UNIVERSITY of CALIFORNIA RIVERSIDE Studies
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-
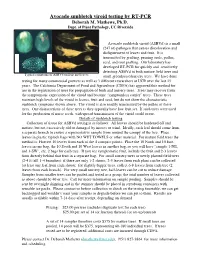
Avocado Sunblotch Viroid Testing by RT-PCR Deborah M
Avocado sunblotch viroid testing by RT-PCR Deborah M. Mathews, Ph.D. Dept. of Plant Pathology, UC Riverside Avocado sunblotch viroid (ASBVd) is a small (247 nt) pathogen that causes discoloration and disfigurement of leaves and fruit. It is transmitted by grafting, pruning tools, pollen, seed, and root grafting. Our laboratory has developed RT-PCR for quickly and sensitively detecting ASBVd in both mature field trees and Typical symptoms of ASBVd on fruit and leaves small greenhouse/nursery trees. We have done testing for many commercial growers as well as 3 different researchers at UCR over the last 15 years. The California Department of Food and Agriculture (CDFA) has approved this method for use in the registration of trees for propagation of buds and nursery trees. Trees may recover from the symptomatic expression of the viroid and become “symptomless carrier” trees. These trees maintain high levels of the viroid in leaves, fruit and seed, but do not show the characteristic sunblotch symptoms shown above. The viroid is also readily transmitted by the pollen of these trees. One characteristic of these trees is they typically have low fruit set. If such trees were used for the production of nurse seeds, widespread transmission of the viroid could occur. Details of sunblotch testing Collection of tissue for ASBVd testing is as follows: All leaves should be hardened off and mature, but not excessively old or damaged by insects or wind. Ideally, each leaf should come from a separate branch to ensure a representative sample from around the canopy of the tree. Place leaves in plastic ziplock bags with NO WET TOWELS or other material. -

Detection of Avocado Sunblotch Viroid and Estimation of Infection Among Accessions in the National Germplasm Collection for Avocado
Proc. Fla. State Hort. Soc. 109:235-237. 1996. DETECTION OF AVOCADO SUNBLOTCH VIROID AND ESTIMATION OF INFECTION AMONG ACCESSIONS IN THE NATIONAL GERMPLASM COLLECTION FOR AVOCADO Catherine M. Running and Raymond J. Schnell National Germplasm Repository U. S. Department of Agriculture Agricultural Research Service 13601 Old Cutler Road, Miami, FL 33158 David N. Kuhn Florida International University Dept. of Biological Sciences Miami, FL 33199 Additional index words. Persea Americana, RT-PCR, viroid, DNA, RNA, sequence variation, germplasm. ABSTRACT Reverse transcription-polymerase chain reaction (RTPCR) was used to determine the incidence of infection by avocado sunblotch viroid (ASBVd) in the germplasm collection at the National Germplasm Repository at Miami (NGR-Mia). Of the 429 avocado plants growing at the repository, 81 (18.9%) are infected with the viroid. The 429 plants represent 237 accessions. There are multiple plants of some accessions and for 42 accessions (17%) every plant is infected with the viroid. There was no apparent relationship between host race and the incidence of infection. Symptoms of sunblotch disease on avocado (Persea Americana Mill.) manifest as a general decline of tree vigor with sunken, yellow areas on the fruit that lessen its marketability. The disease was first described as infectious, rather than physiological, by Home and Parker (1931). The infectious agent has since been determined to be an RNA viroid known as Avocado Sunblotch Viroid (ASBVd) (Dale and Allen, 1979; Thomas and Mohammed, 1979). The viroid is transmitted by budding and grafting, including natural root grafting (Home etal, 1941; Whitsell, 1952), seed (Wallace and Drake, 1962), and pollen (Desjardins et al., 1979). -

