Thioredoxin Targets Fundamental Processes in a Methane-Producing Archaeon, Methanocaldococcus Jannaschii
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

The 2014 Golden Gate National Parks Bioblitz - Data Management and the Event Species List Achieving a Quality Dataset from a Large Scale Event
National Park Service U.S. Department of the Interior Natural Resource Stewardship and Science The 2014 Golden Gate National Parks BioBlitz - Data Management and the Event Species List Achieving a Quality Dataset from a Large Scale Event Natural Resource Report NPS/GOGA/NRR—2016/1147 ON THIS PAGE Photograph of BioBlitz participants conducting data entry into iNaturalist. Photograph courtesy of the National Park Service. ON THE COVER Photograph of BioBlitz participants collecting aquatic species data in the Presidio of San Francisco. Photograph courtesy of National Park Service. The 2014 Golden Gate National Parks BioBlitz - Data Management and the Event Species List Achieving a Quality Dataset from a Large Scale Event Natural Resource Report NPS/GOGA/NRR—2016/1147 Elizabeth Edson1, Michelle O’Herron1, Alison Forrestel2, Daniel George3 1Golden Gate Parks Conservancy Building 201 Fort Mason San Francisco, CA 94129 2National Park Service. Golden Gate National Recreation Area Fort Cronkhite, Bldg. 1061 Sausalito, CA 94965 3National Park Service. San Francisco Bay Area Network Inventory & Monitoring Program Manager Fort Cronkhite, Bldg. 1063 Sausalito, CA 94965 March 2016 U.S. Department of the Interior National Park Service Natural Resource Stewardship and Science Fort Collins, Colorado The National Park Service, Natural Resource Stewardship and Science office in Fort Collins, Colorado, publishes a range of reports that address natural resource topics. These reports are of interest and applicability to a broad audience in the National Park Service and others in natural resource management, including scientists, conservation and environmental constituencies, and the public. The Natural Resource Report Series is used to disseminate comprehensive information and analysis about natural resources and related topics concerning lands managed by the National Park Service. -

Diversity of Understudied Archaeal and Bacterial Populations of Yellowstone National Park: from Genes to Genomes Daniel Colman
University of New Mexico UNM Digital Repository Biology ETDs Electronic Theses and Dissertations 7-1-2015 Diversity of understudied archaeal and bacterial populations of Yellowstone National Park: from genes to genomes Daniel Colman Follow this and additional works at: https://digitalrepository.unm.edu/biol_etds Recommended Citation Colman, Daniel. "Diversity of understudied archaeal and bacterial populations of Yellowstone National Park: from genes to genomes." (2015). https://digitalrepository.unm.edu/biol_etds/18 This Dissertation is brought to you for free and open access by the Electronic Theses and Dissertations at UNM Digital Repository. It has been accepted for inclusion in Biology ETDs by an authorized administrator of UNM Digital Repository. For more information, please contact [email protected]. Daniel Robert Colman Candidate Biology Department This dissertation is approved, and it is acceptable in quality and form for publication: Approved by the Dissertation Committee: Cristina Takacs-Vesbach , Chairperson Robert Sinsabaugh Laura Crossey Diana Northup i Diversity of understudied archaeal and bacterial populations from Yellowstone National Park: from genes to genomes by Daniel Robert Colman B.S. Biology, University of New Mexico, 2009 DISSERTATION Submitted in Partial Fulfillment of the Requirements for the Degree of Doctor of Philosophy Biology The University of New Mexico Albuquerque, New Mexico July 2015 ii DEDICATION I would like to dedicate this dissertation to my late grandfather, Kenneth Leo Colman, associate professor of Animal Science in the Wool laboratory at Montana State University, who even very near the end of his earthly tenure, thought it pertinent to quiz my knowledge of oxidized nitrogen compounds. He was a man of great curiosity about the natural world, and to whom I owe an acknowledgement for his legacy of intellectual (and actual) wanderlust. -

