanford, California; 6University of Copenhagen, -

Screening for Differentially Expressed Genes Between Left‑And Right‑Sided
ONCOLOGY LETTERS 6: 353-358, 2013 Screening for differentially expressed genes between left‑ and right‑sided colon carcinoma by microarray analysis HONG ZHU1, TIAN-CONG WU1, WEI-QIONG CHEN1, LI-JUN ZHOU1, YUE WU1, LIANG ZENG2 and HAI-PING PEI3 1Department of Oncology, Xiangya Hospital, Central South University, Changsha, Hunan 410008; 2Department of Pathology, Hunan Tumor Hospital, Changsha, Hunan 410013; 3Department of Gastrointestinal Surgery, Xiangya Hospital, Central South University, Changsha, Hunan 410008, P.R. China Received December 25, 2012; Accepted May 14, 2013 DOI: 10.3892/ol.2013.1414 Abstract. Left-sided colon carcinoma (LSCC) and right-sided and RSCC. These insights may therefore serve as a basis for colon carcinoma (RSCC) differ in their genetic susceptibili- the identification of novel colon cancer markers and thera- ties to neoplastic transformation. The present study identified peutic targets. 11 genes that were differentially expressed in LSCC and RSCC by expression profiling with microarray analysis. Compared Introduction with RSCC, the human genes for L-lactate dehydrogenase B chain (LDHB), cyclin-dependent kinase 4 inhibitor D Colon cancer is a significant cause of cancer‑related morbidity (CDKN2D), phosphatidylinositol-4-phosphate-3-kinase C2 and mortality, and is the third most fatal malignancy domain-containing subunit α (PI3KC2α), protocadherin fat 1 worldwide (1). In China and other economically developing (FAT; a human protein that closely resembles the Drosophila countries, colon cancer incidence rates have increased over tumor suppressor, fat) and dual specificity protein phospha- the past 20 years; most likely due to changes in the envi- tase 2 (DUSP2) were upregulated in LSCC. By contrast, ronment, individual lifestyle and nutritional habits (2). -

CDKN2C-Null Leiomyosarcoma: a Novel
original reports CDKN2C-Null Leiomyosarcoma: A Novel, Genomically Distinct Class of TP53/ RB1–Wild-Type Tumor With Frequent CIC Genomic Alterations and 1p/19q-Codeletion Erik A. Williams, MD1; Radwa Sharaf, PhD1; Brennan Decker, MD, PhD2; Adrienne J. Werth, MD3; Helen Toma, MD3; Meagan Montesion, PhD1; Ethan S. Sokol, PhD1; Dean C. Pavlick, BS1; Nikunj Shah, BS1; Kevin Jon Williams, MD4; Jeffrey M. Venstrom, MD1; Brian M. Alexander, MD, MPH1; Jeffrey S. Ross, MD1,5; Lee A. Albacker, PhD1; Douglas I. Lin, MD, PhD1; Shakti H. Ramkissoon, MD, PhD1,6; and Julia A. Elvin, MD, PhD1 abstract PURPOSE Leiomyosarcoma (LMS) harbors frequent mutations in TP53 and RB1 but few actionable genomic alterations. Here, we searched for recurrent actionable genomic alterations in LMS that occur in the absence of common untreatable oncogenic drivers. METHODS Tissues from 276,645 unique advanced cancers, including 2,570 uterine and soft tissue LMS, were sequenced by hybrid-capture–based next-generation DNA and RNA sequencing/comprehensive genomic profiling of up to 406 genes. We characterized clinicopathologic features of relevant patient cases. RESULTS Overall, 77 LMS exhibited homozygous copy loss of CDKN2C at chromosome 1p32.3 (3.0% of LMS). Genomic alterations (GAs) in TP53, RB1, and ATRX were rare compared with the remainder of the LMS cohort (11.7% v 73.4%, 0% v 54.5%, 2.6% v 24.5%, respectively; all P , .0001). CDKN2C-null LMS patient cases were significantly enriched for GAs in CIC (40.3% v 1.4%) at 19q13.2, CDKN2A (46.8% v 7.0%), and RAD51B (16.9% v 1.7%; all P , .0001). -