THE MOLECULAR CHARACTERIZATION of the VIRUS and VIRUS-LIKE AGENTS PRESENT in TA TAO 5 GERMPLASM of PRUNUS PERSICA Diana Marini Clemson University, [email protected]
Clemson University TigerPrints All Dissertations Dissertations 12-2007 THE MOLECULAR CHARACTERIZATION OF THE VIRUS AND VIRUS-LIKE AGENTS PRESENT IN TA TAO 5 GERMPLASM OF PRUNUS PERSICA Diana Marini Clemson University, [email protected] Follow this and additional works at: https://tigerprints.clemson.edu/all_dissertations Part of the Agronomy and Crop Sciences Commons Recommended Citation Marini, Diana, "THE MOLECULAR CHARACTERIZATION OF THE VIRUS AND VIRUS-LIKE AGENTS PRESENT IN TA TAO 5 GERMPLASM OF PRUNUS PERSICA " (2007). All Dissertations. 143. https://tigerprints.clemson.edu/all_dissertations/143 This Dissertation is brought to you for free and open access by the Dissertations at TigerPrints. It has been accepted for inclusion in All Dissertations by an authorized administrator of TigerPrints. For more information, please contact [email protected]. THE MOLECULAR CHARACTERIZATION OF THE VIRUS AND VIRUS-LIKE AGENTS PRESENT IN TA TAO 5 GERMPLASM OF PRUNUS PERSICA A Dissertation Presented to the Graduate School of Clemson University In Partial Fulfillment of the Requirements for the Degree Doctor of Philosophy Plant and Environmental Sciences by Diana Beatriz Marini December 2007 Accepted by: Dr. Simon W. Scott, Committee Chair Dr. N. Dwight Camper Dr. Steven N. Jeffers Dr. Gregory L. Reighard ABSTRACT Peach production in the southeastern United States is limited by late spring freezes. Ta Tao 5 germplasm, used either as an interstem or by chip bud inoculation, has been shown to delay bloom and avoid the effects of these late freezes. The growth modification is graft transmissible and the germplasm has been found to be infected with ACLSV, APruV-3, and PLMVd. Using a combination of PCR, cloning, and sequencing techniques, a molecular characterization of the three graft-transmissible agents present in Ta Tao 5 has been completed. -

Replication of Avocado Sunblotch Viroid in the Cyanobacterium Nostoc Sp. PCC 7120
atholog P y & nt a M Latifi et al., J Plant Pathol Microbiol 2016, 7:4 l i P c f r o o DOI: 10.4172/2157-7471.1000341 b l i Journal of a o l n o r g u y o J ISSN: 2157-7471 Plant Pathology & Microbiology Research Article Open Access Replication of Avocado Sunblotch Viroid in the Cyanobacterium Nostoc Sp. PCC 7120 Amel Latifi1*, Christophe Bernard1, Laura da Silva2, Yannick Andéol3, Amine Elleuch4, Véronique Risoul1, Jacques Vergne2 and Marie-Christine Maurel2 1Aix Marseille University, CNRS, UMR Chemistry Laboratory Bacterial 7283. 31 Chemin Joseph Aiguier 13009 Marseille Cedex 20, France 2Institute of Systematics, Evolution, Biodiversity ( ISyEB ), CNRS, MNHN , UPMC, EPHE, UPMC Sorbonne University, 57 rue Cuvier, PO Box 50, F-75005 Paris, France 3Team Enzymology of RNA, UR6, UPMC Sorbonne University, 75252 Paris Cedex 05 France 4Laboratory of Plant Biotechnology, Faculty of Sciences of Sfax, University of Sfax, BP 1171, 3000 Sfax, Tunisia Abstract Viroids are small infectious RNA molecules that replicate in plants via RNA-RNA replication processes. The molecular mechanism responsible for this replication has attracted great interest, and studies on this topic have yielded interesting biological findings on the processes in which RNA is involved. Viroids belonging to the Avsunviroidae family replicate in the chloroplasts of infected hosts. It has by now been established that chloroplasts and cyanobacteria share a common have ancestor. In view of this phylogenetic relationship, we investigated whether a member of the Avsunviroidae family could be replicated in a cyanobacterium. The results obtained here show that Avocado Sunblotch Viroid (ASBVd) RNA is able to replicate in the filamentous cyanobacterium Nostoc PCC 7120. -

Recognizing Avocado Sunblotch Disease
RECOGNIZING AVOCADO SUNBLOTCH DISEASE 1J. A. Dodds, 1D Mathews, 2M. L. Arpaia and 3G. W. Witney. 1Dept. Plant Pathology, University of California, Riverside, CA 92521. A. 2Dept. Botany and Plant Sciences, University of California, Riverside, CA 92521. 3California Avocado Commission, 1251 E. Dyer Rd., Suite 210, Santa Ana, CA 92705. Avocado sunblotch is a disease which has been known for over 60 years. Sunblotch is characterized by abnormal tree growth, reduced yield and a high proportion of small or misshapen fruit. Specific symptoms may be observed on fruit, leaves and stems. Fruit may show overall distortion, sometimes with sunken yellow and red depressions on the surface. These fruit are graded as standards and receive lower returns from the packinghouse (Figure A). Foliar symptoms can include a general thinning of B. the canopy, with individual leaves showing bleaching (white patches), variegation (yellow and green patches), and distortion (Figure B, C). Stems may also show discoloration with yellow, white or pink streaks (Figure D), and in extreme cases may be completely chlorotic (yellow/white). Infected trees often appear stunted and somewhat sprawling. It is possible for trees to “recover” from the disease. The “recovered” tree will have no apparent visual disease symptoms but it still carries the viroid. These trees are termed “symptomless” carriers. Such trees typically have very low fruit yield or at times may set heavy crops of small fruit. If symptomless trees are topworked with disease-free material, the topworked material will become infected and can exhibit classical sunblotch symptoms revealing C. the presence of sunblotch. If a symptomless tree is subject to a stress such as fire or is stumped, the regrowth may once again exhibit sunblotch symptoms. -