Genomics and Energy and Environmental Science Poster 2011
Omics: Exploring the Molecular Universe within Biological Systems Instead of studying one or a few genes or proteins at a time, “omics” collectively Proteomics is the analysis of the proteome—the complete set describes the comprehensive analysis of genes, RNA transcripts, proteins, of proteins expressed by a cell or population of cells. Proteins, metabolites, and other molecules present in a biological system. Ongoing the workhorse molecules of life, catalyze biochemical reactions; advances in computing power and automated technologies for DNA sequencing provide structural support; and recognize, bind, or transport and experiments continue to improve our ability to analyze increasing numbers other molecules throughout the cell. Hundreds of dierent types of molecules and how they function as a system. Systems biology integrates the of proteins can be expressed at a time, and most are part of large data from various omic analyses using computational tools to build predictive complexes made up of models of biological systems. many proteins and other molecules. Transcriptomics is the analysis of the transcriptome—the complete set of RNA Hundreds to molecules present in a cell or population of thousands of different cells. RNA, which is much less stable than protein types exist within DNA, is constantly being synthesized and a cell. This large subunit of a ribosome contains about then broken down to facilitate rapid 3,000 RNA nucleotides changes in paerns of protein expression (gray) and 30 protein that occur as an organism dynamically chains (gold). responds to its environment. In addition to the three major classes of RNA (mRNA, tRNA, and ribosomal RNA), single- stranded RNA is very exible and can fold into complex shapes that carry out specic functions. -

Biological Diversity in the Patent System
Biological Diversity in the Patent System Paul Oldham1,2*, Stephen Hall1,3, Oscar Forero1,4 1 ESRC Centre for Economic and Social Aspects of Genomics (Cesagen), Lancaster University, Lancaster, United Kingdom, 2 Institute of Advanced Studies, United Nations University, Yokohama, Japan, 3 One World Analytics, Lancaster, United Kingdom, 4 Centre for Development, Environment and Policy, SOAS, University of London, London, United Kingdom Abstract Biological diversity in the patent system is an enduring focus of controversy but empirical analysis of the presence of biodiversity in the patent system has been limited. To address this problem we text mined 11 million patent documents for 6 million Latin species names from the Global Names Index (GNI) established by the Global Biodiversity Information Facility (GBIF) and Encyclopedia of Life (EOL). We identified 76,274 full Latin species names from 23,882 genera in 767,955 patent documents. 25,595 species appeared in the claims section of 136,880 patent documents. This reveals that human innovative activity involving biodiversity in the patent system focuses on approximately 4% of taxonomically described species and between 0.8–1% of predicted global species. In this article we identify the major features of the patent landscape for biological diversity by focusing on key areas including pharmaceuticals, neglected diseases, traditional medicines, genetic engineering, foods, biocides, marine genetic resources and Antarctica. We conclude that the narrow focus of human innovative activity and ownership of genetic resources is unlikely to be in the long term interest of humanity. We argue that a broader spectrum of biodiversity needs to be opened up to research and development based on the principles of equitable benefit-sharing, respect for the objectives of the Convention on Biological Diversity, human rights and ethics. -

Phylogenetics of Archaeal Lipids Amy Kelly 9/27/2006 Outline
Phylogenetics of Archaeal Lipids Amy Kelly 9/27/2006 Outline • Phlogenetics of Archaea • Phlogenetics of archaeal lipids • Papers Phyla • Two? main phyla – Euryarchaeota • Methanogens • Extreme halophiles • Extreme thermophiles • Sulfate-reducing – Crenarchaeota • Extreme thermophiles – Korarchaeota? • Hyperthermophiles • indicated only by environmental DNA sequences – Nanoarchaeum? • N. equitans a fast evolving euryarchaeal lineage, not novel, early diverging archaeal phylum – Ancient archael group? • In deepest brances of Crenarchaea? Euryarchaea? Archaeal Lipids • Methanogens – Di- and tetra-ethers of glycerol and isoprenoid alcohols – Core mostly archaeol or caldarchaeol – Core sometimes sn-2- or Images removed due to sn-3-hydroxyarchaeol or copyright considerations. macrocyclic archaeol –PMI • Halophiles – Similar to methanogens – Exclusively synthesize bacterioruberin • Marine Crenarchaea Depositional Archaeal Lipids Biological Origin Environment Crocetane methanotrophs? methane seeps? methanogens, PMI (2,6,10,15,19-pentamethylicosane) methanotrophs hypersaline, anoxic Squalane hypersaline? C31-C40 head-to-head isoprenoids Smit & Mushegian • “Lost” enzymes of MVA pathway must exist – Phosphomevalonate kinase (PMK) – Diphosphomevalonate decarboxylase – Isopentenyl diphosphate isomerase (IPPI) Kaneda et al. 2001 Rohdich et al. 2001 Boucher et al. • Isoprenoid biosynthesis of archaea evolved through a combination of processes – Co-option of ancestral enzymes – Modification of enzymatic specificity – Orthologous and non-orthologous gene -