Increased Expression of Unmethylated CDKN2D by 5-Aza-2'-Deoxycytidine in Human Lung Cancer Cells
Oncogene (2001) 20, 7787 ± 7796 ã 2001 Nature Publishing Group All rights reserved 0950 ± 9232/01 $15.00 www.nature.com/onc Increased expression of unmethylated CDKN2D by 5-aza-2'-deoxycytidine in human lung cancer cells Wei-Guo Zhu1, Zunyan Dai2,3, Haiming Ding1, Kanur Srinivasan1, Julia Hall3, Wenrui Duan1, Miguel A Villalona-Calero1, Christoph Plass3 and Gregory A Otterson*,1 1Division of Hematology/Oncology, Department of Internal Medicine, The Ohio State University-Comprehensive Cancer Center, Columbus, Ohio, OH 43210, USA; 2Department of Pathology, The Ohio State University-Comprehensive Cancer Center, Columbus, Ohio, OH 43210, USA; 3Division of Human Cancer Genetics, Department of Molecular Virology, Immunology and Medical Genetics, The Ohio State University-Comprehensive Cancer Center, Columbus, Ohio, OH 43210, USA DNA hypermethylation of CpG islands in the promoter Introduction region of genes is associated with transcriptional silencing. Treatment with hypo-methylating agents can Methylation of cytosine residues in CpG sequences is a lead to expression of these silenced genes. However, DNA modi®cation that plays a role in normal whether inhibition of DNA methylation in¯uences the mammalian development (Costello and Plass, 2001; expression of unmethylated genes has not been exten- Li et al., 1992), imprinting (Li et al., 1993) and X sively studied. We analysed the methylation status of chromosome inactivation (Pfeifer et al., 1990). To date, CDKN2A and CDKN2D in human lung cancer cell lines four mammalian DNA methyltransferases (DNMT) and demonstrated that the CDKN2A CpG island is have been identi®ed (Bird and Wole, 1999). Disrup- methylated, whereas CDKN2D is unmethylated. Treat- tion of the balance in methylated DNA is a common ment of cells with 5-aza-2'-deoxycytidine (5-Aza-CdR), alteration in cancer (Costello et al., 2000; Costello and an inhibitor of DNA methyltransferase 1, induced a dose Plass, 2001; Issa et al., 1993; Robertson et al., 1999). -

Co-Deletion of P18 and P16 /P14 /P15 in Glioblastoma Multiforme
Review Conspirators in a Capital Crime: Co-deletion of p18INK4c and p16INK4a/p14ARF/p15INK4b in Glioblastoma Multiforme David A. Solomon,1,2 Jung-Sik Kim,2 Walter Jean,3 and Todd Waldman2 1Tumor Biology Training Program, 2Department of Oncology, Lombardi Comprehensive Cancer Center, and 3Department of Neurosurgery, Georgetown University School of Medicine, Washington, District of Columbia Abstract and showing that p18INK4c-deficient mice are tumor prone, these INK4c Glioblastoma multiforme (GBM) is one of the most dreaded recent studies have suggested that inactivation of p18 may cancer diagnoses due to its poor prognosis and the limited play a perhaps underappreciated role in human cancer patho- INK4a genesis. As such, this review will provide a brief history of the treatment options. Homozygous deletion of the p16 / INK4c p14ARF/p15INK4b locus is among the most common genetic discovery of p18 , summarize the studies that link it to cancers in mice and humans, and provide a forward-looking alterations in GBM. Two recent studies have shown that INK4c deletion and mutation of another INK4 family member, assessment of the role of p18 inactivation in the pathogenesis p18INK4c, also drives the pathogenesis of GBM. This minireview of GBM (Fig. 1). INK4c will discuss the known roles for p18 in the initiation and INK4c progression of cancer and suggest opportunities for future Discovery of p18 studies. [Cancer Res 2008;68(21):8657–60] The human p18INK4c gene was initially discovered in 1994 by Guan and colleagues (8) in a yeast two-hybrid screen to identify Introduction proteins that interacted with human cdk6. The mouse homologue was reported the following year by Hirai and colleagues (9). -