The Avocado Sunblotch Viroid: an Invisible Foe of Avocado
viruses Review The Avocado Sunblotch Viroid: An Invisible Foe of Avocado José Ramón Saucedo Carabez 1,*, Daniel Téliz Ortiz 2 , Moisés Roberto Vallejo Pérez 3 and Hugo Beltrán Peña 4 1 Departamento de Investigación Aplicada-Driscolls, Libramiento Sur #1620, Jacona 59833, Michoacán, Mexico 2 Postgrado en Fitosanidad-Fitopatología, Colegio de Postgraduados, Km. 36.5, Montecillo, Texcoco 56230, Estado de México, Mexico; [email protected] 3 Consejo Nacional de Ciencia y Tecnología-Universidad Autónoma de San Luis Potosí, Álvaro Obregón #64, San Luis Potosí 78000, San Luis Potosí, Mexico; [email protected] 4 Departamento de Ciencias Naturales y Exactas-Fitopatología, Universidad Autónoma de Occidente UR Los Mochis, Boulevard Macario Gaxiola, Los Mochis 81223, Sinaloa, Mexico; [email protected] * Correspondence: [email protected] Received: 1 May 2019; Accepted: 25 May 2019; Published: 29 May 2019 Abstract: This review collects information about the history of avocado and the economically important disease, avocado sunblotch, caused by the avocado sunblotch viroid (ASBVd). Sunblotch symptoms are variable, but the most common in fruits are irregular sunken areas of white, yellow, or reddish color. On severely affected fruits, the sunken areas may become necrotic. ASBVd (type species Avocado sunblotch viroid, family Avsunviroidae) replicates and accumulates in the chloroplast, and it is the smallest plant pathogen. This pathogen is a circular single-stranded RNA of 246–251 nucleotides. ASBVd has a restricted host range and only few plant species of the family Lauraceae have been confirmed experimentally as additional hosts. The most reliable method to detect ASBVd in the field is to identify symptomatic fruits, complemented in the laboratory with reliable and sensitive molecular techniques to identify infected but asymptomatic trees. -

Viroid-2018 Book of Abstracts
Viroid-2018 Viroid-2018 International Conference on Viroids and Viroid-Like RNAs 5-7 July 2018, Valencia, Spain Polytechnic City of Innovation, Universitat Politècnica de València Avenida de los Naranjos, 46022 Valencia, Spain http://www.ibmcp.upv.es/viroid-2018/ [email protected] Book of Abstracts Organizer: José-Antonio Daròs Instituto de Biología Molecular y Celular de Plantas (IBMCP) CSIC-Universitat Politècnica de Valencia Avenida de los Naranjos, 46022Valencia, Spain [email protected] http://personales.upv.es/jadaros/ Updated version, 3 July 2018 1 Viroid-2018 INDEX Supporting institutions and contributing companies ……………… 3 Conference venue …………………………………….……….……… 4 Conference program ………………………………………………… 6 List of participants ………………………………………...………… 12 Abstracts, oral presentations …………….………….……….……… 16 Abstracts, poster presentations …………….………….……….…… 47 Index of authors ……………………………………………………… 70 Notes …………………………..……………………….……………… 72 2 Viroid-2018 SUPPORTING INSTITUTIONS Grants from Ministerio de Ciencia, Innovación y Universidades (Spain): BIO2014-54269-R, BIO2017-83184-R and BIO2017-91865-EXP Grant from Generalitat Valenciana: AORG/2018/114 CONTRIBUTING COMPANIES Biotest Diagnosticos S.L., http://www.biotestdiagnosticos.es/ Cultek, http://www.cultek.com/ Durviz, http://durviz.com/ Épica, http://epicasl.com/ Fisher Scientific, http://www.fishersci.com GenoChem, http://geno-chem.com/ IDT, http://www.idtdna.com/pages NZYTech, http://www.nzytech.com/ Werfen, http://es.werfen.com/ VWR, http://www.vwr.com/ 3 Viroid-2018 CONFERENCE VENUE Campus Universitat Politècnica de Valencia, corner between Avenida de los Naranjos and Calle Ingeniero Fausto Elio, Access J, Ciudad Politécnica de la Innovación (CPI), Red Prism. 4 Viroid-2018 5 Viroid-2018 CONFERENCE PROGRAM Thursday, July 5th 09:00-10:00. Registration at CPI Red Prism 10:00-10:20. Opening Chairman: Robert A. -