Archaeology of Eukaryotic DNA Replication
Downloaded from http://cshperspectives.cshlp.org/ on September 25, 2021 - Published by Cold Spring Harbor Laboratory Press Archaeology of Eukaryotic DNA Replication Kira S. Makarova and Eugene V. Koonin National Center for Biotechnology Information, National Library of Medicine, National Institutes of Health, Bethesda, Maryland 20894 Correspondence: [email protected] Recent advances in the characterization of the archaeal DNA replication system together with comparative genomic analysis have led to the identification of several previously un- characterized archaeal proteins involved in replication and currently reveal a nearly com- plete correspondence between the components of the archaeal and eukaryotic replication machineries. It can be inferred that the archaeal ancestor of eukaryotes and even the last common ancestor of all extant archaea possessed replication machineries that were compa- rable in complexity to the eukaryotic replication system. The eukaryotic replication system encompasses multiple paralogs of ancestral components such that heteromeric complexes in eukaryotes replace archaeal homomeric complexes, apparently along with subfunctionali- zation of the eukaryotic complex subunits. In the archaea, parallel, lineage-specific dupli- cations of many genes encoding replication machinery components are detectable as well; most of these archaeal paralogs remain to be functionally characterized. The archaeal rep- lication system shows remarkable plasticity whereby even some essential components such as DNA polymerase and single-stranded DNA-binding protein are displaced by unrelated proteins with analogous activities in some lineages. ouble-stranded DNA is the molecule that Okazaki fragments (Kornberg and Baker 2005; Dcarries genetic information in all cellular Barry and Bell 2006; Hamdan and Richardson life-forms; thus, replication of this genetic ma- 2009; Hamdan and van Oijen 2010). -

A Korarchaeal Genome Reveals Insights Into the Evolution of the Archaea
A korarchaeal genome reveals insights into the evolution of the Archaea James G. Elkinsa,b, Mircea Podarc, David E. Grahamd, Kira S. Makarovae, Yuri Wolfe, Lennart Randauf, Brian P. Hedlundg, Ce´ line Brochier-Armaneth, Victor Kunini, Iain Andersoni, Alla Lapidusi, Eugene Goltsmani, Kerrie Barryi, Eugene V. Koonine, Phil Hugenholtzi, Nikos Kyrpidesi, Gerhard Wannerj, Paul Richardsoni, Martin Kellerc, and Karl O. Stettera,k,l aLehrstuhl fu¨r Mikrobiologie und Archaeenzentrum, Universita¨t Regensburg, D-93053 Regensburg, Germany; cBiosciences Division, Oak Ridge National Laboratory, Oak Ridge, TN 37831; dDepartment of Chemistry and Biochemistry, University of Texas, Austin, TX 78712; eNational Center for Biotechnology Information, National Library of Medicine, National Institutes of Health, Bethesda, MD 20894; fDepartment of Molecular Biophysics and Biochemistry, Yale University, New Haven, CT 06520; gSchool of Life Sciences, University of Nevada, Las Vegas, NV 89154; hLaboratoire de Chimie Bacte´rienne, Unite´ Propre de Recherche 9043, Centre National de la Recherche Scientifique, Universite´de Provence Aix-Marseille I, 13331 Marseille Cedex 3, France; iU.S. Department of Energy Joint Genome Institute, Walnut Creek, CA 94598; jInstitute of Botany, Ludwig Maximilians University of Munich, D-80638 Munich, Germany; and kInstitute of Geophysics and Planetary Physics, University of California, Los Angeles, CA 90095 Communicated by Carl R. Woese, University of Illinois at Urbana–Champaign, Urbana, IL, April 2, 2008 (received for review January 7, 2008) The candidate division Korarchaeota comprises a group of uncul- and sediment samples from Obsidian Pool as an inoculum. The tivated microorganisms that, by their small subunit rRNA phylog- cultivation system supported the stable growth of a mixed commu- eny, may have diverged early from the major archaeal phyla nity of hyperthermophilic bacteria and archaea including an or- Crenarchaeota and Euryarchaeota. -
An Estimate of the Elemental Composition of Luca
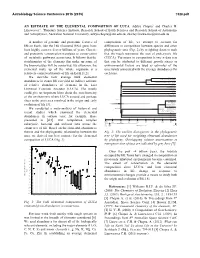
Astrobiology Science Conference 2015 (2015) 7328.pdf AN ESTIMATE OF THE ELEMENTAL COMPOSITION OF LUCA. Aditya Chopra1 and Charles H. Lineweaver1, 1Planetary Science Institute, Research School of Earth Sciences and Research School of Astronomy and Astrophysics, Australian National University, [email protected], [email protected] A number of genomic and proteomic features of composition of life, we attempt to account for life on Earth, like the 16S ribosomal RNA gene, have differences in composition between species and other been highly conserved over billions of years. Genetic phylogenetic taxa (Fig. 2) by weighting datasets such and proteomic conservation translates to conservation that the result represents the root of prokaryotic life of metabolic pathways across taxa. It follows that the (LUCA). Variations in composition between data sets stoichiometry of the elements that make up some of that can be attributed to different growth stages or the biomolecules will be conserved. By extension, the environmental factors are used as estimates of the elemental make up of the whole organism is a uncertainty associated with the average abundances for relatively conserved feature of life on Earth [1,2]. each taxa. We describe how average bulk elemental Euryarchaeota 1a Methanococci, Methanobacteria, Methanopyri 7 Euryarchaeota 1b abundances in extant life can yield an indirect estimate 6 Thermoplasmata, Methanomicrobia, Halobacteria, Archaeoglobi Euryarchaeota 2 4 Thermococci of relative abundances of elements in the Last Crenarchaeota Sulfolobus, Thermoproteus ? Thaumarchaeota Universal Common Ancestor (LUCA). The results Cenarchaeum ? Korarchaeota ARMAN could give us important hints about the stoichiometry ? 2 Archaeal Richmond Mine Acidophilic Nanoorganisms Nanoarchaeota of the environment where LUCA existed and perhaps Archaea Terrabacteria clues to the processes involved in the origin and early Actinobacteria, Deinococcus-Thermus, 1 Cyanobacteria, Life Chloroflexi, 9 evolution of life [3]. -