P63 Modulates the Expression of the WDFY2 Gene Which Is Implicated in Cancer Regulation and Limb Development
Bioscience Reports (2019) 39 BSR20192114 https://doi.org/10.1042/BSR20192114 Research Article P63 modulates the expression of the WDFY2 gene which is implicated in cancer regulation and limb development 1 2 1,* 1 2 3 Paola Monti , Yari Ciribilli , Giorgia Foggetti , Paola Menichini , Alessandra Bisio , Serena Cappato , Downloaded from https://portlandpress.com/bioscirep/article-pdf/39/12/BSR20192114/862944/bsr-2019-2114.pdf by guest on 07 February 2020 Alberto Inga2, Maria Teresa Divizia4, Margherita Lerone4, Renata Bocciardi3,4 and Gilberto Fronza1 1Mutagenesis and Cancer Prevention Unit, IRCCS Ospedale Policlinico San Martino, Largo R. Benzi, 10, Genoa 16132, Italy; 2Department of Cellular, Computational and Integrative Biology (CIBio), University of Trento, Via Sommarive 9, Povo (TN) 38123, Italy; 3Department of Neurosciences, Rehabilitation, Ophthalmology, Genetics, Maternal and Child Health (DiNOGMI), University of Genoa, Largo P. Daneo, 3, Genoa 16132, Italy; 4Medical Genetics Unit, IRCCS Istituto Giannina Gaslini, Via G. Gaslini, 5, Genoa 16147, Italy Correspondence: Paola Monti ([email protected]) or Gilberto Fronza ([email protected]) TP63 is a member of the TP53 gene family, sharing a common gene structure that produces two groups of mRNAs’ encoding proteins with different N-terminal regions (N and TA iso- forms); both transcripts are also subjected to alternative splicing mechanisms at C-terminus, generating a variety of isoforms. p63 is a master regulator of epidermal development and homoeostasis as well as an important player in tumorigenesis and cancer progression with both oncogenic and tumour suppressive roles. A number of studies have aimed at the identi- fication of p63 target genes, allowing the dissection of the molecular pathways orchestrated by the different isoforms. -

Autoimmunity in a Mouse Model of Lupus Deficiency Promotes B1a
Cyclin-Dependent Kinase Inhibitor Cdkn2c Deficiency Promotes B1a Cell Expansion and Autoimmunity in a Mouse Model of Lupus This information is current as Hari-Hara S. K. Potula, Zhiwei Xu, Leilani Zeumer, Allison of September 25, 2021. Sang, Byron P. Croker and Laurence Morel J Immunol 2012; 189:2931-2940; Prepublished online 15 August 2012; doi: 10.4049/jimmunol.1200556 http://www.jimmunol.org/content/189/6/2931 Downloaded from Supplementary http://www.jimmunol.org/content/suppl/2012/08/15/jimmunol.120055 Material 6.DC1 http://www.jimmunol.org/ References This article cites 42 articles, 14 of which you can access for free at: http://www.jimmunol.org/content/189/6/2931.full#ref-list-1 Why The JI? Submit online. • Rapid Reviews! 30 days* from submission to initial decision by guest on September 25, 2021 • No Triage! Every submission reviewed by practicing scientists • Fast Publication! 4 weeks from acceptance to publication *average Subscription Information about subscribing to The Journal of Immunology is online at: http://jimmunol.org/subscription Permissions Submit copyright permission requests at: http://www.aai.org/About/Publications/JI/copyright.html Email Alerts Receive free email-alerts when new articles cite this article. Sign up at: http://jimmunol.org/alerts The Journal of Immunology is published twice each month by The American Association of Immunologists, Inc., 1451 Rockville Pike, Suite 650, Rockville, MD 20852 Copyright © 2012 by The American Association of Immunologists, Inc. All rights reserved. Print ISSN: 0022-1767 Online ISSN: 1550-6606. The Journal of Immunology Cyclin-Dependent Kinase Inhibitor Cdkn2c Deficiency Promotes B1a Cell Expansion and Autoimmunity in a Mouse Model of Lupus Hari-Hara S. -