Results of the 2009 Asbvd Survey of Avocado Accessions of the National Germplasm Collection in Florida
Proc. Fla. State Hort. Soc. 123:5–7. 2010. Results of the 2009 ASBVd Survey of Avocado Accessions of the National Germplasm Collection in Florida CeCile l. Tondo*, Raymond J. SChnell, and david n. Kuhn National Germplasm Repository, USDA ARS, 13601 Old Cutler Road, Miami, FL 33158 AdditionAl index words. Persea americana, avocado diseases, avocado sunblotch viroid, germplasm collection management, viroid indexing The presence of avocado sunblotch viroid (ASBVd) infection among the avocado (Persea americana Mill.) accessions in the National Germplasm Repository at Miami (NGR–Miami) was established in previous studies. An ASBVd specific reverse transcription-polymerase chain reaction (RT-PCR) protocol was used to detect the viroid. Surveys performed in 1996 and in 2000 found that the proportion of ASBVd positive accessions remained unchanged at 19% during that time period. The object of the current study was to assess the spread of infection, if any, and the rate and direction of transmission. For this purpose the collection was screened again for ASBVd in 2009. The germplasm collection increased from 403 to 505 trees. Fifty newly infected trees were detected. Forty-eight percent of the newly infected plants were found to be adjacent to previously infected plants, adjacent to plots from which infected plants had been removed, or adjacent to other newly infected plants that are adjacent to previously infected plants or contaminated plots. No pattern in direction of spread was discerned for non-adjacent new infections. The proportion of plants found to be positive for the viroid in the current study is 21%. Fourteen plants previously found to be infected were found to be negative in this survey. -

The Avocado Conundrum
The Avocado Conundrum Track: Management Education and Teaching Cases Key words: Avocado, Agribusiness, International trade, WTO Abstract This case illustrates the challenges that the Government of Costa Rica faces of whether and how to manage a trade solution under the WTO framework, due to its decision of suspending the phytosanitaty import certificates of avocados from Mexico. The aim is to stimulate a discussion about international trade and agribusiness. This is a case study suitable for use with graduate students in Masters of Business Administration (MBA), Master of Agribusiness, Executive Master of Business Administration or any other post-graduate program related. 2 On 22 April 2015, the State Phytosanitary Service (SFE) from the Costa Rican Government released an official resolution, through which it temporally suspended the emission of phytosanitary import certificates for avocados from eight countries— Australia, Spain, Ghana, Guatemala, Israel, Mexico, South Africa, and Venezuela—plus the state of Florida in the United States1. The SFE stated its decision was an urgent action to protect the national territory and its local producers from the Avocado Sun Blotch Viroid (ASBVd), a plague affecting the quality and yield of avocado crops in these countries. However, the decision from the SFE was subject to considerable criticism from different national and international stakeholders due to many inconsistencies in the process. The SFE had a puzzling organizational structure and role within the national regulatory system. First of all, the SFE held an independent legal status and was not attached to the Ministry of Agriculture (MAG). It charged a fee for their services and maintained self-sustainable operations. -

Detection of Avocado Sunblotch Viroid by Polymerase Chain Reaction (Pcr)
California Avocado Society 1997 Yearbook 81: 91-96 DETECTION OF AVOCADO SUNBLOTCH VIROID BY POLYMERASE CHAIN REACTION (PCR) DM. Mathews, J.A. Heick, and J.A. Dodds Department of Plant Pathology, University of California, Riverside, CA 92521 Abstract Several avocado trees were identified which had been previously biologically indexed as either negative or positive for avocado sunblotch viroid (ASBVd). Each was then tested by RT-PCR for the presence of ASBVd. Of the 52 biologically indexed negative trees, all were subsequently scored as negative for ASBVd by PCR. All positive controls were scored as ASBVd positive by PCR. When testing multiple samples from single trees, all replications were scored as ASBVd positive with one exception, which was scored as a weak positive with further analysis. Two ASBVd positive field trees were identified from approximately 40 that were tested from South Coast Field Station. We have made improvements to the existing PCR test such that extraction time and handling are greatly reduced. INTRODUCTION Avocado sunblotch viroid (ASBVd) is a small RNA-based pathogen which causes sunken yellow or red areas on the fruit, stem discoloration, and bleaching, variegation, and distortion on the leaves of avocado (Figure I)1'2-3. It is transmitted by grafting of infected budwood, pollen, and seed; but it has no known vector. The incidence of ASBVd in commercial trees is low at this time due to a successful certification program of budwood that was performed in the past. ASBVd was traditionally tested for by performing multiple graft inoculations (up to 40 grafts per tree), and observing for 1-4 years for the development of characteristic symptoms. -