Title Genomic Analysis of the Marine Hyperthermophilic Archaeon
Genomic analysis of the marine hyperthermophilic archaeon Title Aeropyrum( Dissertation_全文 ) Author(s) Daifuku, Takashi Citation 京都大学 Issue Date 2015-03-23 URL https://doi.org/10.14989/doctor.k19034 学位規則第9条第2項により要約公開; 許諾条件により本文 Right は2019-08-01に公開 Type Thesis or Dissertation Textversion ETD Kyoto University 1. General introduction Chapter 1 General introduction Gene repertoires and genome organizations differ between closely related microbial organisms depending on the ecological characteristics of each habitat (Cohan and Koeppel 2008). The cyanobacterial Prochlorococcus spp. account for a significant fraction of primary production in the ocean (Goericke and Welschmeyer 1993) and show physiological features relevant to the different ecological niches within a stratified oceanic water column (Moore et al. 1998; West et al. 2001). The whole-genomic comparisons of the Prochlorococcus spp. strains show gross signatures according to this niche differentiation (Rocap et al. 2003). Alpha-proteobacterium Pelagibacter ubique which belongs to the SAR11 clade in the phylogenetic tree based on the 16S rRNA gene is the most abundant microorganism in the ocean (Morris et al. 2002). The genomes of the SAR11 isolates are highly conserved in the core genes that are common to all strains (Medini et al. 2005) and show synteny (the conservation of DNA sequence and gene order) (Bentley and Parkhill 2004). However, variations exist among genes for phosphorus metabolism, glycolysis, and C1 metabolism, suggesting that adaptive specialization in nutrient resource utilization is important for niche partitioning (Grote et al. 2012). This adaptation at the genomic level was also observed in archaea. The members of the genus Pyrococcus are anaerobic and hyperthermophilic archaea (Fiala and Stetter 1 1. -