(CDKN2D) (NM 001800) Human Recombinant Protein Product Data
OriGene Technologies, Inc. 9620 Medical Center Drive, Ste 200 Rockville, MD 20850, US Phone: +1-888-267-4436 [email protected] EU: [email protected] CN: [email protected] Product datasheet for TP314065 p19 INK4d (CDKN2D) (NM_001800) Human Recombinant Protein Product data: Product Type: Recombinant Proteins Description: Recombinant protein of human cyclin-dependent kinase inhibitor 2D (p19, inhibits CDK4) (CDKN2D), transcript variant 1 Species: Human Expression Host: HEK293T Tag: C-Myc/DDK Predicted MW: 17.5 kDa Concentration: >50 ug/mL as determined by microplate BCA method Purity: > 80% as determined by SDS-PAGE and Coomassie blue staining Buffer: 25 mM Tris.HCl, pH 7.3, 100 mM glycine, 10% glycerol Preparation: Recombinant protein was captured through anti-DDK affinity column followed by conventional chromatography steps. Storage: Store at -80°C. Stability: Stable for 12 months from the date of receipt of the product under proper storage and handling conditions. Avoid repeated freeze-thaw cycles. RefSeq: NP_001791 Locus ID: 1032 UniProt ID: P55273, A0A024R796 RefSeq Size: 1416 Cytogenetics: 19p13.2 RefSeq ORF: 498 Synonyms: INK4D; p19; p19-INK4D This product is to be used for laboratory only. Not for diagnostic or therapeutic use. View online » ©2021 OriGene Technologies, Inc., 9620 Medical Center Drive, Ste 200, Rockville, MD 20850, US 1 / 2 p19 INK4d (CDKN2D) (NM_001800) Human Recombinant Protein – TP314065 Summary: The protein encoded by this gene is a member of the INK4 family of cyclin-dependent kinase inhibitors. This protein has been shown to form a stable complex with CDK4 or CDK6, and prevent the activation of the CDK kinases, thus function as a cell growth regulator that controls cell cycle G1 progression. -

Loss of the Nuclear Wnt Pathway Effector TCF7L2 Promotes Migration and Invasion of Human Colorectal Cancer Cells
Oncogene (2020) 39:3893–3909 https://doi.org/10.1038/s41388-020-1259-7 ARTICLE Loss of the nuclear Wnt pathway effector TCF7L2 promotes migration and invasion of human colorectal cancer cells 1,2 1 3 4,5,6 2,5 Janna Wenzel ● Katja Rose ● Elham Bavafaye Haghighi ● Constanze Lamprecht ● Gilles Rauen ● 1 7 3,8,9 1,2,5 Vivien Freihen ● Rebecca Kesselring ● Melanie Boerries ● Andreas Hecht Received: 27 September 2019 / Revised: 3 March 2020 / Accepted: 4 March 2020 / Published online: 20 March 2020 © The Author(s) 2020. This article is published with open access Abstract The transcription factor TCF7L2 is indispensable for intestinal tissue homeostasis where it transmits mitogenic Wnt/ β-Catenin signals in stem and progenitor cells, from which intestinal tumors arise. Yet, TCF7L2 belongs to the most frequently mutated genes in colorectal cancer (CRC), and tumor-suppressive functions of TCF7L2 were proposed. This apparent paradox warrants to clarify the role of TCF7L2 in colorectal carcinogenesis. Here, we investigated TCF7L2 dependence/independence of CRC cells and the cellular and molecular consequences of TCF7L2 loss-of-function. By genome editing we achieved complete TCF7L2 inactivation in several CRC cell lines without loss of viability, showing that fi 1234567890();,: 1234567890();,: CRC cells have widely lost the strict requirement for TCF7L2. TCF7L2 de ciency impaired G1/S progression, reminiscent of the physiological role of TCF7L2. In addition, TCF7L2-negative cells exhibited morphological changes, enhanced migration, invasion, and collagen adhesion, albeit the severity of the phenotypic alterations manifested in a cell-line-specific fashion. To provide a molecular framework for the observed cellular changes, we performed global transcriptome profiling and identified gene-regulatory networks in which TCF7L2 positively regulates the proto-oncogene MYC, while repressing the cell cycle inhibitors CDKN2C/CDKN2D.