Replication of Avocado Sunblotch Viroid in the Cyanobacterium Nostoc Sp. PCC 7120
atholog P y & nt a M Latifi et al., J Plant Pathol Microbiol 2016, 7:4 l i P c f r o o b http://dx.doi.org/10.4172/2157-7471.1000341 l i Journal of a o l n o r g u y o J ISSN: 2157-7471 Plant Pathology & Microbiology Research Article Open Access Replication of Avocado Sunblotch Viroid in the Cyanobacterium Nostoc Sp. PCC 7120 Amel Latifi1*, Christophe Bernard1, Laura da Silva2, Yannick Andéol3, Amine Elleuch4, Véronique Risoul1, Jacques Vergne2 and Marie-Christine Maurel2 1Aix Marseille University, CNRS, UMR Chemistry Laboratory Bacterial 7283. 31 Chemin Joseph Aiguier 13009 Marseille Cedex 20, France 2Institute of Systematics, Evolution, Biodiversity ( ISyEB ), CNRS, MNHN , UPMC, EPHE, UPMC Sorbonne University, 57 rue Cuvier, PO Box 50, F-75005 Paris, France 3Team Enzymology of RNA, UR6, UPMC Sorbonne University, 75252 Paris Cedex 05 France 4Laboratory of Plant Biotechnology, Faculty of Sciences of Sfax, University of Sfax, BP 1171, 3000 Sfax, Tunisia Abstract Viroids are small infectious RNA molecules that replicate in plants via RNA-RNA replication processes. The molecular mechanism responsible for this replication has attracted great interest, and studies on this topic have yielded interesting biological findings on the processes in which RNA is involved. Viroids belonging to the Avsunviroidae family replicate in the chloroplasts of infected hosts. It has by now been established that chloroplasts and cyanobacteria share a common have ancestor. In view of this phylogenetic relationship, we investigated whether a member of the Avsunviroidae family could be replicated in a cyanobacterium. The results obtained here show that Avocado Sunblotch Viroid (ASBVd) RNA is able to replicate in the filamentous cyanobacterium Nostoc PCC 7120. -

AVOCADO SUNBLOTCH VIROID (Asbvd) in CELL AND
AVOCADO SUNBLOTCH VIROID (ASBVd) IN CELL AND TISSUE CULTURES OF AVOCADO ( Persea americana Mill.) By ISIDRO ELIAS SUAREZ PADRON A DISSERTATION PRESENTED TO THE GRADUATE SCHOOL OF THE UNIVERSITY OF FLORIDA IN PARTIAL FULFILLMENT OF THE REQUIREMENTS FOR THE DEGREE OF DOCTOR OF PHILOSOPHY UNIVERSITY OF FLORIDA 2003 Copyright 2003 by Isidro E. Suarez Padron To my wife Sahara, for her love and comprehensive patience during these difficult years of sacrifice, and to my sons, Sebastian and Elias David for my absence in the most precious moments. A mi esposa Sahara por su amor y comprensiva paciencia durante estos dificiles anos de sacrificio, y a mis hijos Sebastian y Elias David por mi ausencia en los momentos mas hermosos de sus vidas. AKNOWLEDGMENTS I wish to express my deepest gratitude to Dr. Richard Litz, my major professor, for his unlimited support, advice, help, patience and time, and particularly for his direction in the completion of this dissertation. His encouragement, guidance and permanent stimulation for better work will always be remembered. I want to extend my gratitude for their help, guidance, and stimulating discussion to the members of my graduate committee, Drs. Raymond J. Schnell, Dennis Gray, Paul Lyrene, and Nigel Harrison. Sincere thanks go to Mrs. Pamela Moon for her help with statistical analyses, graphic designs and lab support. The invaluable cooperation of Drs. David N. Kuhn, Martha Heath and Alan Meerow, and Cecile Olano, Wilber Quintanilla, Cheryl Krol, Mike Winterstein, Jason Clayton, Emilio Power, Tyrone McArthy, Wilhelmina Wasik and Nannette Langiven at the Plant Sciences lab of the ARS-USDA in Miami, FL, will always be appreciated.