Universidad Politécnica De Madrid Escuela Técnica Superior De Ingeniería Agronómica Alimentaria Y De Biosistemas Departamento De Biotecnología-Biología Vegetal
UNIVERSIDAD POLITÉCNICA DE MADRID ESCUELA TÉCNICA SUPERIOR DE INGENIERÍA AGRONÓMICA ALIMENTARIA Y DE BIOSISTEMAS DEPARTAMENTO DE BIOTECNOLOGÍA-BIOLOGÍA VEGETAL TESIS DOCTORAL Caracterización de una nueva proteína hipertermófila NifB y su papel en el mecanismo de síntesis de NifB-co, precursor del grupo metálico de FeMo-co Author: Alessandro Scandurra Director: Prof. Luis Manuel Rubio Herrero Co-director: Dr. Simon Jean Marius Arragain i ii UNIVERSIDAD POLITÉCNICA DE MADRID ESCUELA TÉCNICA SUPERIOR DE INGENIERÍA AGRONÓMICA ALIMENTARIA Y DE BIOSISTEMAS DEPARTAMENTO DE BIOTECNOLOGÍA-BIOLOGÍA VEGETAL Memoria presentada por D. Alessandro Scandurra para optar al grado de Doctor Director Dr. Luis Manuel Rubio Herrero Profesor Titular UPM Madrid, 2017 © This copy oF the thesis has been supplied on condition that anyone who consults it is understood to recognize that its copyright rest with the author and that no quotation From the thesis, or any inFormation derived therefrom may be published without the author’s prior, written consent. I II AcKnowledgments As each journey, also this one has an end (Thanks goodness). It was a long trip, with several changes oF direction. Like For many other people, the PhD adventure was not easy, but I have to admit that I was lucky enough to meet really unique people, that became special Friendships. First oF all, I want to thank my Family, especially my mother and my Father, who always push me to go Forward in my life and to Follow my dreams, ready to pay the sacrifice oF the distance. This expirience has helped me to understand how easy is to be a son and how challenging is to be a parent. -

Research Article Role of Mn2+ and Compatible Solutes in the Radiation Resistance of Thermophilic Bacteria and Archaea
Hindawi Publishing Corporation Archaea Volume 2012, Article ID 845756, 11 pages doi:10.1155/2012/845756 Research Article Role of Mn2+ and Compatible Solutes in the Radiation Resistance of Thermophilic Bacteria and Archaea Kimberly M. Webb and Jocelyne DiRuggiero Department of Biology, Johns Hopkins University, Baltimore, MD 21218, USA Correspondence should be addressed to Jocelyne DiRuggiero, [email protected] Received 25 July 2012; Revised 18 September 2012; Accepted 13 October 2012 Academic Editor: Yoshizumi Ishino Copyright © 2012 K. M. Webb and J. DiRuggiero. This is an open access article distributed under the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited. Radiation-resistant bacteria have garnered a great deal of attention from scientists seeking to expose the mechanisms underlying their incredible survival abilities. Recent analyses showed that the resistance to ionizing radiation (IR) in the archaeon Halobacterium salinarum is dependent upon Mn-antioxidant complexes responsible for the scavenging of reactive oxygen species (ROS) generated by radiation. Here we examined the role of the compatible solutes trehalose, mannosylglycerate, and di-myo- inositol phosphate in the radiation resistance of aerobic and anaerobic thermophiles. We found that the IR resistance of the thermophilic bacteria Rubrobacter xylanophilus and Rubrobacter radiotolerans was highly correlated to the accumulation of high intracellular concentration of trehalose in association with Mn, supporting the model of Mn2+-dependent ROS scavenging in the aerobes. In contrast, the hyperthermophilic archaea Thermococcus gammatolerans and Pyrococcus furiosus did not contain significant amounts of intracellular Mn, and we found no significant antioxidant activity from mannosylglycerate and di-myo- inositol phosphate in vitro. -

Counts Metabolic Yr10.Pdf
Advanced Review Physiological, metabolic and biotechnological features of extremely thermophilic microorganisms James A. Counts,1 Benjamin M. Zeldes,1 Laura L. Lee,1 Christopher T. Straub,1 Michael W.W. Adams2 and Robert M. Kelly1* The current upper thermal limit for life as we know it is approximately 120C. Microorganisms that grow optimally at temperatures of 75C and above are usu- ally referred to as ‘extreme thermophiles’ and include both bacteria and archaea. For over a century, there has been great scientific curiosity in the basic tenets that support life in thermal biotopes on earth and potentially on other solar bodies. Extreme thermophiles can be aerobes, anaerobes, autotrophs, hetero- trophs, or chemolithotrophs, and are found in diverse environments including shallow marine fissures, deep sea hydrothermal vents, terrestrial hot springs— basically, anywhere there is hot water. Initial efforts to study extreme thermo- philes faced challenges with their isolation from difficult to access locales, pro- blems with their cultivation in laboratories, and lack of molecular tools. Fortunately, because of their relatively small genomes, many extreme thermo- philes were among the first organisms to be sequenced, thereby opening up the application of systems biology-based methods to probe their unique physiologi- cal, metabolic and biotechnological features. The bacterial genera Caldicellulosir- uptor, Thermotoga and Thermus, and the archaea belonging to the orders Thermococcales and Sulfolobales, are among the most studied extreme thermo- philes to date. The recent emergence of genetic tools for many of these organ- isms provides the opportunity to move beyond basic discovery and manipulation to biotechnologically relevant applications of metabolic engineering.